Abstract
Carbon nanomaterials field-effect transistor (FET)-based electrical biosensors provide significant advantages over the current gold standards, holding great potential for realizing direct, label-free, real-time electrical detection of biomolecules in a multiplexed manner with ultrahigh sensitivity and excellent selectivity. The feasibility of integrating them with current complementary metal oxide semiconductor platform and a fluid handling module using standard microfabrication technology opens up new opportunities for the development of low-cost, low-noise, portable electrical biosensors for use in practical future devices. In this article, we review recent progress in the rapidly developing area of biomolecular interaction detection using FET-based biosensors based on the carbon nanomaterials single-walled carbon nanotubes (SWNTs) and graphene. Detection scenarios include DNA–DNA hybridization, DNA–protein interaction, protein function and cellular activity. In particular, we will highlight an amazing property of SWNT- or graphene-FETs in biosensing: their ability to detect biomolecules at the single-molecule level or at the single-cell level. This is due to the size comparability and the surface compatibility of the carbon nanomaterials with biological molecules. We also summarize some current challenges the scientific community is facing, including device-to-device heterogeneity and the lack of system integration for uniform device array mass production.
Similar content being viewed by others
Main
A biosensor is an analytical device that incorporates a biological recognition element in direct spatial contact with a transduction element. This integration ensures the rapid and convenient conversion of the biological events to detectable signals.1 With the discovery of rich nanomaterials and the development of exquisite nanofabrication tools, such as electron beam lithography, focused ion beam and nanoimprint lithography, new avenues have been opened up in the field of biosensors in the last two decades.2, 3 In particular, researchers around the world have been tailor-making a multitude of nanomaterials-based electrical biosensors and developing new strategies to apply them in ultrasensitive biosensing. Examples of such nanomaterials include carbon nanotubes,4, 5, 6, 7, 8, 9, 10, 11, 12, 13 nanowires,11, 14, 15, 16, 17, 18, 19, 20, 21 nanoparticles,6, 22, 23, 24, 25 nanopores,26, 27 nanoclusters28 and graphene.5, 29, 30, 31, 32 Compared with conventional optical, biochemical and biophysical methods, nanomaterial-based electronic biosensing offers significant advantages, such as high sensitivity and new sensing mechanisms, high spatial resolution for localized detection, facile integration with standard wafer-scale semiconductor processing and label-free, real-time detection in a nondestructive manner.
Among diverse electrical biosensing architectures, devices based on field-effect transistors (FETs) have attracted great attention because they are an ideal biosensor that can directly translate the interactions between target biological molecules and the FET surface into readable electrical signals.33, 34, 35, 36, 37, 38 In a standard FET, current flows along a semiconductor path (the channel) that is connected to two electrodes, (the source and the drain). The channel conductance between the source and the drain is switched on and off by a third (gate) electrode that is capacitively coupled through a thin dielectric layer. In conventional complementary metal oxide semiconductor-fabricated transistors, the conducting channel is buried inside the substrate; in FET-based biosensors, the channel is in direct contact with the environment, and this gives better control over the surface charge. This implies that surface FET-based biosensors might be more sensitive: biological events occurring at the channel surface could result in the surface potential variation of the semiconductor channel and then modulate the channel conductance. In conjunction with the ease of on-chip integration of device arrays and the cost-effective device fabrication, the surface ultrasensitivity places FET-based biosensors as attractive alternatives to existing biosensor technologies. In this review, we summarize recent progress in ultrasensitive biosensors formed from nanomaterials-based FETs. We pay particular attention to carbon nanomaterials: single-walled carbon nanotube (SWNT) and graphene. This research field is a diverse and rapidly growing one; having limited space and references, we will only be able to cover several of the major contributions. We will highlight some important aspects including strategies to increase sensitivity, dynamic detection in cells and liquid environment, DNA hybridization and single-molecule detection, as these have been neglected in most previous reviews. Fortunately, there are a number of excellent previous review papers in the literature covering various aspects of carbon nanomaterials-based biosensors, which can amend these deficiencies.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 30, 34
Basic introduction to carbon nanomaterials
There has been an explosion of interest in use of carbon nanomaterials in new nanoscale biosensors. Owing to their unique physicochemical properties, SWNTs and, very recently, graphene are in the forefront of this explosion. Graphene is a two-dimensional (2D) crystalline monolayer made of sp2-hybridized carbon atoms arranged in a honeycomb lattice.39, 40, 41 In fact, graphene is the basic building block for graphitic materials of all other dimensionalities. For example, by folding up a graphene sheet into a cylinder along a certain lattice vector, a well-ordered, hollow graphitic nanomaterial, classified as a SWNT, is formed.42 Both of these two allotropes of carbon have the simplest chemical composition and atomic bonding configuration in a 2D manner that maximizes its surface-to-volume ratio. Each carbon atom on the nanocarbon surface is exposed to the environment and any small changes in the environment can result in drastic changes in the electrical properties of the nanocarbon device, thus forming the basis for ultrasensitive biosensing.43, 44, 45 As a single biomolecule or a small collection of biomolecules approach the SWNT or graphene-based device, it or they can alter the electronic properties of the device via one or more of the following mechanisms: (1) surface charge-induced gating, (2) charge transfer or doping between carbon nanomaterials and biomolecules, (3) a scattering potential across carbon nanomaterials, and (4) Schottky barrier modification between carbon nanomaterials and metal electrodes.46, 47, 48, 49 In the case of solution-gate FETs, the modulation of the channel conductance is achieved by applying a gate potential at the solid/liquid interface from a reference electrode placed on top of the channel across the electrolyte that acts as the dielectric. The conductance of SWNTs and graphene shows the strong dependence on the ionic strength, the pH and the type of ions present. The sensitivity to electrolyte composition is attributed to a combination of different mechanisms including electrostatic gating, Schottky barrier medications and changes in gate capacitance.50
The second significant feature of SWNTs and graphenes is that they are 2D conducting nanomaterials that are intrinsically the same size as the molecules. This size compatibility offers the opportunity of detecting single-molecule events.51, 52
Third, SWNTs and graphenes are molecular chemicals entirely composed of carbon atoms, and this suggests a natural match to biomolecules. This also allows controlled functionalization to selectively immobilize bioaffinitive agents on their surfaces. This can establish efficient electrical communication with biological analytes using one of two alternative methods: noncovalent adsorption or wrapping of various bioaffinitive agents on the carbon surfaces through hydrophobic interactions,53 or covalent attachment to the functional groups produced through chemical reactions on the carbon π-conjugated skeleton.10, 54 Ultimately, the nanocarbon reagents are large and can be easily micro/nanofabricated on a large range of substrates. These major advantages of SWNTs and graphenes combined with their high conductivity, high chemical stability and easy availability through bottom-up approaches suggest that carbon nanomaterials-based FETs have promise as paradigms for device architecture, yielding devices that are capable of converting biological information to easily detectable electrical signals.
