Abstract
Aim:
To examine the potential correlation between serum REST corepressor 3 (RCOR3) level and the outcome of patients with hepatitis B.
Methods:
Concanavalin A (ConA)-induced mouse hepatitis model was used. The mRNA level of RCOR3 in mouse liver was measured using GeneChip array and real-time PCR. One hundred seventy-seven patients with hepatitis B and 34 healthy individuals were categorized into six groups including mild chronic hepatitis, moderate chronic hepatitis B, severe hepatitis B (SHB), cirrhosis, hepatocellular carcinoma (HCC) and healthy control. Serum levels of human RCOR3 were measured using ELISA.
Results:
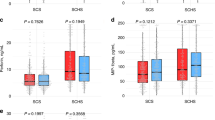
In the mouse hepatitis model, the mRNA level of RCOR3 in liver was reduced early after exposure to ConA, then increased after 6 h of exposure. There was no significant difference in the serum RCOR3 level between the mild chronic hepatitis B and the control groups. The serum RCOR3 level was significantly increased in the moderate chronic hepatitis B group, but significantly reduced in SHB, cirrhosis and HCC groups, as compared with the control group. Moreover, the serum RCOR3 levels in SHB, cirrhosis and liver cancer patients were significantly lower than those in the patients with moderate chronic hepatitis B and with mild chronic hepatitis B. Rank correlation analysis revealed a significant correlation between serum RCOR3 level and total bilirubin (r=-0.305, P<0.01). There was no significant correlation between RCOR3 on one hand, and alanine transaminase (r=0.014, P>0.05) or aspartate transaminase (r=-0.079, P>0.05) on the other hand.
Conclusion:
Serum RCOR3 level may reflect the degree of liver damage, which might be a potential biomarker for the outcome of patients with hepatitis B.
Similar content being viewed by others
Introduction
Chronic infection with hepatitis B virus (HBV) is a common and serious liver disease that affects individuals worldwide. Current treatment of chronic hepatitis B is disappointingly limited. It is reported that there are more than 2 billion HBV carriers at present globally, and among them, about 360 million patients suffer from chronic hepatitis, cirrhosis or hepatocellular carcinoma (HCC)1, 2.
RCOR3 is a member of the REST (repressor element-1-silencing transcription factor) corepressor (CoREST) family that functions as a corepressor for REST, which regulates neuronal gene expression and determines the fate of neural stem cells. The CoREST family consists of RCOR1, also known as CoREST, RCOR2, and RCOR3. REST (also known as neuron-restrictive silencer factor, NRSF) is an essential vertebrate zinc finger transcription factor3, which can bind to the conservative sequence of neuron-restrictive silencer element DNA (NRSE DNA)4. The REST gene is highly expressed in non-neuronal cells and neuronal precursor cells, but it is expressed much less in differentiated neurons5.
To date, it has been shown that REST does not work in-dependently. The dynamic and multiplex function of REST is achieved through the REST-dependent recruitment of distinct modular macromolecular complexes. Thus, REST serves as a hub for recruiting multiple chromatin-modifying proteins, including multiple histone deacetylases (HDACs) and lysine specific demethylases (LSD; for example, LSD1)6, 7, 8. The N-terminal domain of NRSF can combine with mSin3A/B, and the C-terminal domain can bind to CoREST9, 10. The histone modifiers are able to interact directly with either REST or its corepressors, such as CoREST11 or mSin39, 12, 13, 14, and thus block the transcription of certain genes15.
In a previous study, we obtained the gene expression profile of livers from mice in a concanavalin A (ConA)-induced hepatitis model by GeneChip array16, 17. ConA-induced hepatic injury in mice has long been established as a model of immunologically induced hepatocyte injury because its histological features highly resemble those of viral or drug-induced acute hepatitis in humans18. The hepatocyte injury is associated with massive hepatocellular degeneration and lymphoid infiltration of the liver19. The hepatic injury induced by ConA depends mainly upon T-cell activation20, which raises the plasma levels of various cytokines such as tumor necrosis factor-alpha (TNF-α), interferon (IFN)-γ, interleukin (IL)-6, etc21. Hence, we used the ConA-induced hepatitis model and selected microarray to explore the pathogenic mechanism of human fulminant hepatitis. In this study, 26 differentially expressed genes associated with hepatitis B were selected. However, only the expression of RCOR3 attracted our interest, for RCOR3 was suitable to be used as a biomarker. Considering the clinical significance of RCOR3 remains largely unknown, we herein studied the serum RCOR3 levels of patients with hepatitis B to explore the potential correlation between serum RCOR3 and the outcome of hepatitis B patients.
