Abstract
Aim:
To investigate the role of the Notch1 signaling pathway in growth arrest of an esophageal carcinoma cell line (EC109) in vitro and the mechanism involved.
Methods:
An intracellular domain of Notch1 (ICN) was transfected into cultured EC109 cells by lipofectamine transfection. Subsequently, the proliferation of the transfected cells was measured by an MTT assay. Cell cycle distribution was analyzed by flow cytometry. Human papillomavirus type 18 (HPV18) E6/E7 mRNA expression was detected by RT-PCR, and p53 protein expression was detected by Western blot.
Results:
Activation of Notch1 signaling resulted in inhibition of EC109 cell proliferation with the induction of G2/M arrest, downmodulation of HPV18 E6/E7 gene expression, and upregulation of p53 expression.
Conclusion:
Repression of HPV18 E6/E7 expression by Notch1 signaling results in the activation of p53-mediated pathways with concomitant growth suppression of HPV18-positive EC109 cells.
Similar content being viewed by others
Introduction
Since their initial discovery in Drosophila as critical regulators of embryonic development, Notch genes have been found to be conserved in many species, including human. Notch genes encode highly conserved type I transmembrane glycoproteins, which can be activated via direct interaction with transmembrane ligands expressed on the surface of neighboring cells1, 2. Upon activation, Notch is cleaved, releasing an intracellular domain (ICN)3 that then translocates into the nucleus. The ICN associates with transcriptional factors known as Su(H)/CBF1 to regulate the expression of target genes and successively modulate the development and growth of cells3, 4. Constitutive expression of active ICN in targeted cells also results in an “activated” Notch phenotype5, 6.
Notch signaling is involved in a variety of cell specification, proliferation, and apoptosis processes that affect the development and function of many organs7, 8. For example, in the hematopoietic system, Notch is involved in T cell commitment and B cell development9. Overexpression of Notch1 has been demonstrated to promote the self-renewal of hematopoietic stem cells in vivo and in vitro 10, 11. Pathophysiologic alterations in Notch signaling have been associated with tumorigenesis. In human acute T-lymphoblastic leukemia and lymphomas, Notch1 transcripts encode a series of truncated Notch1 polypeptides containing at least the cytosolic domain of Notch11. Similar truncations caused by the insertion of wild-type mouse mammary tumor virus within the Notch1 and Notch4 genes are associated with mammary tumors3, 12. These observations suggest that dysfunction of intracellular Notch prevents differentiation and predisposes undifferentiated cells to malignant transformation13. On the other hand, constitutive activation of Notch1 signaling can cause a profound growth arrest in small cell lung cancer cells, associated with a G1 cell cycle arrest14. Overexpression of active Notch1 inhibits the proliferation of various prostate cancer cells15, suggesting that Notch activation can also induce growth arrest and reduce the neoplastic potential of tumors.
Esophageal cancer is the sixth most common cancer worldwide and the third most common malignancy of the gastrointestinal tract, and has a very poor prognosis. It is estimated that about 30 000 new cases of esophageal cancer are diagnosed annually in the world, approximately half of which occur in China. Esophageal cancer has two major histological types: adenocarcinoma (AC) and squamous cell carcinoma (ESCC); the latter is the type seen more frequently in China. Lu et al16 demonstrated that Notchl protein was overexpressed in normal esophageal tissues and underexpressed in human ESCC. Hence, there may be a link between the specific downmodulation of Notch1 expression in ESCC and their deregulated growth.
EC109, a well-differentiated human ESCC cell line, was established in 1976 by the National Laboratory of Molecular Oncology, Cancer Institute and Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College. It was reported that EC109 was a human papillomavirus type 18 (HPV18)-positive cell line17. In particular, the E6 and E7 oncoproteins of HPV18 perturb normal cell cycle control through their interactions with a number of cellular proteins, such as p53 and p105-Rb. The role of Notch1 is well studied in another HPV-positive human cervical cancer cell line, HeLa. It has been shown that expression of activated Notch1 strongly inhibits the growth of HPV-positive HeLa cells by downmodulation of the E6 and E7 genes18.
This study aims to assess whether increased Notch1 signaling via transfection with an exogenous intracellular domain of Notch1 (ICN) can directly affect the growth of EC109 cells and expressions of E6/E7 and p53.
Materials and methods
Reagents and plasmid
Thiazoyl blue tetrazolium bromide (MTT) was purchased from Amresco (Solon, OH, USA). Lipofectamine 2000 and anti-His antibody were purchased from Invitrogen (Carlsbad, CA, USA). Anti-p53 (FL-393) antibody, anti-β-actin antibody, biotinylated goat anti-mouse secondary antibody, and FITC-labeled goat anti-mouse secondary antibody were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). The FACSort flow cytometer was from Becton Dickinson (San Jose, CA, USA).