SWNT FET-based biosensors
SWNT−dynamic detection in living cells
Nanostructured carbon materials have shown promising potential as novel nanoelectronic biosensors for biomolecular detection; they are extraordinarily sensitive and their detection schemes are quite simple. Most of the realizations so far, however, have been limited to static in vitro measurement. The dynamic detection of the release of biomolecules from living cells in real time is important both in fundamental studies and in the evaluation of drugs for the treatment of secretion-related diseases. To this goal, Huang et al.55 utilized a SWNT network to directly interface with living neuroglial astrocytes and, without labels, detect the triggered release of ATP from these cells. This detection scheme showed high temporal resolution. Highly charged ATP molecules secreted from the astrocyte diffused into the conductive channel of the FET and electrostatically modulated the SWNT conductance, leading to measurable current responses. Heller et al.56 used SWNTs in a contact-passivated, suspended layout to allow close contact between the cell and the SWNT. They followed the process of phagocytosis in real time by simultaneously monitoring both changes in transistor conductance (FET signal) and changes in the electrochemical current (signal), which suggests successful detection of cellular activity. They also demonstrated that the sensitivity for certain electrochemical processes could be enhanced when the SWNT was coated with catalytic platinum nanoparticles. The goal of another study by the Chen and colleagues57 was to improve the biocompatible interactions between SWNTs and living cells; they demonstrated that noncovalent functionalization of SWNTs with bioactive sugar moieties conferred biocompatibility without compromising the sensing capabilities of their devices. The SWNT network was first surface-functionalized via π–π interactions with bioactive sugars (N-acetyl-D-glucosamine) to allow PC12 cells to adhere and grow on the SWNT-net substrate. When the solution with high K+ content was administrated to the cells to evoke Ca2+ influx through voltage-gated Ca2+ channels and consequently trigger exocytosis of secretory vesicles, single-cell secretion of catecholamine molecules occurred, and this resulted in current responses (spikes) of glycosylated SWNT-net FETs (Figures 1a and b). This is because the aromatic rings of the catecholamines at the cell–nanotube junction attach noncovalently to the nanotube sidewall and thereby impose a p-doping effect that increases nanotube conductance. A similar approach of surface functionalization was also reported previously by Wang et al.,58 who used single-SWNT-based FETs to detect the acute release of chromogranin A at low concentration (1 nM) from living cortical neurons and monitor dose-dependent chromogranin A release from a single bovine chromaffin cell positioned above the sensing region by a micropipette and stimulated by histamine.59 These studies provide a real-time and noninvasive measurement platform to examine subtle cellular activities from living cells with high temporal resolution and ease of detection.
(a) Triggered exocytosis and SWNT-net detection. (b) Nanotube responses to exocytosis of PC12 cells triggered by high K+ stimulation. The SWNT-net was biased at Vds=0.4 V. Adapted from Sudibya et al.57 (© 2009 Wiley-VCH). (c) Schematic of the side view of a SWNT FET inside a polydimethylsiloxane (PDMS) microfluidic channel. (d) Scaled schematic of a carbon nanotube under a 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) bilayer. The DOPC lipid molecules are shown with two light green fatty acid chains. A toxin protein, represented in a ribbon model, binds to a membrane-embedded ganglioside molecule that is shown with two dark green chains. Adapted from Zhou et al.60 (© 2007 NPG).
SWNT−functionalization to increase sensitivity
The properties of SWNTs can be leveraged by combining them with other functional materials, such as small molecules, lipid membrane or aptamers, to enhance the biosensing performance and install new functionalities. For example, Zhou et al.60 showed the integration of supported lipid bilayers with SWNT FETs (Figures 1c and d). They first demonstrated membrane continuity and lipid diffusion over the tube. Then they showed that the nanotube acted as a diffusion barrier for a membrane-bound tetanus toxin protein, and that the strength of the barrier depended on the diameter of the nanotube. Finally, they present results on the electrical detection of specific binding of streptavidin to biotinylated lipids. The formation of fluidic supported lipid bilayers on SWNTs allowed the study of lipid–SWNT interactions and sensing of analytes binding to specific receptors embedded in the supported lipid bilayers. Kim et al.61 coated SWNT surfaces with human olfactory receptor 2AG1 and reported the real-time detection of specific odorant molecules (such as amyl butyrate, butyl butyrate, propyl butyrate and pentyl valerate). The observed femtomolar sensitivity resulted from the deformation of the human olfactory receptor 2AG1 protein when the WNT-bound protein and the odorants docked. Vedala et al.62 used SWNT FETs that were noncovalently functionalized with porphyrin-based glycoconjugates synthesized using ‘click’ azide-alkyne chemistry. They realized a highly selective electrical detection of interactions between glycoconjugates and bacterial lectins (PA-IL and PA-IIL from Pseudomonas aeruginosa and a plant lectin concanavalin A, respectively), which exhibit specific multivalent binding to carbohydrates. So et al.63 functionalized SWNT FETs with RNA-based Escherichia coli aptamers, which can be selectively recognized by E. coli. By using these aptamer-functionalized SWNT FETs combined with the most probable number method, they demonstrated a reliable screening tool for microorganisms such as E. coli. However, there is a difference of two orders of magnitude in titer estimation between the culture method and the most probable number-combined SWNT FETs. This is probably due to the small number of examples studied and the small sensing surfaces of devices. Villamizar et al.64 reported a method of using anti-Salmonella antibodies to functionalize SWNT network-based FETs, which were subsequently protected by Tween 20 to prevent nonspecific binding (NSB) of other proteins, for the selective determination of S. infantis at low concentration (at least 100 c.f.u. ml−1). These studies show that functionalized SWNT FET-based promise to be useful platforms for detecting microorganisms, such as antibodies, fungi, bacteria or viruses, with high selectivity.
SWNT–DNA hybridization
The detection of DNA hybridization has been a topic of central importance owing to its use in the diagnosis of pathogenic and genetic diseases. Previous reports indicated that single-stranded DNA can form a stable complex with individual SWNTs by wrapping around them, the interaction being driven by the aromatic interactions between nucleotide bases and SWNT sidewalls. Label-free electrical detection of DNA hybridization utilizing SWNT FET-based biosensors suggests a new generation of DNA chips that can give direct electrical readouts. This could provide fast and inexpensive analyses of nucleic acid samples. In 2006, Star et al.65 used SWNT FETs to study interactions between single-stranded DNA oligonucleotides and SWNTs and the subsequent DNA hybridization processes that occurred on the device surface. They showed that SWNT FETs could be selectively modified with DNA oligonucleotides and maintain hybridization specificity. Then they demonstrated that SWNT FETs with immobilized synthetic oligonucleotides could recognize target DNA sequences with single-nucleotide polymorphism and that electronic signal output from the SWNT FET biochips clearly differentiated between mutant and wild-type alleles of the HFE gene (the gene that is responsible for hereditary hemochromatosis). To ensure specific adsorption of DNA to SWNTs, Martinez et al.66 developed a new approach for making functionalized SWNT FET arrays to detect DNA hybridization (Figure 2a). In that study aminated single-strand DNA was covalently attached to the polymer poly (methylmethacrylate0.6-co-poly(ethyleneglycol)methacrylate0.15-co-N-succinimidylmethacrylate0.25) that was noncovalently absorbed to the nanotube surface. This method of anchoring the probe DNA can prevent the nonspecific adsorption of DNA molecules onto SWNTs that can occur by virtue of aggregation on the sidewalls of the contact electrodes. They demonstrated that DNA hybridization led to statistical changes in the threshold voltages resulting from the charge trapping character of hybridized DNA. Although label-free electrical detection of DNA and biomolecules using SWNT FETs has been successfully achieved, the detection limits are typically on the order of ca. 1 nM of DNA. To improve the sensitivity, Dong et al.67 reported a novel ‘nanoparticle enhancement’ approach (Figure 2b), in which the target DNAs are hybridized with probe DNAs on the device, and reporter DNAs labeled with Au nanoparticles flank a segment of the target DNA sequence. Before detection, they blocked the SiO2 surface by octyltrichlorosilane treatment to reduce possible NSB of DNA to SiO2 and vacant SWNT surfaces by using polyethylene glycol to reduce the NSB of DNA to SWNTs. With careful experiments, they demonstrated that the detection sensitivity of SWNT FET-based biosensors for DNA can be improved to ca. 100 fM.