Materials and methods
ConA-induced hepatitis model in mice
Male BALB/c mice, weighing about 25 g, were housed under standard animal laboratory conditions in the Animal Center of Zhejiang Academy of Medical Sciences. This study was approved by the local Ethical Committee of the Academy. The mice were subjected to a tail vein injection of ConA (type V, Sigma Chemical Co, USA) at 20 mg/kg body weight, and phosphate-buffered saline (PBS) was used as a control. The mice were sacrificed at 1, 3, or 6 h separately. All animals received humane care in accordance with the guidelines established by the National Science Council of China. Mouse liver samples were collected and fixed in 4% phosphate-buffered paraformaldehyde. Paraffin tissue sections were prepared and subjected to hematoxylin-eosin (H&E) staining for standard histological examination. The extent of liver injury was also evaluated by determining serum ALT, AST, STB, and LDH using the standard Reitman-Frankel method. The results can be seen in our previous articles16, 17.
Microarray analysis
Total RNA extracted from mouse liver samples was reverse transcribed and labeled with biotin. The biotinylated cDNAs were then hybridized to 12 individual Affymetrix GeneChips Mouse 430 2.0 Arrays (Affymetrix, Inc, Santa Clara, CA, USA), which contain 45101 probes in total according to the manufacturer's instructions. Significant differentially expressed genes at each time point were filtered by analysis of variance (ANOVA) and corrected by the random variance model (RVM)22. To assess whether the significance of one particular sample occurred by random chance, the false discovery rate (FDR) was estimated to determine whether the differences of certain genes were truly significant by repeating the comparison test and the permutation test 1000 times. A P-value<0.05 and FDR<0.05 were considered the threshold for true significance.
Real-time PCR
The original amount of RCOR3 mRNA was measured by real-time PCR with a SYBR Green PCR Kit (Applied Biosystems, Foster City, CA, USA) using the ABI PRISM 7900 sequence detector (Applied Biosystems, Foster City, CA, USA). A volume of 20 μL of amplification reactions contained 2×SYBR Green Realtime PCR Master Mix, 0.4 μmol/L primers, and 1 μL of template cDNA. The mRNA of the housekeeping gene β-actin was used as an endogenous control. The sequences for the primers used are as follows: RCOR3 forward primer, 5′-AGTATCCCAGATGCCAAAT-3′; RCOR3 reverse primer, 5′-GGAAAGGGCGTGAAGTTA-3′; β-actin forward primer, 5′-CGTTGACATCCGTAAAGACC-3′; and β-actin reverse primer, 5′-AACAGTCCGCCTAGAAGCAC-3′. Thermal cycling was carried out for 5 min at 95 °C, followed by 40 cycles of 30 s at 95 °C, 30 s at 57 °C, and 30 s at 72 °C. We performed the PCR amplifications in duplicate for each sample and the expression of RCOR3 was normalized against β-actin. Analysis of variance (ANOVA) was used to compare the differences of gene expression levels between groups. A P-value<0.05 was considered statistically significant.
Patients and samples
One hundred and seventy-seven patients with hepatitis B were selected from the Infectious Department of the First Affiliated Hospital of College of Medicine, Zhejiang University, referring to the guideline of prevention and treatment for chronic hepatitis B (2010 version)23. Thirty-four healthy people who had undertaken a medical examination in 2009 at the First Affiliated Hospital of College of Medicine, Zhejiang University, were included as the control group. The demographic characteristics of all subjects are summarized in Table 1. All patients provided informed consent prior to participating in the study; the experimental protocols used in this study were approved by the Ethics Committee of the First Affiliated Hospital of College of Medicine, Zhejiang University.
ELISA
Commercially available ELISA kits (Groundwork Biotechnology Diagnosticate Ltd) were utilized to determine serum levels of human RCOR3 following the manufacturer's instruction. ELISA samples were read at the wavelength of 450 nm and a standard curve was constructed using the OD values. Assays were performed in duplicate.
