Abstract
CRISPR–Cas9 is poised to become the gene editing tool of choice in clinical contexts. Thus far, exploration of Cas9-induced genetic alterations has been limited to the immediate vicinity of the target site and distal off-target sequences, leading to the conclusion that CRISPR–Cas9 was reasonably specific. Here we report significant on-target mutagenesis, such as large deletions and more complex genomic rearrangements at the targeted sites in mouse embryonic stem cells, mouse hematopoietic progenitors and a human differentiated cell line. Using long-read sequencing and long-range PCR genotyping, we show that DNA breaks introduced by single-guide RNA/Cas9 frequently resolved into deletions extending over many kilobases. Furthermore, lesions distal to the cut site and crossover events were identified. The observed genomic damage in mitotically active cells caused by CRISPR–Cas9 editing may have pathogenic consequences.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
31 July 2018
In the version of this article initially published online, four figure citations were incorrect on p.2: left-hand column, after "complex rearrangements," "Supplementary Fig. 2a,b" should have been "Fig. 2a,b"; right-hand column, in three places, the citation for "Supplementary Fig. 3..." should have been for "Supplementary Fig. 2." The errors have been corrected for the print, PDF and HTML versions of this article.
01 October 2018
Nat. Biotechnol. 10.1038/nbt.4192; corrected online 31 July 2018 In the version of this article initially published online, four figure citations were incorrect on p.2: left-hand column, after “complex rearrangements,” “Supplementary Fig. 2a,b” should have been “Fig. 2a,b”; right-hand column, in three places, the citation for “Supplementary Fig.
References
Cornu, T.I., Mussolino, C. & Cathomen, T. Refining strategies to translate genome editing to the clinic. Nat. Med. 23, 415–423 (2017).
Kim, S., Kim, D., Cho, S.W., Kim, J. & Kim, J.-S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Komor, A.C., Kim, Y.B., Packer, M.S., Zuris, J.A. & Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Frock, R.L. et al. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 33, 179–186 (2015).
Xie, F. et al. Seamless gene correction of β-thalassemia mutations in patient-specific iPSCs using CRISPR/Cas9 and piggyBac. Genome Res. 24, 1526–1533 (2014).
Guilinger, J.P., Thompson, D.B. & Liu, D.R. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat. Biotechnol. 32, 577–582 (2014).
Kleinstiver, B.P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Tsai, S.Q. et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 32, 569–576 (2014).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Koike-Yusa, H., Li, Y., Tan, E.-P. & Velasco-Herrera, M.D.C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
van Overbeek, M. et al. DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks. Mol. Cell 63, 633–646 (2016).
Tan, E.P., Li, Y., Velasco-Herrera, M.D.C., Yusa, K. & Bradley, A. Off-target assessment of CRISPR-Cas9 guiding RNAs in human iPS and mouse ES cells. Genesis 53, 225–236 (2015).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Ghezraoui, H. et al. Chromosomal translocations in human cells are generated by canonical nonhomologous end-joining. Mol. Cell 55, 829–842 (2014).
Weinstock, D.M., Elliott, B. & Jasin, M. A model of oncogenic rearrangements: differences between chromosomal translocation mechanisms and simple double-strand break repair. Blood 107, 777–780 (2006).
Canver, M.C. et al. Characterization of genomic deletion efficiency mediated by CRISPR/Cas9 in mammalian cells. J. Biol. Chem. 289, 21312–21324 (2014).
Kraft, K. et al. Deletions, inversions, duplications: engineering of structural variants using CRISPR/Cas in mice. Cell Rep. 10, 833–839 (2015).
Boroviak, K., Doe, B., Banerjee, R., Yang, F. & Bradley, A. Chromosome engineering in zygotes with CRISPR/Cas9. Genesis 54, 78–85 (2016).
Boroviak, K., Fu, B., Yang, F., Doe, B. & Bradley, A. Revealing hidden complexities of genomic rearrangements generated with Cas9. Sci. Rep. 7, 12867 (2017).