(a) Schematic representation of the bonding of the polymer to SWNT (top right and left) and DNA hybridization (lower). Adapted from Martinez et al.66 (© 2009 ACS). (b) Schematic illustration of DNA detection enhancement by reporter DNA–Au nanoparticle conjugates. The bottom right panel illustrates the possible molecular binding on SWNTs. Adapted from Dong et al.67 (© 2008 Wiley-VCH).
SWNT−single-molecule detection
A key goal of modern bioscience is monitoring biomolecular interactions with high sensitivity and high selectivity in real time, with the ultimate aim of detecting single-molecule events in natural samples. SWNT FET-based biosensors hold the promise of detecting single-molecule processes because of the size compatibility of SWNTs with biological entities. In the following discussion, we will highlight the recent advances of the burgeoning field of single-molecule biosensing. Most of this research thus far has dealt with proofs of concept and has used biological solutions or samples made in the laboratory rather than natural samples.
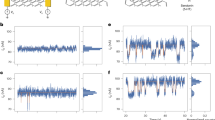
Recently, Goldsmith et al.51 have developed an electrochemical method to create single point defects in SWNTs in a controllable manner and then covalently bind biomolecules at this scattering site. Owing to the real-time monitoring of conductance during the defect generation, these point-functionalized SWNT FETs can be prepared in high yield. Using these devices, they also demonstrated continuous, multihour monitoring of the binding of a single molecule with high sensitivity (a conductance change of >100 nS for binding of a reactive carbodiimide). This sensitivity is due to the Coulomb interaction between the molecule and the defect that modulates scattering in the 1D channel.52 This approach provides a new electronic platform for studying biomolecular interactions and kinetics that are hidden in ensemble measurements, as demonstrated by Sorgenfrei et al.68 In this study, they covalently attached a single-stranded probe DNA sequence, which was terminated with an amine group, to a carboxylic acid-functionalized point defect in a carbon nanotube using a standard amide-formation coupling reaction. After probe DNA was attached, these devices were used to study the kinetics and thermodynamics of DNA hybridization with the experimental setup shown in Figure 3a. In the absence of target DNA, the devices did not show any particular features in a conductance dominated by flicker (1/f) noise, as shown in Figures 3b. When the device was immersed in buffer containing complementary target DNA, however, reproducible large-amplitude two-level fluctuations appeared at different temperatures, as shown in Figures 3d. Conductance differences reached ∼60–100 nS and the signal-to-noise ratio >3 (over the 1/f noise background) over a time interval of 30 s. This observation can be explained by the proposed model that the device conductance is controlled by probe-target hybridization that decreases the device conductance because of increased scattering and charge transfer at the position of target DNA binding. This effect would be partially offset by the Debye screening from the dissolved solution counter ions for longer DNA strands. Further kinetic investigations of the system as a function of temperature demonstrated non-Arrhenius behavior; this agrees with DNA hybridization experiments using fluorescence correlation spectroscopy. This technique is label-free and could be used to probe single-molecule dynamics at microsecond timescales.
(a) Schematic of the nanotube device used for studying the kinetics and thermodynamics of DNA hybridization (with an external circular heater/refrigerator to control temperature). (b, d, h) Conductance recordings of devices over one 30-s interval with DNA oligonucleotide probe H2N-5′-GGAAAAAAGG-3′ without exposure to complementary DNA target at 35.0 °C (b), after exposure to complementary DNA target at 33.0 °C (d) and after exposure to complementary DNA target at 21.0 °C (h). The source–drain bias is 100 mV and VPt is zero. (c, e, i) Conductance-based histograms of time intervals shown in b, d and h. The two levels in e and i are fit to the Gaussian distributions. (f, g) Representative short time interval (40 ms) for real-time conductance recording for probe only and after exposure to complementary target DNA. Adapted from Sorgenfrei et al.68 (© 2011 NPG).
An alternative to the aforementioned single-point functionalization for single-molecule biosensing is the fabrication of molecular electronic devices from single molecules that can be subsequently derivatized with a probe molecule that is capable of biomolecular recognition. To do this, a new system was recently developed for measuring the conductance of a single molecule covalently immobilized within a nanotube gap (Figure 4a).69 In this system, gaps are formed in carboxylic-acid-functionalized SWNTs that can be reconnected by one or a few molecules attached to both sides of the gap through amide bond formation. Consequently, the devices are sufficiently robust so that a wide range of chemistries and conditions can be applied. By using this method, we have made molecular devices that detect the binding between proteins and substrates at the single-event level, and that probe the dependence of charge transport of a single intact DNA duplex on its Π-stacking integrity.70 However, biomolecular interactions cannot be measured in real time. To do this, a useful strategy has been recently developed to create an integrated system that can combine rapid real-time measurements with single-molecule sensitivity.71 In this study, individual DNA aptamers were coupled with SWNTs as point contacts to form single-molecule devices that allow us to selectively and reversibly detect a target protein, thrombin (Figure 4b). After further thrombin treatment, these fresh aptamer-functionalized devices showed consistent conductance increases originating from the enhanced DNA charge transfer that is due to the rigidification of DNA conformation by DNA–thrombin interactions. To achieve real-time measurements, a repeating pattern that consists of 79 identical SWNT transistors by a double photolithographic process was designed and fabricated (Figure 4c). Combining this design with microfluidics (Figure 4c, inset) allowed us both to detect proteins and to monitor stochastic DNA–protein interactions in real time (Figure 4d). Reversible and equivalent conductance changes at different thrombin concentrations (from 2.6 fM to 2.6 pM and 2.6 nM; Figure 4d) were observed, thus demonstrating single-molecule sensitivity. Further delivery of elastase (3.4 nM) did not lead to any detectable conductance change in the same device. In a separate experiment, we observed negligible conductance changes upon thrombin injection using the devices reconnected with a different DNA (Con-A), which could not bind human thrombin, (2.6 fM; Figure 4e). Both control experiments demonstrated that this protein detection scheme has excellent selectivity. These results distinguished this method as a valuable platform to achieve real-time, label-free, reversible detection of DNA–protein interactions with high selectivity and real single-molecule sensitivity.