Plasmid pcDNA3.1C-ICN (denoted as ICN) was a gift from Professor Tom Kadesch (Department of Genetics, University of Pennsylvania School of Medicine, USA). ICN was generated by PCR and cloned into the vector pcDNA3.1C at the Xho I or Kpn I site, which contains a consensus kozak start site, and cloned in-frame with the myc-his tags. The gene sequence contained in this plasmid was confirmed by gene sequencing analysis.
Cell culture
The human ESCC cell line, EC109, was purchased from the Shanghai Institute of Cell Biology, the Chinese Academy of Sciences. EC109 cells were cultured in Iscove's modified Dulbecco's medium (IMDM) supplemented with 2 mol/L glutamine, 10% fetal calf serum, 100 U/mL penicillin G, and 100 U/mL streptomycin at 37 °C in a 5% CO2 atmosphere.
Transfection of ICN
The EC109 cells were inoculated into 24- or 96-well plates, and ICN or empty plasmid transfection was performed as soon as the cells reached above 80% in confluence. The transfection of ICN was performed using lipofectamine 2000 according to the manufacturer's instructions. Three experimental groups were established: a non-transfected group, an empty plasmid-transfected group, and an ICN-transfected group. All experiments were performed in triplicate.
Detection of ICN protein expression in EC109 cells by flow cytometry
The EC109 cells were harvested 72 h after transfection, washed twice in PBS, and fixed with 2% paraformaldehyde containing 0.1% Triton X-100 at 4 °C for 30 min. Next, normal goat serum was added and incubated at 4 °C for 30 min, after which the supernatant was discarded. The diluted anti-His antibody (1:100) was added and incubated at 4 °C for 30 min. Subsequently, the EC109 cells were washed twice in PBS. Next, the FITC-labeled secondary antibody (1:50) was added and incubated at 4 °C for 30 min, followed by two washes in PBS and detection by flow cytometry.
Proliferation rate of EC109 cells measured by the MTT assay
EC109 cells in 180 μL IMDM were inoculated onto 96-well plates. Four parallel rows of wells were set up for each group. The MTT assay was performed 24, 48, and 72 h after transfection, respectively. The absorbance (OD) of each well was measured at 570 nm with enzyme-linked immunosorbent detector (DG3022A).
Cell cycle analysis of EC109 cells by flow cytometry
EC109 cells were collected three days after transfection. Cell cycle analysis was performed with a flow cytometer, and the data were analyzed with Multicycle software (Phoenix Flow Systems Inc, San Diego, CA, USA).
Detection of HPV18 E6/E7 mRNA expression in EC109 cells by RT-PCR
EC109 cells were collected three days after transfection and HPV18 E6/E7 mRNA expressions were detected by RT-PCR. Total RNA was isolated from nucleated cells with the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's protocol and the RNA was dissolved in RNase-free water. RNA quality and quantity were assessed by ethidium bromideagarose gel electrophoresis and by relative absorbance at 260 nm versus 280 nm. Complementary DNA (cDNA) was synthesized in a volume of 20 μL with a cDNA synthesis kit (Fermenters, Glen Burnie, MD, USA) according to the manufacturer's protocol. The obtained cDNA was amplified by a regular PCR. The sequences of primers used were: TGTCAAAAACCGTTGTGTCC (sense) and GAGCTGTCGCTTAATTGCTC (anti-sense) for HPV18 E6/E7; ACCACAGTCCATGCCATCAC (sense) and TCCACCACCCTGTTGCTGTA (anti-sense) for GAPDH. The PCR conditions were: 94 °C for 3 min; 30 cycles of 94 °C for 30 s, 58 °C for 60 s, and 72 °C for 90 s, followed by a final extension at 72 °C for 10 min. The amplified PCR products were electrophoresed on agarose gels and the fragments were analyzed using the MUVB-20 gel analysis system (Ultralum Inc, Claremont, CA, USA). The absorbance (A) value of the gene band/GAPDH band was taken as the relative amount of the target gene. The sizes of the E6/E7 and GAPDH amplification products were 270 bp and 451 bp, respectively.
