Abstract
Aim:
To characterize the metabolism of cardamonin by the P450 enzymes in human and animal liver microsomes.
Methods:
Cardamonin was incubated with both human and animal liver microsomal incubation systems containing P450 reaction factors. High performance liquid chromatography coupled with ion trap mass spectrometry was used to identify the metabolites. Serial cardamonin dilutions were used to perform a kinetic study in human liver microsomes. Selective inhibitors of 7 of the major P450 isozymes were used to inhibit cardamonin hydroxylation to identify the isozymes involved in cardamonin metabolism. The cardamonin hydroxylation metabolic capacities of human and various other animals were investigated using the liver microsomal incubation system.
Results:
Two metabolites generated by the liver microsome system were detected and identified as hydroxylated cardamonin. The Km and Vmax values for cardamonin hydroxylation were calculated as 32 μmol/L and 35 pmol·min−1·mg−1, respectively. Furafylline and clomethiazole significantly inhibited cardamonin hydroxylation. Guinea pigs showed the highest similarity to humans with respect to the metabolism of cardamonin.
Conclusion:
CYP 1A2 and 2E1 were identified as the P450 isozymes involved in the metabolism of cardamonin in human liver microsomes. Furthermore, our research suggests that guinea pigs could be used in the advanced pharmacokinetic studies of cardamonin in vivo.
Similar content being viewed by others
Introduction
Cardamonin (Figure 1), a chalcone that is present at high content1 in commonly used traditional Chinese medicines2, such as Alpinia katsumadai3 and Amomum subulatum4, shows broad-spectrum preclinical pharmacological activities such as anti-platelet aggregation5, vasorelaxant effects6, anti-inflammation7 and inhibition of NO release and iNOS expression8. Because of these significant pharmacological activities, cardamonin shows some potential to be a drug candidate.
For the advanced development of drug candidates, not only pharmacological activities, but also pharmacokinetic properties should be considered. Metabolism studies are an important part of the whole pharmacokinetic study because a structural change after metabolism can modulate either the pharmacological activity or the toxicity of the drug candidates9, 10. Among all of the metabolic processes, metabolisms catalyzed by the cytochrome P450 (CYP) enzymes are the most important because almost 70%−80% of the known Phase I metabolisms are attributed to them11. In recent years, in vitro metabolism studies involving P450 enzymes have become more common and have been suggested to be an important part of preclinical drug development12, 13, 14 so that the potential failure of a drug induced by its metabolic properties can be avoided in the advanced clinical drug development studies.
However, until now, few studies have been carried out on the metabolism of chalcones, cardamonin in particular, although metabolism studies of other flavonoids, including flavones15, flavonols16, isoflavones17, 18 and flavonones19, have been performed extensively. In the present study, an in vitro metabolism study of cardamonin in human and animal liver microsomes was performed. The isozymes involved in the metabolism of cardamonin were identified and the differences among species with respect to cardamonin metabolism were noted. The results of the present study will help to understand the basic metabolic properties of cardamonin, which will further help to advance its pharmacokinetic study in vivo.
Materials and methods
Chemicals and Materials
Cardamonin was isolated from Alpiniae Katsumadai by our lab and the purity was determined to be more than 98% as tested by a high performance liquid chromatography (HPLC) system coupled with a photodiode array (PDA) detector. Glucose-6-phosphate (G-6-P), G-6-P dehydrogenase and β-nicotinamide-adenine dinucleotide phosphate (NADP) were purchased from Sigma-Aldrich (St Louis, MO, USA). Testosterone, which was used as a substrate for the positive control, was purchased from Acros (Geel, Belgium). Reagents for the chemical inhibition study, including sulfaphenazole, quinidine, clomethiazole, furafylline and 8-methoxypsoralen, were purchased from Sigma-Aldrich (St Louis, MO, USA) and ketoconazole and S-mephenytoin were obtained from ICN Biomedicals Inc (Aurora, OH, USA) and Toronto Research Chemicals Inc (North York, Canada), respectively. All other reagents were of HPLC grade or the highest commercially-available grade.