(a) Schematic of single-molecule devices using SWNT point contacts. Adapted from Feldman et al.70 (© 2008 ACS). (b) Schematic representation of the sensing design showing how single-molecule devices can detect proteins at the single-molecule level. (c) Scanning electron microscopy images showing highly integrated identical SWNT devices. Inset shows an optical image of a single-molecule device during real-time measurements. (d) Current-versus-time data recorded for an aptamer-rejoined device upon alternate additions of the thrombin Tris-HCl buffer solution at different concentrations (from 2.6 fM to 2.6 pM and 2.6 nM), the 6-M guanidine HCl solution, and finally the elastase (3.4 nM) Tris-HCl buffer solution. (e) I–V plot recorded for a Con-A-rejoined device upon injection of 2.6 fM thrombin in Tris-HCl buffer solution. All the measurements were performed at VD=−50 mV and VG=0 V. Adapted from Liu et al.71 (© 2011 Wiley-VCH).
Graphene FET-based biosensors
Graphene, a new allotrope of carbon made of sp2-hybridized carbon atoms arranged in a honeycomb lattice, is an ideal 2D system with remarkable electronic, physical and chemical properties. When incorporated into electrical devices, graphene shows high mobilities of both holes and electrons. It is structurally uniform, that is, it shows no structural variations analogous to those of diameter and chirality that characterize SWNTs. Even though it is chemically quite stable, its reaction chemistry has been developed to the extent that flexible chemical modifications are possible. High-quality graphenes are available from mechanical exfoliation from graphite, reduction of graphene oxides (GOs), chemical vapor deposition growth and chemical synthesis. It has also been proven to be easily incorporated into devices. In conjunction with its subnanoscale thickness and surface compatibility with various biological species, these properties make graphene a promising building block for developing new types of nanoscale biosensors. Recently, the increasing interest in graphene for biosensing applications seems to have overtaken the corresponding interest in SWNTs.
Graphene−strategies to increase sensitivity
The first graphene FET-based electrical biosensors were demonstrated by Mohanty et al.,72 who made graphene transistors from chemically modified graphenes (CMGs), such as GOs or graphene amines (GAs) obtained by treating GOs with nitrogenous plasmas or ethylenediamne. They immobilized GO and GA sheets on silica substrates with either predeposited or postdeposited gold electrodes, and used silicon as a backgate for electrical measurements. They exploited the functional groups on GOs or GAs to fabricate three CMG-based biosensors: (i) a single-bacterium biodevice, (ii) a label-free DNA sensor and (iii) a bacterial DNA/protein, and a polyelectrolyte chemical transistor. The bacteria biodevice was highly sensitive; just a single bacterium attachment caused a sharp (42%) increase in conductivity of CMG sheets (Figure 5a). This can be attributed to the p-type characteristic of CMGs, where the attachment of a negatively charged species, such as bacteria, is equivalent to a negative potential gating, and this gating increases the hole density and thus the conductivity of CMG. Similarly, single-stranded DNA tethered on graphene reversibly increased the device conductance during cycles of dehybridization and rehybridization with its complementary DNA strand (Figure 5b). Finally, they demonstrated that adsorption of polyelectrolyte molecules on the modified graphenes resulted in polarity-dependent changes in electrical conductivity.
(a) The conductivity of the p-type GA device increases upon attachment of a single bacterial cell on the surface of GA (the device is shown in inset 1). (b) DNA transistor: single-stranded DNA (ssDNA) tethering on GO increases the conductivity of the device. Successive hybridization and dehybridization of DNA on the G-DNA device results in completely reversible increase and restoration of conductivity. Inset shows a G-DNA(ds) sheet with wrinkles and folds clearly visible. Adapted from Mohanty et al.72 (© 2008 ACS).
To improve the detection sensitivity, Dong et al.73 used large, CVD-grown graphene films to fabricate biosensors, thereby achieving DNA sensing with a concentration as low as 0.01 nM. They attribute this marked increase in sensitivity to the electronic n-doping to the devices. They further found that adding AuNPs on the surface of graphene devices could extend the upper limit of DNA detection from 10 to 500 nM due to the increase in loading of probe DNA molecules onto the AuNPs. This AuNP strategy is apparently widely applicable: Mao et al.74 demonstrated a specific protein detection biosensor using reduced GO sheets decorated with Au nanoparticle–antibody conjugates (Figure 6a). The response of the sensor increases with the increase of protein concentrations and saturates at 0.02 mg ml−1. The lower detection limit of the sensor is on the order of ng ml−1 and could be further improved by optimizing the device structure. To achieve real-time biomolecular sensing, Huang et al.75 demonstrated the use of large-sized CVD-grown graphene films configured as field-effect transistors for detecting glucose or glutamate molecules by the conductance changes of the graphene transistors as the molecules were being oxidized by the specific redox enzyme (glucose oxidase or glutamic dehydrogenase) attached to the graphene film.
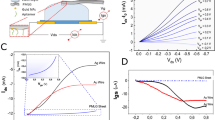
(a) Schematic of a reduced GO FET. Anti-immunoglobulin G (IgG) was anchored to the reduced GO sheet surface through Au nanoparticles and functions as a specific recognition group for the IgG binding. Adapted from Mao et al.74 (© 2010 Wiley-VCH). (b) Schematic illustration of the device structure for solution-gated graphene FETs. Adapted from Ang et al.77 (© 2008 ACS). (c) Schematic illustration of apatmer-modified electrolyte-gated graphene FETs. (d) Time course of ID for an aptamer-modified graphene FET. At 10-min intervals, various concentrations of IgE were injected. Adapted from Ohno et al.79 (© 2010 ACS).
Graphene−detection in liquid environment
So far, most graphene FETs have operated either under vacuum or atmospheric conditions, not in solution. In some cases, high-quality graphene can be grown epitaxially on SiC substrates. Because of the thickness of the intrinsic substrate on SiC, to achieve field effect responses, top gating of the epitaxial graphene is required in most cases except that based on nitrogen implantation into a SiC wafer before graphene growth.76 Previous reports have proved that an efficient approach for top gating is ‘solution top-gating’. Only very recently has the operation of graphene in aqueous electrolytes (Figure 6b) for use in biosensors and bioelectronics been reported by several groups.77, 78, 79, 80, 81, 82, 83, 84, 85, 86 For example, Ang et al.77 first demonstrated the use of solution-gated epitaxial graphene as a pH sensor. Ohno et al.78 reported on electrolyte-gated graphene field-effect transistors for detecting pH and protein adsorption. In another report by Ohno et al.,79 they demonstrated label-free immunosensing based on an aptamer-modified graphene FETs (Figure 6c). Immunoglobulin E aptamers with an approximate height of 3 nm were successfully immobilized on a graphene surface, as confirmed by atomic force microscopy. The aptamer-modified graphene FETs showed selective electrical detection of immunoglobulin E protein. From the dependence of the drain current variation on the immunoglobulin E concentration, the dissociation constant was estimated to be 47 nM, indicating good affinity (Figure 6d). In a similar way, Agarwal et al.87 recently demonstrated the biocompatibility of reduced GOs with proteins and further used them after protein functionalization to create biosensors for detecting various metals in real time with high sensitivity.81 These reports already confirm the potential of graphene for sensing in aqueous electrolytes, however, detailed understandings of the graphene/electrolyte interface and the effect of the electrolyte on the electronic transport in graphene are still lacking. To address these issues, Dankerl et al.82 developed a facile method for the scalable fabrication of graphene FET arrays and provided a comprehensive characterization of operation of these devices in aqueous electrolytes. By using in-solution Hall-effect measurements and taking into account the microscopic structure of water at the interface, they demonstrated that charge carrier mobilities and concentrations as a function of electrolyte gate potential can be directly determined. They also showed that graphene FETs exhibited a high transconductance and correspondingly high sensitivity together with an effective gate noise as low as tens of μV. These studies demonstrated that graphene FETs, with their ease of fabrication, high transconductance, and low noise, hold great promise for biosensor and bioelectronic applications.