Detection of p53 protein expression in EC109 cells by Western blot analysis
The proteins from cell lysates of EC109 cells were separated by SDS-polyacrylamide gel electrophoresis and transferred onto PVDF membranes (Millipore, Bedford, MA, USA). The blots were incubated with the anti-p53 antibody or the anti-β-actin antibody (1:100 dilution in a blocking solution) at 4 °C overnight, then washed with a solution containing 5% milk, 20 mmol/L Tris-HCl (pH 7.5), 500 mmol/L NaCl, and 0.05% Tween-20 (TBS-T). After incubation with a secondary antibody (1:500 dilution in a blocking solution, 3 h at room temperature), positive signal bands were detected by the addition of nitroblue tetrazolium (NBT)-5-bromo-4-chloro-3-indolylphosphate (BCIP, Sigma, Saint Louis, MO, USA). The blots were scanned using Quantity One 4.4.1 (Bio-Rad Technical Service Department, Hercules, CA, USA).
Statistical analysis
All experiments were conducted in triplicate. The results are presented as mean±SD. All data were processed with the SPSS10.0 program (SPSS Inc.Chicago, Illinois, USA), and statistical differences were determined by ANOVA followed by the q test. P <0.05 was considered statistically significant.
Results
ICN expression rate in EC109 cells at 72 h post-transfection
The EC109 cells were transfected with ICN-containing plasmids. After 72 h, the ICN expression rate was 64.71%±5.97% as detected by flow cytometry (Figure 1).
Constitutive activation of Notch1 signaling inhibits proliferation of EC109 cells
MTT assays showed that the cell proliferation of the ICN-transfected group was significantly inhibited in comparison with the control groups (P<0.05; Figure 2).
Constitutive activation of Notch1 signaling in EC109 cells induces a G2/M arrest
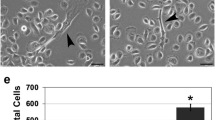
EC109 cells were collected three days after transfection. Cell cycle analysis was performed with a flow cytometer. In the non-transfected group, the percentages of cells in G0/G1 phase, G2/M phase, and S phase were 49.05%±1.57%, 1.88%±0.66%, and 49.07%±1.34%, respectively. In the empty plasmid-transfected group, the percentages of cells in G0/G1 phase, G2/M phase, and S phase were 49.93%±1.99%, 1.99%±1.02%, and 48.08%±0.99%, respectively. In the ICN-transfected group, the percentages of cells in G0/G1 phase, G2/M phase, and S phase were 52.16%±2.82%, 42.57%±1.57%, and 5.07%±1.59%, respectively. There was a significant difference in the percentage of G2/M or S phase cells between the ICN-transfected group and the non-transfected or empty plasmid-transfected groups (P<0.01), but not between the non-transfected group and the empty plasmid-transfected group. There was a decrease in S phase cells and an increase in G2/M phase cells, suggesting that constitutive activation of Notch signaling inhibits the proliferation of EC109 cells by blocking the cells at the G2/M phase of the cell cycle (Figure 3).
Constitutive activation of Notch1 signaling downmodulates HPV18 E6/E7 gene expression
As the growth of HPV-transformed cancer cells is dependent on sustained expressions of E6 and E718, we tested whether the specific growth inhibitory effects of activated Notch1 on EC109 cells could be explained by downmodulation of the E6 and E7 genes. RT-PCR analysis of EC109 cells at 72 h after transient transfection revealed a drastic inhibition of E6/E7 mRNA expression as a consequence of activated Notch1 expression (Figure 4).
Constitutive activation of Notch1 signaling upregulates p53 expression
To understand the molecular basis for Notch1-induced cell cycle arrest in EC109 cells, we assayed the protein expression of p53 at 72 h after transient transfection. Western blot analysis showed that p53 expression in the ICN-transfected group (2.15±0.23) was obviously higher than that in the non-transfected and empty plasmid-transfected groups (0.46±0.02) (P<0.01; Figure 5).
Discussion
Members of the Notch family of transmembrane receptors play an important role in cell fate determination. Over the past decade, a role for Notch in the pathogenesis of hematologic and solid malignancies has become apparent. Numerous cellular functions and microenvironmental cues associated with tumorigenesis are modulated by Notch signaling, including proliferation, apoptosis, adhesion, epithelial-to-mesenchymal transition, and angiogenesis. It is becoming increasingly evident that Notch1 signaling can be both oncogenic and tumor suppressive. To study the role of the Notch signaling pathway in EC109 cells, we transferred an activated form of Notch1 (ICN) into cultured EC109 cells, and found that activation of the Notch1 signaling pathway led to an arrest of EC109 cell proliferation in the G2/M phase of the cell cycle. A previous study had shown that constitutive overexpression of Notch1 signaling caused a G1 arrest in lung cells14. This difference may be due to the different experimental models utilized since Notch signaling functions in a cell- and context-specific manner19.