Preparation of liver microsomes of human and animals
Human livers were obtained from autopsy samples (n=5 Chinese males, 27 to 48 years old) from Dalian Medical University (Dalian, China), with the approval of the ethics committee of Dalian Medical University. The medication history of the donors was not known. Research involving human subjects was done under full compliance with government policies and the Helsinki Declaration. Cattle, sheep and pig livers were freshly obtained from animals in abattoirs. Mouse, rat, guinea pig and dog livers were freshly obtained from the experimental animal center of Shanghai University of Traditional Chinese Medicine (Shanghai, China). After being harvested, the liver specimens were immediately stored in liquid nitrogen until preparation of the microsomes.
Microsomes were prepared from liver tissue by differential ultracentrifugation as described previously20. Protein concentration was determined by using bovine serum albumin as standards21. Liver microsomes were diluted to 10 g/L and stored at -80 °C.
Parameters for chromatography and mass spectrometry
A Survyor HPLC system coupled with a LCQ ion trap mass spectrometry (Thermo Finnigan, San Jose, CA, USA) via an electronic spray ionization (ESI) ion source was used to perform the metabolic profile study and characterize the structure of the metabolites. The HPLC-MSn system consists of a quaternary pump, an auto-sampler and a column compartment. An Intersile C18 column (150×2.1 mm, 5 μm, Japan) was used for separation. Acetonitrile and formic acid solution (0.1% in water) were used as mobile phase A and B, respectively. The following HPLC eluting gradient was derived from our published method22: initiated from 10% of A: 0–5 min, 10%–35% of A; 5–20 min, 35% of A; 20–30 min, 35%–50% of A; 30–35 min, 50%–70% of A. The eluting flow rate was set to be 0.15 mL/min. The negative ion monitor mode was used and mass parameters on HPLC-MSn were set as follows: capillary temperature was 300 °C, sheath gas flow was 20 unit, aux/sweep gas flow was 5 unit, source voltage was 4.7 kV, and capillary voltage was -15 V. A full scan was used to trap the metabolites and parent compound to obtain their molecular weight values, and daughter scans of [M-H]− ions of cardamonin and its metabolites were used to obtain their second order mass (MS2) spectra.
An ultra performance liquid chromatography (UPLC) system (Waters Acquity) coupled with single quadrupole mass spectrometry (MS, Version: ZQ) via an ESI ion source was used to quantitatively determine the cardamonin metabolites in studies to identify the P450 isozymes involved in the metabolism, kinetics and species difference of cardamonin hydroxylation. The eluting gradient was similar to that used on the HPLC system, but the flow rate was changed to 0.3 mL/min and the time in the gradient on the UPLC system was shortened by 10 times compared with the HPLC system. A Waters BEH C18 (50×2.1 mm, 1.7 μm) column was used for separation in the UPLC system. Negative ion monitor mode was adopted and mass parameters for UPLC-MS were set as follows: capillary voltage was 3.2 kV, cone voltage was -40 V, extractor voltage was -1.59 V, source and desolvation temperature were set at 100 and 350 °C, as well cone and desolvation gas flow were set at 50 and 550 L/h. A selective ion monitor (SIM) mode was used to trap the metabolites at m/z 284 ([M-H]− ion) and the parent compound at m/z 269 ([M-H]− ion), respectively.
Incubation system for identification of metabolites in human liver microsomes
A standard incubation system included human liver microsomes (HLM, 0.5 g/L, 10 μL), G-6-P (1 mmol/L, 20 μL), G-6-P dehydrogenase (1 unit/mL, 20 μL), phosphate buffer (100 mmol/L, pH 7.4, 108 μL), MgCl2 (4 mmol/L, 20 μL), and cardamonin (100 μmol/L, 2 μL). Cardamonin was dissolved in methanol, but all other reagents were dissolved in phosphate buffer. The total volume of the incubation system was 200 μL and the total organic volume was less than 1% of the system. The reaction was initiated by adding NADP (1 mmol/L, 20 μL). After incubation for 60 min, 100 μL of acetonitrile was added to stop the reaction. The supernatant was transferred to perform analysis after centrifugation at 20 000×g and 4 °C for 10 min.