Graphene−detection in cells
To further investigate the biocompatibility of graphene with live cells or tissue, Cohen-Karni et al.83 demonstrated for the first time recording from eletrogenic cells using single-layer graphene formed by mechanical exfoliation from graphite and carried out simultaneous recording using graphene and silicon nanowire FETs (Figure 7a). They found that graphene FET conductance signals recorded from spontaneously beating embryonic chicken cardiomyocytes yielded well-defined extracellular signals with signal-to-noise ratio routinely >4 (Figure 7b). Water gate (Vwg)-dependent experiments demonstrated that the conductance signal amplitude could be tuned over nearly an order of magnitude, thus showing a robust graphene/cell interface. Furthermore, by varying Vwg across the Dirac point, they achieved the expected signal polarity flip, thus allowing both n- and p-type recording to be achieved with the same device simply by offsetting Vwg. Finally, they compared peak-to-peak recorded signal widths (made as a function of graphene FET device size) with those made using silicon nanowire-FETs and showed that the widths increased with the area of graphene FET devices. This indicates that they were measuring a signal that was averaged from different points across the outer membrane of the beating cells. In another recent work, Agarwal et al.87 demonstrated the biocompatibility of reduced GOs with live cells, such as neuroendocrine PC12 cells, and further used them to create biosensors for detecting the dynamic secretion of the hormonal catecholamine molecules from living cells.80 Pursuing the development of a graphene-based technology that can detect action potentials from electrically active cells, Hess et al.84 recently reported using arrays of CVD-grown graphene FETs for the extracellular detection of action potentials from electrogenic cells (Figures 7c and d). The action potentials of cardiomyocyte-like HL-1 cells could be effectively resolved and tracked across the transistor array as shown in Figure 7e. The low-noise characteristic of graphene FETs together with the large transconductive sensitivity of these devices clearly indicates an advantage of graphene FETs in terms of signal-to-noise ratio. These studies suggest that the outstanding performance of graphene FETs together with the feasibility of easily integrating graphene electronics with flexible substrates can pave the way for a true breakthrough in bioelectronics, in particular for electrically functional neural prostheses.
(a) Representation of the relative size of cardiomyocyte cell interfaced to typical Graphene and silicon nanowire-FET devices. (b) Recorded traces at different applied water gate potentials. The following blue, green and red traces were recorded at +0.05, +0.10 and +0.15 V, respectively. The corresponding sensitivities are 2020, 398 and 2290 μS V−1, respectively. Adapted from Cohen-Karni et al.83 (© 2010 ACS). (c) Schematic view of a graphene FET with a cell on the gate area. The graphene is shown between the drain and source metal contacts, which are protected by a chemically resistant layer. (d) Combination of an optical microscopy image of a transistor array and a fluorescence image of the calcein-stained cell layer on the same array. The scale bar is 100 μm. (e) Simultaneous current recordings of eight transistors in one FET array over tens of seconds (left) and hundreds of milliseconds (right). Adapted from Hess et al.84 (© 2011 Wiley-VCH).
Summary and perspective
In this article, we reviewed some recent advances in the rapidly developing area of biomolecule detection using carbon nanomaterial FET-based biosensors. We feel these biosensors have had substantial impact and have genuine potential for future applications. By taking advantage of the outstanding electrical properties and the environmental ultrasensitivity of carbon nanomaterials many strategies have been developed to create SWNT or graphene FET-based biosensors for directly detecting DNA–DNA hybridizations, DNA–protein interactions, protein functions and cellular activities in real time with high sensitivity and excellent selectivity. In particular, because of the size comparability and the surface compatibility with biological molecules, an amazing feature of using SWNT or graphene FETs in biosensing is their ability to detect biomolecules at the single-molecule level, as well as at the single-cell level. Although SWNT and graphene share some common functionalities, such as high conductivity and high chemical stability, each has its own advantages and disadvantages. For example, SWNTs are 1D and can be produced in bulk quantities while large-scale production of high-quality graphene is still difficult. In contrast, owing to its 2D nature graphene is ideal for forming larger interfaces with biomolecules, and graphene-based could be produced using traditional lithographic techniques down to the nanoscale. However, when incorporated into the proper device architecture and combined with direct electrical transduction, the associated signal processing circuitry, and a fluid handling module (fabricated using standard microfabrication technology), the distinct and complementary capabilities of SWNT or graphene FETs could open up new opportunities for the development of bioelectronics toward practical applications in the future.
Despite this remarkable progress, none of the nanocarbon FET-based electrical biosensors has yet advanced to the production scale. The major challenge primarily stems from the device-to-device heterogeneity of baseline electronic properties. This is because of the absence of diameter and chirality control for SWNTs and the difficulty of controlling layer number and surface cleanness of graphene. Developing a reliable and scalable fabrication technique for mass-producing identical carbon nanomaterials arrays and integrating individuals into functional devices with high yields are some of the technical issues to be addressed in future. The next problem is system integration. Most of the studies reported in the literature used a sample delivery system, such as a syringe pump that was neither integrated to the sensing platform nor to the readout module. Furthermore, except for a few cases, most of the measurements were carried out in ideal media such as pure buffer solutions. The real physiological sample is far more complex and will definitely introduce a range of interfering and fouling effects. Multiplexing is also an attractive goal, and toward this end the fabrication of parallel and integrated biosensor arrays that have spatially resolved specific functionalities prepared using spatially resolved surface functionalizations remains challenging. Nevertheless, compared with the current gold standards, that is, PCR, mass spectrometry and enzyme-linked immunosorbent assay, nanocarbon FET-based electrical biosensors hold several significant advantages: simplicity, low-cost, portability, ultrahigh sensitivity, excellent selectivity and label-free, real-time, electrical detection. This suggests a thriving research field that can anticipate countless applications in a wide variety of areas: clinic diagnostics, environment preservation, health improvement, national defense and bioterrorism prevention.
References
Thevenot, D. R., Toth, K., Durst, R. A. & Wilson, G. S. Electrochemical biosensors: recommended definitions and classification. Biosens. Bioelectron. 16, 121–131 (2001).
Liu, J. W., Cao, Z. H. & Lu, Y. Functional nucleic acid sensors. Chem. Rev. 109, 1948–1998 (2009).
Rosi, N. L. & Mirkin, C. A. Nanostructures in biodiagnostics. Chem. Rev. 105, 1547–1562 (2005).
