Abstract
Aim:
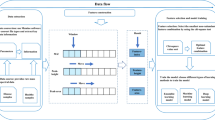
A discrimination analysis has been explored for the probabilistic classification of healthy versus ovarian cancer serum samples using proteomics data from mass spectrometry (MS).
Methods:
The method employs data normalization, clustering, and a linear discriminant analysis on surface-enhanced laser desorption ionization (SELDI) time-of-flight MS data. The probabilistic classification method computes the optimal linear discriminant using the complex human blood serum SELDI spectra. Cross-validation and training/testing data-split experiments are conducted to verify the optimal discriminant and demonstrate the accuracy and robustness of the method.
Results:
The cluster discrimination method achieves excellent performance. The sensitivity, specificity, and positive predictive values are above 97% on ovarian cancer. The protein fraction peaks, which significantly contribute to the classification, can be available from the analysis process.
Conclusion:
The discrimination analysis helps the molecular identities of differentially expressed proteins and peptides between the healthy and ovarian patients.
Similar content being viewed by others
Article PDF
References
Liotta LA, Ferrari M, Petricoin E . Clinical proteomics: written in blood. Nature 2003; 425: 905.
Rai AJ, Chan DW . Cancer proteomics: Serum diagnostics for tumor marker discovery. Ann N Y Acad Sci 2004; 1022: 286–94.
Diamandis EP . Analysis of serum proteomic patterns for early cancer diagnosis: Drawing attention to potential problems. J Natl Cancer Inst 2004; 96: 353–6.
Ransohoff DF . Lessons from controversy: ovarian cancer screening and serum proteomics. J Natl Cancer Inst 2005; 97: 315–9.
Colantonio DA, Chan DW . The clinical application of proteomics. Clin Chim Acta 2005; 357: 151–8.
Diamandis EP . Mass spectrometry as a diagnostic and a cancer biomarker discovery tool: opportunities and potential limitations. Mol Cell Proteomics 2004; 3: 367–78.
Aebersold R . Mann M . Mass spectrometry-based proteomics. Nature 2003; 422: 198–207.
Pusch W, Flocco MT, Leung S, Thiele H, Kostrzewa M . Mass spectrometry-based clinical proteomics. Pharmacogenomics 2003; 4: 463–76.
Diamandis EP, van der Merwe D . Plasma protein profiling by mass spectrometry for cancer diagnosis: Opportunities and limitations. Clin Cancer Res 2005; 11: 963–5.
Soltys SG, Le QT, Shi G, Tibshirani R, Giaccia AJ, Koong AC . The use of plasma surface-enhanced laser desorption/ionization time-of-flight mass spectrometry proteomic patterns for detection of head and neck squamous cell cancers. Clin Cancer Res 2004; 10: 4806–12.
Gramolini AO, Peterman SM, Kislinger T . Mass spectrometry-based proteomics: a useful tool for biomarker discovery? Clin Pharmacol Ther 2008; 83: 758–60.
Petricoin E III, Ardekani A, Hitt B, Levine P, Fusaro V, Steinberg S, et al. Use of proteomic patterns in serum to identify ovarian cancer. The Lancet 2002; 359: 572–7.
Banks RE, Stanley AJ, Cairns DA, Barrett JH, Clarke P, Thompson D, et al. Influences of blood sample processing on low-molecular-weight proteome identified by surface-enhanced laser desorption/ionization mass spectrometry. Clin Chem 2005; 51: 1637–49.
Baggerly KA, Morris JS, Edmonson SR, Coombes KR . Signal in noise: evaluating reported reproducibility of serum proteomic tests for ovarian cancer. J Nat Cancer Inst 2005; 97: 307–9.
Ransohoff DF . Bias as a threat to the validity of cancer molecular-marker research. Nat Rev Cancer 2005; 5: 142–9.
White CN, Zhang Z, Chan DW . Quality control for SELDI analysis. Clin Chem Lab Med 2005; 43: 125–6.
Wong JW, Sullivan MJ, Cartwright HM, Cagney G . msmsEval: tandem mass spectral quality assignment for high-throughput proteomics. BMC Bioinformatics 2007; 8: 51.
Flikka K, Martens L, Vandekerckhovel J, Gevaert K, Eidhammer I . Improving the reliability and throughput of mass spectrometry-based proteomics by spectrum quality filtering. Proteomics 2006; 6: 2086–94.
Author information
Authors and Affiliations
Corresponding author
Additional information
This project was supported by the Zhejiang Provincial Science and Technology Foundation of China (No 2005C13026).
Rights and permissions
About this article
Cite this article
Hong, Yj., Wang, Xd., Shen, D. et al. Discrimination analysis of mass spectrometry proteomics for ovarian cancer detection. Acta Pharmacol Sin 29, 1240–1246 (2008). https://doi.org/10.1111/j.1745-7254.2008.00861.x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1111/j.1745-7254.2008.00861.x
Keywords
This article is cited by
-
Identification of MST1 as a potential early detection biomarker for colorectal cancer through a proteomic approach
Scientific Reports (2017)
-
Proteomic Study of Hepatic Nuclear Extracts in an Adaptive Acetaminophen Tolerance Model
Clinical Proteomics (2009)