Analysis of metabolites
Four experiment groups were set up in the present study: a reaction group, which included cardamonin, HLM, NADP, and all other factors described above; a negative control group, which included cardamonin and all other reaction factors except NADP (which was replaced by phosphate buffer with the same volume); a blank group, which included all reaction factors without cardamonin but the same volume of methanol; and a positive control in which cardamonin was replaced by testosterone, which is a probe substrate of CYP3A4. Peaks appearing only in the total ion chromatogram (TIC) or extractive ion chromatogram (EIC) of the reaction group, but not in the negative control and blank groups, were considered the metabolites of cardamonin. MS2 spectra of the metabolites were compared with that of cardamonin to determine their possible structures. The nomenclature of the fragment ions, which have been referred to in several references23, 24, was modified slightly in the present study (Figure 1).
Kinetics study of metabolite generation in human liver microsomes
Preliminary experiments were carried out to ensure that the formation of the metabolites was linear with the reaction time and the concentration of the protein in human liver microsomes. All experiments were carried out in duplicate and the precision of the parallel samples was relatively good (with RSD values generally below 10%). Because the quantity of M2 was so small and could not be detected in 7 of the animal liver microsomes, the kinetic study was only conducted on the dominant metabolite M1. Concentrations of the cardamonin dilutions were set at 2.5, 4, 6.3, 10, 16, 25, 32, 40, 50, 63, 75, and 85 μmol/L. Reaction velocities and substrate concentrations were used to calculate the apparent Km and Vmax values according to nonlinear regression from the Michaelis-Menten equation (Equation 1). Nonlinear regression was performed using Origin software (OrginLab Corporation, Northampton, MA and Version 7.5)25. To determine whether the metabolism was monophasic or biphasic, the result was graphically represented on Eadie-Hofstee plots (velocities versus ratios of velocities to substrate concentrations). Intrinsic clearance (CLint), a part of the total body clearance representing the capacity of the metabolizing enzyme for clearing drugs, was calculated from Equation 226, 27,


where Vmax was the maximum reaction velocity, Km was the Michaelis constant that represented the substrate concentration at which the velocity was half of the Vmax, V was the reaction velocity, [S] was the substrate concentration, W was body weight, YL was liver weight per kilogram of body weight, YP was the microsome protein content in the human livers (mg/g), and W, YL, and YP were 60 kg, 4.5 g/kg and 45 mg/g for Eastern Asian people, respectively28.
Species differences of cardamonin hydroxylation
To compare the species differences for the metabolism of cardamonin and suggest a suitable animal species to perform its pharmacokinetics in vivo, cardamonin was incubated in the liver microsome incubation systems of humans and several other animals, including dog, rat, mouse, guinea pig, pig, cattle, and sheep. The concentration of cardamonin used in the incubation system was 30 μmol/L (near the Km value). The protein concentrations of the microsomes from humans and all other animal livers used in the incubation system were 0.5 g/L. The incubation time was 25 min. The quantities of M1 were used to compare the metabolic capacity of humans with that of other animals. Microsomes used to determine the species differences for each of the species were from 6 individuals, and average values of the 6 individuals were used for comparison. Group t-tests were performed between each of the animals and humans to examine the significance of the difference when the probabilities value (P) was less than 0.05.
Chemical inhibition of cardamonin hydroxylation in human liver microsomes
Selective inhibitors, including furafylline (10 μmol/L), 8-methoxypsoralen (2.5 μmol/L), quercetin (10 μmol/L), sulfaphenazole (10 μmol/L), quinidine (10 μmol/L), clomethiazole (50 μmol/L) and ketoconazole (1 μmol/L), were used to inhibit the 7 main P450s, including CYP 1A2, 2A6, 2C8, 2C9, 2D6, 2E1, and CYP3A4, respectively, in human liver microsomes29, 30, 31. All of the inhibitors added to the incubation system were dissolved in a 1 μL volume of methanol, which was less than 1% of the total incubation volume; other factors in the incubation system were the same as that in the kinetic study. An incubation without any inhibitor, but with the dissolving medium was set as a solvent control. The reaction system was pre-incubated for 3 min at 37 °C and the reaction was initiated by the addition of cardamonin. The quantities of M1 in the inhibited incubations were compared with the solvent control incubation, which was normalized to 100 %. The compared value was defined as the remaining enzyme activity (REA) and used as the parameter to evaluate the catalytic capacity of an isozyme. However, the chemical inhibition study was conducted only on the formation of M1 because the quantity of M2 was too small to determine quantitatively.