We also investigated the probable mechanism of Notch1-induced inhibition of EC109 cells, and found that activation of the Notch1 signaling pathway led to increased p53 protein expression and decreased E6/E7 gene expression. It has been reported that EC109 is a human papillomavirus type 18 (HPV18)-positive cell line17. Human papillomaviruses (HPVs), especially the high-risk types 16 and 18, have been identified as causative agents of at least 90% of cervical cancer cases and are linked to more than 50% of other anogenital cancers20. The HPV genome consists of approximately 8000 bp of closed-circular double-stranded DNA containing up to nine genes, and is functionally divided into three regions: a long control region (LCR) covering about 10% of the genome, an early (E) region, and a late (L) region21. The regulation of viral gene expression is complex and controlled by multiple cellular and viral transcription factors. Most of the regulation occurs within the LCR, which varies substantially in nucleotide composition between individual HPV types. Within the LCR, cis-active elements regulate transcription of the E6/E7 genes, which are the transforming genes responsible for immortalization and maintenance of the malignant phenotype in HPV-positive cancer cells. Transcription of the E6 and E7 genes is driven by the viral upstream regulatory region (URR) promoter, which is maintained intact and active in HPV-transformed cancer cells20. The AP-1 complex plays a key role in initiating and maintaining transcription from the URR promoter22. AP-1 is a heterodimeric DNA-binding complex formed by proteins of the c-Jun and c-Fos families. It was reported that increased Notch1 signaling caused a dramatic downmodulation of HPV-driven transcription of the E6/E7 viral genes, through suppression of AP-1 activity by upregulation of the Fra-1 family member and decreased c-Fos expression19. In HPV-positive cells, the p53 level is regulated by the continuous expression of E6. The E6 oncoprotein has been shown to recruit the cellular ubiquitin-protein ligase E6-AP to target the p53 tumor suppressor protein for ubiquitin-proteasome-mediated degradation23. p53 can arrest cells at the G2 checkpoint24, 25. We conclude that the proliferation arrest of EC109 cells caused by activation of the Notch1 signaling pathway may be associated with increased p53 expression following lowered E6/E7 expression.
In summary, our results demonstrate that Notch1-mediated repression of HPV18 E6/E7 expression results in activation of p53 pathways and concomitant growth suppression of HPV18-positive EC109 cells in vitro, which leads to growth arrest in the G2/M phase. Combined with results from previous studies26, our observations raise the possibility that downmodulation of Notch1 expression may be one of the mechanisms of esophageal squamous cell carcinoma tumorigenesis. Therefore, the Notch1 gene could be a new target for the treatment of squamous cell carcinoma.
Author contribution
Wen-li LIU, Ke-jie ZHANG designed research; Ke-jie ZHANG, Quan-yi LU, Xiao-qing NIU, Peng ZHANG , Jia-sheng HU, Pu LI performed research; Wen-li LIU contributed new analytical tools and reagents; Jiang-ning ZHAO, Zhao WANG, analyzed data; Ke-jie ZHANG wrote the paper.
References
Aster JC, Xu L, Karnell FG, Patriub V, Pui JC, Pear JC, et al. Essential roles for ankyrin repeat and transactivation domains in induction of T-cell leukemia by notch1. Mol Cell Biol 2000; 20: 7505–15.
Bresnick EH, Chu J, Christensen HM, Lin B, Norton J . Linking Notch signaling, chromatin remodeling, and T-cell leukemogenesis. J Cell Biochem 2000; 35 (Suppl): 46–53.
Yanagawa S, Lee JS, Kakimi K, Matsuda Y, Honjo T, Ishimoto A . Identification of Notch1 as a frequent target for provirus insertional mutagenesis in T-cell lymphomas induced by leukemogenic mutants of mouse mammary tumor virus. J Virol 2000; 74: 9786–91.
Osborne B, Miele L . Notch and the immune system. Immunity 1999; 11: 653–63.
Bigas A, Martin DI, Milner LA . Notch1 and Notch2 inhibit myeloid differentiation in response to different cytokines. Mol Cell Biol 1998; 18: 2324–33.
Milner LA, Bigas A, Kopan R, Brashem-Stein C, Bernstein ID, Martin DI . Inhibition of granulocytic differentiation by mNotch1. Proc Natl Acad Sci USA 1996; 93: 13014–9.
Ohishi K, Katayama N, Shiku H, Varnum-Finney B, Bernstein ID . Notch signalling in hematopoiesis. Semin Cell Dev Biol 2003; 14: 143–50.
Weijzen S, Rizzo P, Braid M, Vaishnav R, Jonkheer SM, Zlobin A, et al. Activation of Notch-1 signaling maintains the neoplastic phenotype in human Ras-transformed cells. Nat Med 2002; 8: 979–86.