Statistical analysis
All of the data were processed by SPSS 13.0 software and presented as means±SD. An ANOVA and LSD test were used for comparisons among the groups and between paired data, respectively. When the data were not normally distributed, the Mann-Whitney U test and the one-way non parametric ANOVA (Kruskal Wallis test) were used to compare quantitative variables between two groups of observation and in more than two groups of data, respectively. Spearman rank correlation test was used for correlation analysis.
Results
Expression of RCOR3 in ConA-induced mouse hepatitis
A GeneChip array was performed to screen differentially expressed genes in a model of ConA-induced mouse hepatitis. The data were analyzed by ANOVA and corrected with the randomized variance model. As a result, 1473 genes with significant P-values and a false discovery rate (FDR) of less than 0.05 were identified; among these genes, we selected 26 differentially expressed genes that were associated with hepatitis B. However, other genes, with the exception of RCOR3, were either validated to be meaningless or not secretory proteins and cannot be used as biomarkers. The expression of RCOR3 attracted our interest, and its differential expression was very notable. Intriguingly, the expression of RCOR3 was decreased at 1 h and 3 h post ConA exposure, but increased at the 6 h time point (Figure 1).
GeneChip array data on the expression of RCOR3 in a model of ConA-induced mouse hepatitis at the indicated time points. The horizontal axis represents time, and the vertical axis shows the mRNA levels of RCOR3. cP<0.01 compared with the 0 h group; fP<0.01 compared with the 1 h group; iP<0.01 compared with the 3 h group.
Validation of RCOR3 expression by real-time PCR
Real-time PCR was performed to verify the expression of RCOR3 in a model of ConA-induced mouse hepatitis. As shown in Figure 2, RCOR3 mRNA was significantly induced at the 6 h time point in comparison with the 0 h group (P<0.05). Meanwhile, there was a significant increase in the mRNA level of RCOR3 at the 3 h and 6 h time points compared with the 1 h group (P<0.05 and P<0.01, respectively), and these results (Figure 2) showed some differences in comparison with the results of the GeneChips Arrays, which showed that the expression of RCOR3 was decreased at 1 h and 3 h post ConA exposure, but increased at the 6 h time point. However, the mRNA level of RCOR3 in real-time PCR array was significantly reduced at the 1 h time point, but was induced in the liver after ConA exposure at the 3 h and 6 h time points in comparison with the 0 h group. Despite the minor difference with the results of the GeneChips Arrays, the overall trend was that the mRNA level of RCOR3 was reduced early after ConA exposure and then induced after 6 h of exposure.
Comparison of hepatitis-associated markers among patients with hepatitis B
As shown in Table 2, the serum ALT levels of mild chronic hepatitis B patients, moderate chronic hepatitis B patients, severe hepatitis B (SHB) patients, cirrhosis patients and HCC patients were significantly increased compared with the healthy control group. Meanwhile, there was a significant increase in serum ALT levels in patients with moderate chronic hepatitis B and SHB in comparison with patients with mild chronic hepatitis B. Moreover, serum ALT levels in cirrhosis and liver cancer patients remarkably declined compared with moderate chronic hepatitis B patients and severe hepatitis patients. Compared with the healthy control group, the serum AST levels of mild chronic hepatitis B patients, moderate chronic hepatitis B patients, SHB patients, cirrhosis patients and HCC patients were evidently increased. Meanwhile, there was a significant increase in serum AST levels in patients with moderate chronic hepatitis B, SHB, cirrhosis and liver cancer in comparison with patients with mild chronic hepatitis B. In addition, the serum TB levels in moderate chronic hepatitis B patients, SHB patients, cirrhosis patients and HCC patients were significantly higher than that of the healthy control group. An increase of serum TB levels in patients with moderate chronic hepatitis B, SHB, cirrhosis and liver cancer was observed in comparison with the mild chronic hepatitis B patients. In comparison with the moderate chronic hepatitis B patients, the serum TB levels of SHB, cirrhosis and liver cancer patients were remarkably increased. Moreover, the serum TB levels in cirrhosis and liver cancer patients remarkably declined compared with severe hepatitis patients.