To understand the inhibitory patterns of cardamonin hydroxylation inhibited by furafylline and clomethiazole, the kinetics of cardamonin hydroxylation was analyzed at various concentrations of these two inhibitors. Concentrations of the cardamonin dilutions were used at 5, 10, 30, 50, and 80 μmol/L. Inhibitor concentrations were set to 0, 0.5, 2, and 10 μmol/L for furafylline and at 0, 1, 50, and 100 μmol/L for clomethiazole. Data were plotted on a Lineweaver-Burk plots (double-reciprocal plot of reaction velocity and substrate concentration) to understand the inhibitory patterns of cardamonin hydroxylation inhibited by furafylline and clomethiazole.
Results
MS2 fragmentation of cardamonin
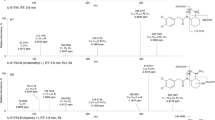
To elucidate the structures of the metabolites, the MS2 fragmentation of the parent compound was first obtained as the reference. The fragment ions and MS2 spectrum of cardamonin are shown in Table 1 and Figure 2A, respectively. In the MS2 spectrum of the [M-H]− ion at m/z 269, three representative ions at m/z 124, 139, and 165 were observed (Figure 2A). The ion at m/z 165 was generated by cleavage between C8 and C9. Serial neutral loss from the ion at m/z 165 produced another two fragment ions (Table 1) at m/z 139 (165-CO) and 124 (165-CO-CH3·), respectively. All the three ions could be valuable to identify the structure of ring A. The fragment ions that represent Part I (Table 1) of the cardamonin structure were also observed at m/z 177 and 179 (Figure 2A).
Identification of metabolites
Comparing the chromatograms of the reaction group with the negative control and the blank group samples, two NADPH-dependent metabolites marked as M1 and M2 were detected (Figure 3A). The presence of [M-H]− ions at m/z 285 suggesting that both the MW values of M1 and M2 were 16 Da higher than that of cardamonin, which indicated that these two metabolites were two hydroxylates. Fragment ions and MS2 spectra of M1 and M2 are shown in Table 1 and Figure 2B and 2C.
In the MS2 spectra of M1 (Figure 2B), the presence of the charateristic ions at m/z 124, 139 and 165, which were the same as those of cardamonin (Figure 2A), suggested that ring A of cardamonin was not metabolically modified (Table 1). The ion at m/z 119 in the MS2 spectra of M1 (Figure 2B) indicates that the hydroxylation site is on part II of the cardamonin structure including ring B, C7 or C8 (Table 1). The characteristic fragment ion at m/z 179 (Figure 2B), which is the same as that of cardamonin, suggests that the structure of M1 contain the part I (Table 1) of the cardamonin and that C7 and C8, which are contained in the structure of both part I and part II, are not substituted by the hydroxyl. Thus, the structure of M1 was tentatively assigned as (ring B)-hydroxylation cardamonin.
With respect to the identification of M2, the MS2 spectra of [M-H]− ion at m/z 285 showed two characteristic fragment ions of cardamonin at m/z 165 and 139 (Table 1), suggesting that ring A of M2 was also unchanged. The ion at m/z 119 indicates that the substituted site is on part II of the cardamonin structure. However, a different fragmentation pattern from M1 and cardamonin was obtained that the ions at m/z 177 and 179 (Figure 2C) in the the MS2 spectra of M2 disappeared, indicating that part I (Table 1) of M2 was substituted by a hydroxyl. According to the analysis described above, the substituted site should be C7 or C8, because the two carbons were both contained in the structures of part I and II (Table 1). Furthermore, the MS2 spectra of M2 showed the lose of H2O to yield the product ion at m/z 267. As a molecule of H2O may be lost more easily between the C6'-hydroxyl and the C7-hydroxyl than that of the C8-hydroxyl (Table 1), M2 is putatively characterized as 7-hydroxylation cardamonin.