Huang, Y. X. & Chen, P. Nanoelectronic biosensing of dynamic cellular activities based on nanostructured materials. Adv. Mater. 22, 2818–2823 (2010).
Yang, W. R., Ratinac, K. R., Ringer, S. P., Thordarson, P., Gooding, J. J. & Braet, F. Carbon nanomaterials in biosensors: should you use nanotubes or graphene? Angew. Chem. Int. Ed. 49, 2114–2138 (2010).
Wang, J. Nanomaterial-based electrochemical biosensors. Analyst 130, 421–426 (2005).
Allen, B. L., Kichambare, P. D. & Star, A. Carbon nanotube field-effect-transistor-based biosensors. Adv. Mater. 19, 1439–1451 (2007).
Lahiff, E., Lynam, C., Gilmartin, N., O'Kennedy, R. & Diamond, D. The increasing importance of carbon nanotubes and nanostructured conducting polymers in biosensors. Anal. Bioanal. Chem. 398, 1575–1589 (2010).
Liu, Z., Tabakman, S., Welsher, K. & Dai, H. J. Carbon nanotubes in biology and medicine: in vitro and in vivo detection, imaging and drug delivery. Nano Res. 2, 85–120 (2009).
Liu, S., Shen, Q., Cao, Y., Gan, L., Wang, Z. X., Steigerwald, M. L. & Guo, X. F. Chemical functionalization of single-walled carbon nanotube field-effect transistors as switches and sensors. Coord. Chem. Rev. 254, 1101–1116 (2010).
Roy, S. & Gao, Z. Q. Nanostructure-based electrical biosensors. Nano Today 4, 318–334 (2009).
Gruner, G. Carbon nanotube transistors for biosensing applications. Anal. Bioanal. Chem. 384, 322–335 (2006).
Kauffman, D. R. & Star, A. Electronically monitoring biological interactions with carbon nanotub field-effect transistors. Chem. Soc. Rev. 37, 1197–1206 (2008).
Arlett, J. L., Myers, E. B. & Roukes, M. L. Comparative advantages of mechanical biosensors. Nat. Nanotechnol. 6, 203–215 (2011).
Song, S. P., Qin, Y., He, Y., Huang, Q., Fan, C. H. & Chen, H.-Y. Functional nanoprobes for ultrasensitive detection of biomolecules. Chem. Soc. Rev. 39, 4234–4243 (2010).
He, Y., Fan, C. H. & Lee, S.-T. Silicon nanostructures for bioapplications. Nano Today 5, 282–295 (2010).
Chen, K.-I., Li, B.-R. & Chen, Y.-T. Silicon nanowire field-effect transistor-based biosensors for biomedical diagnosis and cellular recording investigation. Nano Today 6, 131–154 (2011).
Zheng, G. F., Patolsky, F., Cui, Y., Wang, W. U. & Lieber, C. M. Multiplexed electrical detection of cancer markers with nanowire sensor arrays. Nat. Biotechnol. 23, 1294–1301 (2005).
Stern, E., Klemic, J. F., Routenberg, D. A., Wyrembak, P. N., Turner-Evans, D. B., Hamilton, A. D., LaVan, D. A., Fahmy, T. M. & Reed, M. A. Label-free immunodetection with CMOS-compatible semiconducting nanowires. Nature 445, 519–522 (2007).
Stern, E., Vacic, A., Rajan, N. K., Criscione, J. M., Park, J., Ilic, B. R., Mooney, D. J., Reed, M. A. & Fahmy, T. M. Label-free biomarker detection from whole blood. Nat. Nanotechnol. 5, 138–142 (2010).
Tian, B. Z., Cohen-Karni, T., Qing, Q., Duan, X. J., Xie, P. & Lieber, C. M. Three-dimensional, flexible nanoscale field-effect transistors as localized bioprobes. Science 329, 830–834 (2010).
Medintz, I. L., Uyeda, H. T., Goldman, E. R. & Mattoussi, H. Quantum dot bioconjugates for imaging, labelling and sensing. Nat. Mater. 4, 435–446 (2005).
Wilson, R. The use of gold nanoparticles in diagnostics and detection. Chem. Soc. Rev. 37, 2028–2045 (2008).
Nam, J.-M., Thaxton, C. S. & Mirkin, C. A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 301, 1884–1886 (2003).
Alivisatos, P. The use of nanocrystals in biological detection. Nat. Biotechnol. 22, 47–52 (2004).
Dekker, C. Solid-state nanopores. Nat. Nanotechnol. 2, 209–215 (2007).
Howorka, S. & Siwy, Z. Nanopore analytics: sensing of single molecules. Chem. Soc. Rev. 38, 2360–2384 (2009).
Soleymani, L., Fang, Z. C., Sargent, E. H. & Kelley, S. O. Programming the detection limits of biosensors through controlled nanostructuring. Nat. Nanotechnol. 4, 844–848 (2009).
Ratinac, K. R., Yang, W. R., Gooding, J. J., Thordarson, P. & Braet, F. Graphene and related materials in electrochemical sensing. Electroanalysis 23, 803–826 (2011).
Ohno, Y., Maehashi, K. & Matsumoto, K. Chemical and biological sensing applications based on graphene field-effect transistors. Biosens. Bioelectron. 26, 1727–1730 (2010).
Huang, X., Yin, Z. Y., Wu, S. X., Qi, X. Y., He, Q. Y., Zhang, Q. C., Yan, Q. Y., Boey, F. & Zhang, H. Graphene-based materials: synthesis, characterization, properties, and applications. Small 7, 1876–1902 (2011).
He, Q. Y., Wu, S. X., Yin, Z. Y. & Zhang, H. Graphene-based electronic sensors. Chem. Sci. 3, 1764–1772 (2012).
Cui, Y., Wei, Q. Q., Park, H. K. & Lieber, C. M. Nanowire nanosensors for highly sensitive and selective detection of biological and chemical species. Science 293, 1289–1292 (2001).
Patolsky, F., Zheng, G. F. & Lieber, C. M. Fabrication of silicon nanowire devices for ultrasensitive, label-free, real-time detection of biological and chemical speciesw. Nat. Protoc. 1, 1711–1724 (2006).
Wang, W. U., Chen, C., Lin, K.-H., Fang, Y. & Lieber, C. M. Label-free detection of small-molecule-protein interactions by using nanowire nanosensors. Proc. Natl. Acad. Sci. USA 102, 3208–3212 (2005).
Patolsky, F., Timko, B. P., Yu, G. H., Fang, Y., Greytak, A. B., Zhang, G. F. & Lieber, C. M. Detection, stimulation, and inhibition of neuronal signals with high-density nanowire transistor arrays. Science 313, 1100–1104 (2006).
Chen, R. J., Bangsaruntip, S., Drouvalakis, K. A., Kam, N. W. S., Shim, M., Li, Y. M., Kim, W., Utz, P. J. & Dai, H. J. Noncovalent functionalization of carbon nanotubes for highly specific electronic biosensors. Proc. Natl. Acad. Sci. USA 100, 4984–4989 (2003).
Besteman, K., Lee, J.-O., Wiertz, F. G. M., Heering, H. A. & Dekker, C. Enzyme-coated carbon nanotubes as single-molecule biosensors. Nano Lett. 3, 727–730 (2003).