Serum RCOR3 levels in patients with hepatitis B
As shown in Table 3, there was no significant difference in serum RCOR3 levels between the mild chronic hepatitis B patients and the healthy control group. However, compared with the healthy control group, the serum RCOR3 levels were significantly increased in patients with moderate chronic hepatitis B and significantly decreased in patients with SHB, cirrhosis and HCC. Moreover, the serum RCOR3 levels in SHB, cirrhosis and liver cancer patients remarkably declined compared with the moderate chronic hepatitis B patients and mild chronic hepatitis B patients. Taken together, serum RCOR3 levels were increased in patients with mild chronic hepatitis B and moderate chronic hepatitis B and were decreased in patients with SHB, cirrhosis and liver cancer.
Correlation of serum RCOR3 and hepatitis B-associated serum markers
As shown in Table 4, rank correlation analysis demonstrated that there was a significant correlation between serum RCOR3 and TB (r=-0.305, P<0.01), while there was no significant correlation between ROCR3 and ALT (r=0.014, P>0.05) or AST (r=-0.079, P>0.05).
Discussion
REST, also known as NRSF or X box repressor (XBR), is the first identified system-wide transcription repressor implicated in vertebrate neuronal development4, 24, 25, 26, 27. However, more and more studies have provided evidence that REST plays an essential role in non-neuronal tissues28 by functioning as a transcription silencer or activator29, 30, 31, 32, 33, 34. For example, REST was reported to be a critical regulator in medulloblastoma cells and may serve as a therapeutic target for medulloblastoma35. The profound context-specificity of the functional repertoire of REST is further highlighted by its dual role as a tumor suppressor and oncoprotein36, 37, 38.
The results of the GeneChip Arrays and real-time PCR indicated that the mRNA level of RCOR3 was initially reduced and then subsequently induced following ConA treatment. However, an RCOR3 ELISA on human serum showed that RCOR3 levels were initially induced and then reduced. It is possible that different samples and distinct RCOR3 detection methods (mouse liver tissue for GeneChip Arrays and real-time PCR; human plasma for ELISA) are responsible for the observed differences in RCOR3 levels.
RCOR3 is a member of the CoREST family and functions as a corepressor for REST, which mediates repression by REST. The significant increase of serum RCOR3 levels in mild chronic hepatitis B or moderate chronic hepatitis B patients in comparison with normal cases indicates the enhanced function of RCOR3 at the very early stage of hepatitis B, which may lead to the repression of certain genes and thus be helpful to inhibit the progression of the disease. However, the significant decline of serum RCOR3 levels in patients with SHB, cirrhosis and HCC with the development of disease indicates the impaired function of RCOR3, which may aggravate certain diseases.
To summarize, our data showed that serum RCOR3 levels may reflect the degree of liver damage, and the assessment of serum RCOR3 in patients with hepatitis B could be helpful to the classification and intervention of the disease. Compared with the most common serum markers of hepatitis, such as ALT, AST, or TB, RCOR3 might be a more ideal biomarker that reflects the process or outcome of HB because the variation in ALT, AST or TB levels is rather intensive among the HB patients. This study also provides new insights into the potential role of the NRSE-NRSF system in non-neuronal tissues at both the physiological and pathological conditions. Nevertheless, the role of RCOR3 (REST) in the development of hepatitis B remains obscure. More systematic and comprehensive investigations are still needed to deepen our understanding on the biological significance of RCOR3. In future research, we would like to explore the mechanisms of how RCOR3 (REST) participates in the development of hepatitis B. Because REST regulates different groups of target genes in different cell types39, we intend to identify the potential REST targets and test whether liver injury could be reversed by interfering with the expression of RCOR3.
Author contribution
Feng CHEN and Zhi CHEN designed the experiments; Ji-hua XUE, Min ZHENG, Xiao-wei XU, and Shan-shan WU performed the experiments; Ji-hua XUE and Shan-shan WU analyzed the data; and Ji-hua XUE and Feng CHEN wrote the paper.
References
Shepard CW, Simard EP, Finelli L, Fiore AE, Bell BP . Hepatitis B virus infection: epidemiology and vaccination. Epidemiol Rev 2006; 28: 112–25.
Da Villa G, Nuri B, Ghisetti V, Picciotto L, Ponzetto A, Angoni R, et al. Epidemiology of hepatitis B and delta virus infection in Albania: an approach to universal vaccination. Res Virol 1995; 146: 245–8.
Chen ZF, Paquette AJ, Anderson DJ . NRSF/REST is required in vivo for repression of multiple neuronal target genes during embryogenesis. Nat Genet 1998; 20: 136–42.