UPLC-MS analysis of cardamonin and its metabolites
Analysis of all of the chemical inhibition studies, species differences and kinetic studies on cardamonin hydroxylation were performed based on the developed analytical method. Chromatograms are shown in Figure 4. Comparing the chromatograms of the reaction group with the negative control and blank groups, M1 and M2 were identified as two peaks eluted out at 2.20 and 2.62 min, respectively, in the channel of metabolites (Figure 4A); and cardamonin was identified as the peak eluted out at 4.03 min in the channel of the parent compound (Figure 4B). The total analytical time was shortened by ten times compared with HPLC analysis and each peak was still adequately separated.
Kinetic study in human liver microsomes
Cardamonin dilutions were used for incubation with HLM. A protein concentration of 0.5 g/L and a reaction time of 25 min were linear with the cardamonin hydroxylation according to preliminary experiments (data not shown). Reaction velocities showed significant concentration-dependent characteristics (Figure 5A). According to the Eadie-Hofstee plots (Figure 5B), cardamonin hydroxylation showed a monophasic feature, suggesting that cardamonin is metabolized by only one isozyme or two isozymes with the same Km value. The Eadie-Hofstee plots also indicated that cardamonin hydroxylation in human liver microsomes followed the typical Michaelis-Menten equation (Equation 2). Vmax and Km values were calculated as 35 pmol−1·min−1·mg−1 and 32 μmol/L via nonlinear regression from the Michaelis-Menten equation using Origin software. Intrinsic clearance (CLint), calculated from equation 1, was 13.37 mL/min for people of Eastern Asian descent.
Species differences of cardamonin hydroxylation
M1, the dominant metabolite in the human liver microsome incubation, was also generated by all of the animal liver microsome incubations. Neither M2 nor the other metabolites was observed in the animal incubations. Therefore, M1 was the only animal metabolite. Quantities of M1 in the animal incubations were compared with its counterpart from human incubation, which was normalized to 100%. As a result, the quantity of M1 generated by guinea pig was nearly the same as that in humans with no significant difference; conversely, the difference between sheep and humans was the largest. Mouse and rat, the two species that are normally used experimental animal species, generated the metabolite at quantities only around 50% of the human levels. The metabolic capacities of most animals, except the guinea pig, are significantly different from that of humans (Figure 6), with a P value less than 0.05 according to t-tests (group).
Chemical inhibition study of cardamonin hydroxylation
The selective inhibitors of CYP 1A2, 2A6, 2C8, 2C9, 2D6, 2E1, and 3A4 were used to inhibit cardamonin hydroxylation. REA on cardamonin hydroxylation in each of the inhibited incubations was 16.14%, 73.97%, 55.88%, 63.53%, 80.41%, 13.99%, and 84.33% for CYP 1A2, 2A6, 2C8, 2C9, 2D6, 2E1, and 3A4, respectively (Figure 7). Results indicated that CYP 1A2 and 2E1 might play the most important roles during cardamonin hydroxylation. The inhibitory kinetics of cardamonin hydroxylation by furafylline and clomethiazole was analyzed. For furafylline, the trend lines of the Lineweaver-Burk plots at various inhibitor concentrations intersected the Y axis at the same plot (Figure 8A), which indicates that furafylline shows a competitive inhibition on cardamonin hydroxylation. For clomethiazole, the trend lines of the Lineweaver-Burk plots intersected the X axis at the same plot (Figure 8B), indicating that clomethiazole shows noncompetitive inhibition on cardamonin hydroxylation.
Discussion
The elucidation of the structure of the metabolites was performed via LC-MS/MS technology. Evidence shows that there is a hydroxyl substituted on ring B of cardamonin for M1, but the exact site on the benzene ring cannot be confirmed by mass spectrometry data alone. The most probable substitution site is suggested to be site 4 of ring B because the para position is the most easily substituted site of the benzene ring, which could be proven by analyzing the metabolite structures of other flavonoids, such as chrysin and galangin16, 32.