Maillard I, He Y, Warren S . Pear from the yolk sac to the spleen: new roles for notch in regulating hematopoiesis. Immunity 2003; 18: 587–9.
Stier S, Cheng T, Dombkowski D, Carlesso N, Scadden DT . Notch1 activation increases hematopoietic stem cell self-renewal in vivo and favors lymphoid over myeloid lineage outcome. Blood 2002; 99: 2369–78.
Varnum-Finney B, Xu L, Brashem-Stein C, Nourigat C, Flowers D, Bakkour S, et al. Pluripotent, cytokine-dependent, hematopoietic stem cells are immortalized by constitutive Notch1 signaling. Nat Med 2000; 6: 1278–81.
Joutel A, Tournier-Lasserve E . Notch signalling pathway and human diseases. Semin Cell Dev Biol 1998; 9: 619–25.
Pear WS, Aster JC, Scott ML, Hasserjian RP, Soffer B, Sklar J, et al. Exclusive development of T cell neoplasms in mice transplanted with bone marrow expressing activated Notch alleles. J Exp Med 1996; 183: 2283–91.
Sriuranpong V, Borges MW, Ravi RK, Arnold DR, Nelkin BD, Baylin SB, et al. Notch signaling induces cell cycle arrest in small cell lung cancer cells. Cancer Res 2001; 61: 3200–5.
Shou J, Ross S, Koeppen H, de Sauvage FJ, Gao WQ . Dynamics of notch expression during murine prostate development and tumorigenesis. Cancer Res 2001; 61: 7291–7.
Lu ZM, Liu HT, Gao Y, Xu PR, Yan WH, Xue LX . Notch1 mRNA and protein expression in human esophageal squamous cell carcinoma and normal esophageal tissues. J Huazhong Normal Univ (Natural Science) 2007; 41: 263–7.
Qi ZL, Huo X, Zhang B, Yang HW, Qiu B, Peng L, et al. Esophageal carcinoma cell line-EC109 is confirmed a human papillomavirus type 18 positive cell line. J Shantou Univ Med Coll 2006; 19: 136–8.
Talora C, Sgroi DC, Crum CP, Dotto GP . Specific down-modulation of Notch1 signaling in cervical cancer cells is required for sustained HPV-E6/E7 expression and late steps of malignant transformation. Genes Dev 2002; 16: 2252–63.
Nefedova Y, Cheng P, Alsina M, Dalton WS, Gabrilovich DI . Involvement of Notch-1 signaling in bone marrow stroma-mediated de novo drug resistance of myeloma and other malignant lymphoid cell lines. Blood 2004; 103: 3503–10.
Zur HH . Papillomaviruses causing cancer: Evasion from host-cell control in early events in carcinogenesis. J Natl Cancer Inst 2000; 92: 690–8.
Zur HH . Papillomavirus infections-a major cause of human cancers. Biochim Biophys Acta 1996; 1288: 55–78.
Thierry F, Spyrou G, Yaniv M, Howley P . Two AP1 sites binding JunB are essential for human papillomavirus type 18 transcription in keratinocytes. J Virol 1992; 66: 3740–8.
Scheffner M, Werness BA, Huibregtse JM, Levine AJ, Howley PM . The E6 oncoprotein encoded by human papillomavirus types 16 and 18 promotes the degradation of p53. Cell 1990; 63: 1129–36.
Taylor WR, Tark GR . Regulation of the G2/M transition by p53. Oncogene 2001; 20: 1803–15.
Bunz F, Dutriaux A, Lengauer C, Waldman T, Zhou S, Brown JP, et al. Requirement for p53 and p21 to sustain G2 arrest after DNA damage. Science 1998; 282: 1497–501.
Lu ZM, Liu HT, Xu PR, Hou GQ, Xue LX . Expression of Notch1 gene in esophageal squamous cell carcinoma cell line EC9706 and its efect on cell apoptosis. Ai Zheng 2007; 26: 1074–9.
Acknowledgements
This project was supported by a grant from the National Natural Science Foundation of China (30570773).
We are grateful to Prof Tom KADESCH (Department of Genetics, University of Pennsylvania School of Medicine, Philadelphia, PA, USA) for providing the pcDNA3.1C-ICN plasmid.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, Kj., Lu, Qy., Niu, Xq. et al. Notch1 signaling inhibits growth of EC109 esophageal carcinoma cells through downmodulation of HPV18 E6/E7 gene expression. Acta Pharmacol Sin 30, 153–158 (2009). https://doi.org/10.1038/aps.2008.16
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2008.16