Schoenherr CJ, Anderson DJ . The neuron-restrictive silencer factor (NRSF): a coordinate repressor of multiple neuron-specific genes. Science 1995; 267: 1360–3.
Palm K, Belluardo N, Metsis M, Timmusk T . Neuronal expression of zinc finger transcription factor REST/NRSF/XBR gene. J Neurosci 1998; 18: 1280–96.
Ooi L, Wood IC . Chromatin crosstalk in development and disease: lessons from REST. Nat Rev Genet 2007; 8: 544–54.
Ballas N, Mandel G . The many faces of REST oversee epigenetic programming of neuronal genes. Curr Opin Neurobiol 2005; 15: 500–6.
Hsieh J, Gage FH . Epigenetic control of neural stem cell fate. Curr Opin Genet Dev 2004; 14: 461–9.
Roopra A, Sharling L, Wood IC, Briggs T, Bachfischer U, Paquette AJ, et al. Transcriptional repression by neuron-restrictive silencer factor is mediated via the Sin3-histone deacetylase complex. Mol Cell Biol 2000; 20: 2147–57.
You A, Tong JK, Grozinger CM, Schreiber SL . CoREST is an integral component of the CoREST- human histone deacetylase complex. Proc Natl Acad Sci U S A 2001; 98: 1454–8.
Andres ME, Burger C, Peral-Rubio MJ, Battaglioli E, Anderson ME, Grimes J, et al. CoREST: a functional corepressor required for regulation of neural-specific gene expression. Proc Natl Acad Sci U S A 1999; 96: 9873–8.
Grimes JA, Nielsen SJ, Battaglioli E, Miska EA, Speh JC, Berry DL, et al. The co-repressor mSin3A is a functional component of the REST-CoREST repressor complex. J Biol Chem 2000; 275: 9461–7.
Huang Y, Myers SJ, Dingledine R . Transcriptional repression by REST: recruitment of Sin3A and histone deacetylase to neuronal genes. Nat Neurosci 1999; 2: 867–72.
Naruse Y, Aoki T, Kojima T, Mori N . Neural restrictive silencer factor recruits mSin3 and histone deacetylase complex to repress neuron-specific target genes. Proc Natl Acad Sci U S A 1999; 96: 13691–6.
Roopra A, Qazi R, Schoenike B, Daley TJ, Morrison JF . Localized domains of G9a-mediated histone methylation are required for silencing of neuronal genes. Mol Cell 2004; 14: 727–38.
Chen F, Zhu HH, Zhou LF, Li J, Zhao LY, Wu SS, et al. Genes related to the very early stage of ConA-induced fulminant hepatitis: a gene-chip-based study in a mouse model. BMC Genomics 2010; 11: 240.
Cao QY, Chen F, Li J, Wu SS, Wang J, Chen Z . A microarray analysis of early activated pathways in concanavalin A-induced hepatitis. J Zhejiang Univ Sci B 2010; 11: 366–77.
Kaneko Y, Harada M, Kawano T, Yamashita M, Shibata Y, Gejyo F, et al. Augmentation of Valpha14 NKT cell-mediated cytotoxicity by interleukin 4 in an autocrine mechanism resulting in the development of concanavalin A-induced hepatitis. J Exp Med 2000; 191: 105–14.
Miyazawa Y, Tsutsui H, Mizuhara H, Fujiwara H, Kaneda K . Involvement of intrasinusoidal hemostasis in the development of concanavalin A-induced hepatic injury in mice. Hepatology 1998; 27: 497–506.
Tiegs G, Hentschel J, Wendel A . A T cell-dependent experimental liver injury in mice inducible by concanavalin A. J Clin Invest 1992; 90: 196–203.
Gantner F, Leist M, Lohse AW, Germann PG, Tiegs G . Concanavalin A-induced T-cell-mediated hepatic injury in mice: the role of tumor necrosis factor. Hepatology 1995; 21: 190–8.
Wright GW, Simon RM . A random variance model for detection of differential gene expression in small microarray experiments. Bioinformatics 2003; 19: 2448–55.
Chinese Society of Hepatology and Chinese Society of Infectious Diseases, Chinese Medical Association. The guideline of prevention and treatment for chronic hepatitis B. Zhonghua Liu Xing Bing Xue Za Zhi 2011; 32: 405–15.