Metabolism may alter the pharmacokinetic conduct, activity or even toxicity of drugs in vivo. To avoid failure in an advanced in vivo study stage, a suitable animal species with metabolic properties similar to those of humans should be chosen first. In vitro metabolism studies could also be used to help rapidly and conveniently screen animal species. In the present study, cardamonin was incubated with liver microsomes from human and 7 animals. According to the results, the most commonly used experimental animal species, including mouse and rat, were significantly different from humans with regard to cardamonin hydroxylation, but guinea pigs showed the highest similarity to humans.
A chemical inhibition method was used to identify the P450 isozymes involved in cardamonin hydroxylation. As a result, two selective inhibitors named furafylline and clomethiazole both showed a significant inhibitory effect on cardamonin hydroxylation. To further understand the inhibitory mechanism of these two inhibitors on cardamonin hydroxylation, an inhibitory kinetics was analyzed. Data showed that furafylline showed a competitive inhibition on cardamonin hydroxylation, which corresponded with the inhibition pattern of furafylline on CYP 1A2. Furafylline is a mechanism-based inhibitor of CYP 1A233. In this kind of inhibition, the actual inhibitor was not intact furafylline, but its metabolite catalyzed by CYP 1A234, suggesting that furafylline is also one of the substrates of this isozyme. Thus, furafylline could compete for the active site of CYP 1A2 with cardamonin, and then it showed competitive inhibition of cardamonin hydroxylation. Clomethiazole, the selective inhibitor of CYP 2E1, showed a noncompetitive inhibition on cardamonin hydroxylation, corresponding with the published paper in which clomethiazole was demonstrated to be a noncompetitive inhibitor of CYP 2E135. The inhibitory patterns of furafylline and clomethiazole on cardamonin hydroxylation additionally proved that CYP 1A2 and 2E1 are involved in the metabolism of cardamonin.
CYP 1A2 was identified as one of the main isozymes involved in cardamonin hydroxylation, which corresponds well with reports of other flavonoids, such as hesperitin, kaempferol, naringenin and apigenin16, 19, 36. CYP 2E1 also catalyzed cardamonin hydroxylation in the present study, although this isozyme has only been shown to catalyze the hydroxylations of a few other flavonoids. However, except CYP 1A2 and 2E1, the other isozymes also acted as minor metabolizing enzymes for cardamonin hydroxylation, including the two CYP 2C isozymes. Cardamonin hydroxylation was also slightly inhibited by the selective inhibitors of CYP 2A6, 2D6, and 3A4. It was concluded from these result that hydroxylation of cardamonin is not a specific reaction, but is catalyzed by multiple P450 isozymes. However, such a suggestion should be confirmed by other studies, such as recombinant P450s metabolism and correlation analysis.
In the present study, two hydroxylates of cardamonin were observed in the human liver microsome incubation system. CYP 1A2 and 2E1 were the two major isozymes involved in the metabolism of cardamonin. Although chemical inhibitors could also inhibit CYPs not mentioned in this study, the results are confined to the inhibition of the seven main CYPs investigated in the present study. The species difference studies on the metabolism of cardamonin showed that the guinea pig possesses the metabolic capacity that is the most similar to that of humans and could be used for advanced pharmacokinetic studies in vivo.
Author contribution
Yu-qi HE, Li YANG, Yong LIU, Chang-hong WANG, Ling YANG, and Zheng-tao WANG designed the research. Yu-qi HE performed the main experiments and wrote the paper. Jiang-wei ZHANG and Jun TANG revised the paper. Yuan-yuan LI isolated cardamonin. Juan SU and Yan-liu LU participated in part of the experiments.
References
Wang S, Zhou L, He W, Hu Z . Separation and determination of alpinetin and cardamonin by reverse micelle electrokinetic capillary chromatography. J Pharm Biomed Anal 2007; 43: 1557–61.
Chinese Pharmacopoeia Commission (China). Pharmacopoeia of the People's Republic of China. Beijing: People's Medical Publishing House; 2005.
Kimura Y, Takahashi S, Yoshida I . Studies on the constituents of Alpinia: XII: on the constituents of the seeds of Alpinia katsumadai hayata: I: the structure of cardamomin. Yakugaku Zasshi 1968; 88: 239–41.