Novoselov, K. S., Geim, A. K., Morozov, S. V., Jiang, D., Zhang, Y., Dubonos, S. V., Grigorieva, I. V. & Firsov, A. A. Electric field effect in atomically thin carbon films. Science 306, 666–669 (2004).
Zhang, Y. B., Tan, Y.-W., Stormer, H. L. & Kim, P. Experimental observation of the quantum hall effect and berry’s phase in graphene. Nature 438, 201–204 (2005).
Huang, X., Qi, X. Y., Boey, F. & Zhang, H. Graphene-based composites. Chem. Soc. Rev. 41, 666–686 (2012).
Dai, H. J. Carbon nanotubes: synthesis, integration, and properties. Acc. Chem. Res. 35, 1035–1044 (2002).
Schedin, F., Geim, A. K., Morozov, S. V., Hill, E. W., Blake, P., Katsnelson, M. I. & Novoselov, K. S. Detection of individual gas molecules adsorbed on graphene. Nat. Mater. 6, 652–655 (2007).
Wang, Q., Guo, X. F., Cai, L. C., Cao, Y., Gan, L., Liu, S., Wang, Z. X., Zhang, H. T. & Li, L. D. TiO2-decorated graphenes as efficient photoswitches with high oxygen sensitivity. Chem. Sci. 2, 1860–1864 (2011).
Guo, X. F., Huang, L. M., O’Brien, S., Kim, P. & Nuckolls, C. Directing and sensing changes in molecular conformation on individual carbon nanotube field effect transistors. J. Am. Chem. Soc. 127, 15045–15047 (2005).
Chen, R. J., Choi, H. C., Bangsaruntip, S., Yenilmez, E., Tang, X. W., Wang, Q., Chang, Y.-L. & Dai, H. J. An investigation of the mechanisms of electronic sensing of protein adsorption on carbon nanotube devices. J. Am. Chem. Soc. 126, 1563–1568 (2004).
Tang, X. W., Bansaruntip, S., Nakayama, N., Yenilmez, E., Chang, Y.-L. & Wang, Q. Carbon nanotube DNA sensor and sensing mechanism. Nano Lett. 6, 1632–1636 (2006).
Gui, E. L., Li, L.-J., Zhang, K. K., Xu, Y. P., Dong, X. C., Ho, X. N., Lee, P. S., Kasim, J., Shen, Z. X., Rogers, J. A. & Mhaisalkar, S. G. DNA sensing by field-effect transistors based on networks of carbon nanotubes. J. Am. Chem. Soc. 129, 14427–14432 (2007).
Byon, H. R. & Choi, H. C. Network single-walled carbon nanotube-field effect transistors (SWNT-FETs) with increased schottky contact area for highly sensitive biosensor applications. J. Am. Chem. Soc 128, 2188–2189 (2006).
Heller, I., Chatoor, S., Mannik, J., Zevenbergen, M. A. G., Dekker, C. & Lemay, S. G. Influence of electrolyte composition on liquid-gated carbon nanotube and graphene transistors. J. Am. Chem. Soc. 132, 17149–17156 (2010).
Goldsmith, B. R., Coroneus, J. G., Khalap, K. R., Kane, A. A., Weiss, G. A. & Collins, P. G. Conductance-controlled point functionalization of single-walled carbon nanotubes. Science 315, 77–81 (2007).
Goldsmith, B. R., Coroneus, J. G., Kane, A. A., Weiss, G. A. & Collins, P. G. Monitoring single-molecule reactivity on a carbon nanotube. Nano Lett. 8, 189–194 (2008).
Banerjee, S., Hemraj-Benny, T. & Wong, S. S. Covalent surface chemistry of single-walled carbon nanotubes. Adv. Mater. 17, 17–29 (2005).
Zhao, Y.-L. & Stoddart, J. F. Noncovalent functionalization of single-walled carbon nanotubes. Acc. Chem. Res. 42, 1161–1171 (2009).
Huang, Y. X., Sudibya, H. G., Fu, D. L., Xue, R. H., Dong, X. C., Li, L.-J. & Chen, P. Label-free detection of ATP release from living astrocytes with high temporal resolution using carbon nanotube network. Biosens. Bioelectron. 24, 2716–2720 (2009).
Heller, I., Smaal, W. T. T., Lemay, S. G. & Dekker, C. Probing macrophage activity with carbon-nanotube sensors. Small 5, 2528–2532 (2009).
Sudibya, H. G., Ma, J. M., Dong, X. C., Ng, S., Li, L.-J., Liu, X.-W. & Chen, P. Interfacing glycosylated carbon-nanotube-network devices with living cells to detect dynamic secretion of biomolecules. Angew. Chem. Int. Ed. 48, 2723–2726 (2009).
Wang, C.-W., Pan, C.-Y., Wu, H.-C., Shih, P.-Y., Tsai, C.-C., Liao, K.-T., Lu, L.-L., Hsieh, W.-H., Chen, C.-D. & Chen, Y.-T. In situ detection of chromogranin a released from living neurons with a single-walled carbon-nanotube field-effect transistor. Small 3, 1350–1355 (2007).
Tsai, C.-C., Yang, C.-C., Shih, P.-Y., Wu, C.-S., Chen, C.-D., Pan, C.-Y. & Chen, Y.-T. Exocytosis of a single bovine adrenal chromaffin cell: the electrical and morphological studies. J. Phys. Chem. B 112, 9165–9173 (2008).
Zhou, X. J., Moran-Mirabal, J. M., Craighead, H. G. & McEuen, P. L. Supported lipid bilayer/carbon nanotube hybrids. Nat. Nanotechnol. 2, 185–190 (2007).
Kim, T. H., Lee, S. H., Lee, J., Song, H. S., Oh, E. H., Park, T. H. & Hong, S. Single-carbon-atomic-resolution detection of odorant molecules using a human olfactory receptor-based bioelectronic nose. Adv. Mater. 21, 91–94 (2009).
Vedala, H., Chen, Y. N., Cecioni, S., Imberty, A., Vidal, S. & Star, A. Nanoelectronic detection of lectin-carbohydrate interactions using carbon nanotubes. Nano Lett. 11, 170–175 (2011).
So, H.-M., Park, D.-W., Jeon, E.-K., Kim, Y.-H., Kim, B. S., Lee, C.-K., Choi, S. Y., Kim, S. C., Chang, H. & Lee, J.-O. Detection and titer estimation of Escherichia coli using aptamer-functionalized single-walled carbon-nanotube field-effect transistors. Small 4, 197–201 (2008).
Villamizar, R. A., Maroto, A., Rius, F. X., Inza, I. & Figueras, M. J. Fast detection of salmonella infantis with carbon nanotube field effect transistors. Biosens. Bioelectron. 24, 279–283 (2008).
Star, A., Tu, E., Niemann, J., Gabriel, J.-C. P., Joiner, C. S. & Valcke, C. Label-free detection of DNA hybridization using carbon nanotube network field-effect transistors. Proc. Natl. Acad. Sci. USA 103, 921–926 (2006).