Chong JA, Tapia-Ramirez J, Kim S, Toledo-Aral JJ, Zheng Y, Boutros MC, et al. REST: a mammalian silencer protein that restricts sodium channel gene expression to neurons. Cell 1995; 80: 949–57.
Mori N, Schoenherr C, Vandenbergh DJ, Anderson DJ . A common silencer element in the SCG10 and type II Na+ channel genes binds a factor present in nonneuronal cells but not in neuronal cells. Neuron 1992; 9: 45–54.
Scholl T, Stevens MB, Mahanta S, Strominger JL . A zinc finger protein that represses transcription of the human MHC class II gene, DPA. J Immunol 1996; 156: 1448–57.
Kraner SD, Chong JA, Tsay HJ, Mandel G . Silencing the type II sodium channel gene: a model for neural-specific gene regulation. Neuron 1992; 9: 37–44.
Kuwahara K, Saito Y, Ogawa E, Takahashi N, Nakagawa Y, Naruse Y, et al. The neuron-restrictive silencer element-neuron-restrictive silencer factor system regulates basal and endothelin 1-inducible atrial natriuretic peptide gene expression in ventricular myocytes. Mol Cell Biol 2001; 21: 2085–97.
Kuwabara T, Hsieh J, Nakashima K, Taira K, Gage FH . A small modulatory dsRNA specifies the fate of adult neural stem cells. Cell 2004; 116: 779–93.
Sun YM, Greenway DJ, Johnson R, Street M, Belyaev ND, Deuchars J, et al. Distinct profiles of REST interactions with its target genes at different stages of neuronal development. Mol Biol Cell 2005; 16: 5630–8.
Ballas N, Grunseich C, Lu DD, Speh JC, Mandel G . REST and its corepressors mediate plasticity of neuronal gene chromatin throughout neurogenesis. Cell 2005; 121: 645–57.
Bessis A, Champtiaux N, Chatelin L, Changeux JP . The neuron-restrictive silencer element: a dual enhancer/silencer crucial for patterned expression of a nicotinic receptor gene in the brain. Proc Natl Acad Sci U S A 1997; 94: 5906–11.
Yoo J, Jeong MJ, Lee SS, Lee KI, Kwon BM, Kim DS, et al. The neuron restrictive silencer factor can act as an activator for dynamin I gene promoter activity in neuronal cells. Biochem Biophys Res Commun 2001; 283: 928–32.
Su X, Kameoka S, Lentz S, Majumder S . Activation of REST/NRSF target genes in neural stem cells is sufficient to cause neuronal differentiation. Mol Cell Biol 2004; 24: 8018–25.
Lawinger P, Venugopal R, Guo ZS, Immaneni A, Sengupta D, Lu W, et al. The neuronal repressor REST/NRSF is an essential regulator in medulloblastoma cells. Nat Med 2000; 6: 826–31.
Guardavaccaro D, Frescas D, Dorrello NV, Peschiaroli A, Multani AS, Cardozo T, et al. Control of chromosome stability by the beta-TrCP-REST-Mad2 axis. Nature 2008; 452: 365–9.
Majumder S . REST in good times and bad: roles in tumor suppressor and oncogenic activities. Cell Cycle 2006; 5: 1929–35.
Singh SK, Kagalwala MN, Parker-Thornburg J, Adams H, Majumder S . REST maintains self-renewal and pluripotency of embryonic stem cells. Nature 2008; 453: 223–7.
Liu Z, Liu M, Niu G, Cheng Y, Fei J . Genome-wide identification of target genes repressed by the zinc finger transcription factor REST/NRSF in the HEK 293 cell line. Acta Biochim Biophys Sin (Shanghai) 2009; 41: 1008–17.
Acknowledgements
This work was supported by the National Basic Research Program 2007CB512905, National Natural Science Foundation 30571664, and the State S&T Projects (11th Five Year) 2008ZX10002-007, Zhejiang Province Science & Technology Program 2008C23045.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xue, Jh., Zheng, M., Xu, Xw. et al. Involvement of REST corepressor 3 in prognosis of human hepatitis B. Acta Pharmacol Sin 32, 1019–1024 (2011). https://doi.org/10.1038/aps.2011.49
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2011.49