Rao CB, Rao N, Suryaprakasam S . Cardamonin and alpinetin from the seeds of Amomum subulatum. Planta Med 1976; 29: 391–2.
Dong H, Chen SX, Xu HX . A new antiplatelet diarylheptanoid from Alpinia blepharocalyx. J Nat Prod 1998; 61: 142–4.
Wang ZT, Lau CW, Chan FL, Yao X, Chen ZY, He ZD, et al. Vasorelaxant effects of cardamonin and alpinetin from Alpinia henryi K. Schum. J Cardiovasc Pharmacol 2001; 37: 596–606.
Ahmad S, Israfa DA, Lajis NH, Shaari K, Mohamed H, Wahab AA, et al. Cardamonin, inhibits pro-inflammatory mediators in activated RAW 264.7 cells and whole blood. Eur J Pharmacol 2006; 538: 188–94.
Hatziieremia S, Gray AI, Ferro VA, Paul A, Plevin R . The effects of cardamonin on lipopolysaccharide-induced inflammatory protein production and MAP kinase and NFkappaB signalling pathways in monocytes/macrophages. Br J Pharmacol 2006; 149: 188–98.
Walsky RL, Boldt SE . In vitro cytochrome P450 inhibition and induction. Curr Drug Metab 2008; 9: 928–39.
Froy O . Cytochrome P450 and the biological clock in mammals. Curr Drug Metab 2009; 10: 104–15.
Nebert DW, Dalton TP . The role of cytochrome P450 enzymes in endogenous signalling pathways and environmental carcinogenesis. Nature 2006; 6: 947–60.
Bachmann KA, Ghosh R . The use of in vitro methods to predict in vivo pharmacokinetics and drug interactions. Curr Drug Metab 2001; 2: 299–314.
Howgate EM, Rowland Yeo K, Proctor NJ, Tucker GT, Rostami-Hodjegan A. Prediction of in vivo drug clearance from in vitro data. I: impact of inter-individual variability. Xenobiotica 2006; 36: 473–97.
Obach RS, Walsky RL, Venkatakrishnan K, Gaman EA, Houston JB, Tremaine LM . The utility of in vitro cytochrome P450 inhibition data in the prediction of drug-drug interactions. J Pharmacol Exp Ther 2006; 316: 336–48.
Gradolatto A, Canivenc-Lavier MC, Basly JP, Siess MH, Teyssier C . Metabolism of apigenin by rat liver phase I and phase II enzymes and by isolated perfused rat liver. Drug Metab Dispos 2004; 32: 58–65.
Silva ID, Rodrigues AS, Gaspar J, Laires A, Rueff J . Metabolism of galangin by rat cytochromes P450: relevance to the genotoxicity of galangin. Mutat Res 1997; 393: 247–57.
Tolleson WH, Doerge DR, Churchwell MI, Marques MM, Roberts DW . Metabolism of biochanin A and formononetin by human liver microsomes in vitro. J Agric Food Chem 2002; 50: 4783–90.
Kulling SE, Honig DM, Metzler M . Oxidative metabolism of the soy isoflavones daidzein and genistein in humans in vitro and in vivo. J Agric Food Chem 2001; 49: 3024–33.
Breinholt VM, Offord EA, Brouwer C, Nielsen SE, Brosen K, Friedberg T . In vitro investigation of cytochrome P450-mediated metabolism of dietary flavonoids. Food Chem Toxicol 2002; 40: 609–16.
Sanderink G, Bournique B, Stevens J, Petry M, Martinet M . Involvement of human CYP1A isoenzymes in the metabolism and drug interactions of riluzole in vitro. J Pharmacol Exp Ther 1997; 282: 1465–72.
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ . Protein measurement with the Folin phenol reagent. J Biol Chem 1951; 193: 265–75.
Wang Y, Yang L, He YQ, Wang CH, Welbeck EW, Bligh SWA, et al. Characterization of fifty-one flavonoids in a Chinese herbal prescription Longdan Xiegan Decoction by high-performance liquid chromatography coupled to electrospray ionization tandem mass spectrometry and photodiode array detection. Rapid Commun Mass Spectrom 2008; 22: 1767–78.