Martinez, M. T., Tseng, Y.-C., Ormategui, N., Loinaz, I., Eritja, R. & Bokor, J. Label-free DNA biosensors based on functionalized carbon nanotube field effect transistors. Nano Lett. 9, 530–536 (2009).
Dong, X. C., Lau, C. M., Lohani, A., Mhaisalkar, S. G., Kasim, J., Shen, Z. X., Ho, X. N., Rogers, J. A. & Li, L.-J. Electrical detection of femtomolar DNA via gold-nanoparticle enhancement in carbon-nanotube-network field-effect transistors. Adv. Mater. 20, 2389–2393 (2008).
Sorgenfrei, S., Chiu, C.-Y., Gonzalez, R. L., Yu, Y.-J., Kim, P., Nuckolls, C. & Shepard, K. L. Label-free single-molecule detection of DNA-hybridization kinetics with a carbon nanotube field-effect transistor. Nat. Nanotechnol. 6, 125–132 (2011).
Guo, X. F., Small, J. P., Klare, J. E., Wang, Y. L., Purewal, M. S., Tam, I. W., Hong, B. H., Caldwell, R., Huang, L. M., O'Brien, S., Yan, J. M., Breslow, R., Wind, S. J., Hone, J., Kim, P. & Nuckolls, C. Covalently bridging gaps in single-walled carbon nanotubes with conducting molecules. Science 311, 356–359 (2006).
Feldman, A. K., Steigerwald, M. L., Guo, X. F. & Nuckolls, C. Molecular electronic devices based on single-walled carbon nanotube electrodes. Acc. Chem. Res. 41, 1731–1741 (2008).
Liu, S., Zhang, X. Y., Luo, W. X., Wang, Z. X., Guo, X. F., Steigerwald, M. L. & Fang, X. H. Single-molecule detection of proteins using aptamer-functionalized molecular electronic devices. Angew. Chem. Int. Ed. 50, 2496–2502 (2011).
Mohanty, N. & Berry, V. Graphene-based single-bacterium resolution biodevice and DNA transistor: interfacing graphene derivatives with nanoscale and microscale biocomponents. Nano Lett. 8, 4469–4476 (2008).
Dong, X. C., Shi, Y. M., Huang, W., Chen, P. & Li, L.-J. Electrical detection of DNA hybridization with single-base specificity using transistors based on CVD-grown graphene sheets. Adv. Mater. 22, 1649–1653 (2010).
Mao, S., Lu, G. H., Yu, K. H., Bo, Z. & Chen, J. H. Specific protein detection using thermally reduced graphene oxide sheet decorated with gold nanoparticle-antibody conjugates. Adv. Mater. 22, 3521–3526 (2010).
Huang, Y. X., Dong, X. C., Shi, Y. M., Li, C. M., Li, L.-J. & Chen, P. Nanoelectronic biosensors based on CVD grown graphene. Nanoscale 2, 1485–1488 (2010).
Waldmann, D., Jobst, J., Speck, F., Seyller, T., Krieger, M. & Weber, H. B. Bottom-gated epitaxial graphene. Nat. Mater. 10, 357–360 (2011).
Ang, P. K., Chen, W., Wee, A. T. S. & Loh, K. P. Solution-gated epitaxial graphene as pH sensor. J. Am. Chem. Soc. 130, 14392–14393 (2008).
Ohno, Y., Maehashi, K., Yamashiro, Y. & Matsumoto, K. Electrolyte-gated graphene field-effect transistors for detecting pH and protein adsorption. Nano Lett. 9, 3318–3322 (2009).
Ohno, Y., Maehashi, K. & Matsumoto, K. Label-free biosensors based on aptamer-modified graphene field-effect transistors. J. Am. Chem. Soc. 132, 18012–18013 (2010).
He, Q. Y., Sudibya, H. G., Yin, Z. Y., Wu, S. X., Li, H., Boey, F., Huang, W., Chen, P. & Zhang, H. Centimeter-long and large-scale micropatterns of reduced graphene oxide films: fabrication and sensing applications. ACS Nano 4, 3201–3208 (2010).
Sudibya, H. G., He, Q. Y., Zhang, H. & Chen, P. Electrical detection of metal ions using filed-effect transistors based on micropatterned reduced graphene oxide films. ACS Nano 5, 1990–1994 (2011).
Dankerl, M., Hauf, M. V., Lippert, A., Hess, L. H., Birner, S., Sharp, I. D., Mahmood, A., Mallet, P., Veuillen, J.-Y., Stutzmann, M. & Garrido, J. A. Graphene solution-gated field-effect transistor array for sensing applications. Adv. Funct. Mater. 20, 3117–3124 (2010).
Cohen-Karni, T., Qing, Q., Li, Q., Fang, Y. & Lieber, C. M. Graphene and nanowire transistors for cellular interfaces and electrical recording. Nano Lett. 10, 1098–1102 (2010).
Hess, L. H., Jansen, M., Maybeck, V., Hauf, M. V., Seifert, M., Stutzmann, M., Sharp, I. D., Offenhausser, A. & Garrido, J. A. Graphene transistor arrays for recording action potentials from electrogenic cells. Adv. Mater. 23, 5045–5049 (2011).
Cao, X. H., He, Q. Y., Shi, W. H., Li, B., Zeng, Z. Y., Shi, Y. M., Yan, Q. Y. & Zhang, H. Graphene oxide as a carbon source for controlled growth of carbon nanowires. Small 7, 1199–1202 (2011).
Yin, Z. Y., He, Q. Y., Huang, X., Zhang, J., Wu, S. X., Chen, P., Lu, G., Chen, P., Zhang, Q. C., Yan, Q. Y., Zhang, H. & Real-time, D. N. A. detection using Pt nanoparticle-decorated reduced graphene oxide field-effect transistors. Nanoscale 4, 293–297 (2012).
Agarwal, S., Zhou, X. Z., Ye, F., He, Q. Y., Chen, G. C. K., Soo, J., Boey, F., Zhang, H. & Chen, P. Interfacing live cells with nanocarbon substrates. Langmuir 26, 2244–2247 (2010).
Acknowledgements
We acknowledge primary financial support from MOST (2009CB623703 and 2012CB921404), NSFC (20833001, 51121091, 2112016 and 21003002), FANEDD (2007B21), 111 Project (B08001) and BSTSP (2009A01). We also acknowledge Michael Steigerwald (Columbia University) for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Liu, S., Guo, X. Carbon nanomaterials field-effect-transistor-based biosensors. NPG Asia Mater 4, e23 (2012). https://doi.org/10.1038/am.2012.42
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/am.2012.42
Keywords
This article is cited by
-
Electrochemical field-effect bio-transistor based on a multi-scale electronic nanomesh of single-walled carbon nanotubes
Carbon Letters (2023)
-
Recent Progress in Rapid Biosensor Fabrication Methods: Focus on Electrical Potential Application
BioChip Journal (2023)
-
Hetero Channel Double Gate MOSFET for Label-free Biosensing Application
Silicon (2022)
-
pH sensor based on AuNPs/ ITO membrane as extended gate field-effect transistor
Applied Physics B (2022)
-
Stretchable transistors and functional circuits for human-integrated electronics
Nature Electronics (2021)