Wang DL, Liang Y, Chen WD, Xie L, Wang GJ, Liu XD . Identification of ginkgolide B metabolites in urine and rat liver cytochrome P450 enzymes responsible for their formation in vitro. Acta Pharmacol Sin 2008; 29: 376–84.
Fabre N, Rustan I, de Hoffmann E, Quetin-Leclercq J . Determination of flavone, flavonol, and flavanone aglycones by negative ion liquid chromatography electrospray ion trap mass spectrometry. J Am Soc Mass Spectrom 2001; 12: 707–15.
Liu HX, Liu Y, Zhang JW, Li W, Liu HT, Yang L . UDP-glucuronosyltransferase 1A6 is the major isozyme responsible for protocatechuic aldehyde glucuronidation in human liver microsomes. Drug Metab Dispos 2008; 36: 1562–9.
Liu L, Pang KS . An integrated approach to model hepatic drug clearance. Eur J Pharm Sci 2006; 29: 215–30.
Obach RS, Baxter JG, Liston TE, Silber BM, Jones BC, Macintyre F, et al. The prediction of human pharmacokinetic parameters from preclinical and in vitro metabolism data. J Pharmacol Exp Ther 1997; 283: 46–58.
Houston JB . Utility of in vitro drug metabolism data in predicting in vivo metabolic clearance. Biochem Pharmacol 1994; 47: 1469–79.
Bjornsson Td, Callaghan Jt, Einolf Hj, Fischer V, Gan L, Grimm S, et al. The conduct of in vitro and in vivo drug-drug interaction studies: a phrma perspective. Drug Metab Dispos 2003; 31: 815–32.
Huang SM, Temple R, Throckmorton DC, Lesko LJ . Drug interaction studies: study design, data analysis, and implications for dosing and labeling. Clin Pharmacol Ther 2007; 81: 298–304.
Harris JW, Rahman A, Kim BR, Guengerich FP, Collins JM . Metabolism of taxol by human hepatic microsomes and liver slices: participation of cytochrome P450 3A4 and an unknown P450 enzyme. Cancer Res 1994; 54: 4026–35.
Griffiths LA, Smith GE . Metabolism of apigenin and related compounds in the rat. Metabolite formation in vivo and by the intestinal microflora in vitro. Biochem J 1972; 128: 901–11.
Walsky RL, Obach RS . Validated assays for human cytochrome p450 activities. Drug Metab Dispos 2004; 32: 647–60.
Racha JK, Rettie AE, Kunze KL . Mechanism-based inactivation of human cytochrome P450 1A2 by furafylline: detection of a 1:1 adduct to protein and evidence for the formation of a novel imidazomethide intermediate. Biochemistry 1998; 37: 7407–19.
Gebhardt AC, Lucas D, Menez JF, Seitz HK . Chlormethiazole inhibition of cytochrome P450 2E1 as assessed by chlorzoxazone hydroxylation in humans. Hepatology 1997; 26: 957–61.
Doostdar H, Burke MD, Mayer RT . Bioflavonoids: selective substrates and inhibitors for cytochrome P450 CYP1A and CYP1B1. Toxicology 2000; 144: 31–8.
Acknowledgements
This work was partially supported by grants from the National Natural Science Foundation of China (No 30530840, 30772608), Key Project of Chinese National Programs for Fundamental Research and Development (973 Program, No 2006CB504704) and Shanghai Science & Technology Development Foundation (No 04DZ19851), the Greeting Fund of Dalian Institute of Chemical Physics, Chinese Academy of Science (No K2006A23).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
He, Yq., Yang, L., Liu, Y. et al. Characterization of cardamonin metabolism by P450 in different species via HPLC-ESI-ion trap and UPLC-ESI-quadrupole mass spectrometry. Acta Pharmacol Sin 30, 1462–1470 (2009). https://doi.org/10.1038/aps.2009.127
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2009.127
Keywords
This article is cited by
-
Pharmacokinetic interactions of clopidogrel with quercetin, telmisartan, and cyclosporine A in rats and dogs
Archives of Pharmacal Research (2012)