Parikh, B.A., Beckman, D.L., Patel, S.J., White, J.M. & Yokoyama, W.M. Detailed phenotypic and molecular analyses of genetically modified mice generated by CRISPR-Cas9-mediated editing. PLoS One 10, e0116484 (2015).
Shin, H.Y. et al. CRISPR/Cas9 targeting events cause complex deletions and insertions at 17 sites in the mouse genome. Nat. Commun. 8, 15464 (2017).
Gasperini, M. et al. CRISPR/Cas9-mediated scanning for regulatory elements required for HPRT1 expression via thousands of large, programmed genomic deletions. Am. J. Hum. Genet. 101, 192–205 (2017).
Roberts, S.A. et al. Clustered mutations in yeast and in human cancers can arise from damaged long single-strand DNA regions. Mol. Cell 46, 424–435 (2012).
Sinha, S. et al. Microhomology-mediated end joining induces hypermutagenesis at breakpoint junctions. PLoS Genet. 13, e1006714 (2017).
Yang, Y., Sterling, J., Storici, F., Resnick, M.A. & Gordenin, D.A. Hypermutability of damaged single-strand DNA formed at double-strand breaks and uncapped telomeres in yeast Saccharomyces cerevisiae. PLoS Genet. 4, e1000264 (2008).
Tichy, E.D. et al. Mouse embryonic stem cells, but not somatic cells, predominantly use homologous recombination to repair double-strand DNA breaks. Stem Cells Dev. 19, 1699–1711 (2010).
Hacein-Bey-Abina, S. et al. A serious adverse event after successful gene therapy for X-linked severe combined immunodeficiency. N. Engl. J. Med. 348, 255–256 (2003).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Strogantsev, R. et al. Allele-specific binding of ZFP57 in the epigenetic regulation of imprinted and non-imprinted monoallelic expression. Genome Biol. 16, 112 (2015).
Pettitt, S.J. et al. Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat. Methods 6, 493–495 (2009).
Yusa, K., Zhou, L., Li, M.A., Bradley, A. & Craig, N.L. A hyperactive piggyBac transposase for mammalian applications. Proc. Natl. Acad. Sci. USA 108, 1531–1536 (2011).
Moreno-Mateos, M.A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Hill, J.T. et al. Poly peak parser: method and software for identification of unknown indels using sanger sequencing of polymerase chain reaction products. Dev. Dyn. 243, 1632–1636 (2014).
Platt, R.J. et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Acknowledgements
We wish to thank M. Friedrich for sharing his gRNA expression construct and technical advice, E. Metzakopian for technical advice and critical reading of the manuscript, G. Rutledge for critical reading of the early manuscript, A. Ferguson-Smith for the CAST/B6 hybrid ES cells, P. Liu and X. Gao for mCherry/GFP reporter cells, S. Jackson's group for the Cas9-expressing RPE1 cell line and the Cytometry Core Facility for assistance with cell sorting. This work was supported by the Wellcome Trust Grant number 098051.
Author information
Authors and Affiliations
Contributions
M.K. performed most of the experiments and analyzed the data. K.T. performed the primary cell work. A.B. supervised the project. All authors contributed to writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Frequency of PigA loss upon editing with RNP in mouse ES cells and structure of the recovered alleles.
(A) Frequency of PigA loss caused by Cas9 with intronic and exonic gRNAs. Each experiment was conducted in biological triplicate and technical duplicate. (B) Recovered alleles from the “Neon electroporation” experiment. Display conventions as in Fig. 4. Alleles are sorted by total length. See Table S2 for detailed descriptions.
Supplementary Figure 2 Examples of insertion alleles at the PigA locus.
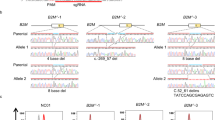
Panels, A, B and C are illustrations of different alleles. The bottom of each panel is a representation of the PigA reference allele around exon 2, the top of each panel shows the structure of the sequenced allele. Black horizontal line – direct reference match, orange bar – inversion, blue bar – insertion from another part of the genome, black arrowhead – gRNA target site, violet bar – duplication, black vertical bar – small indel within the insertion. Grey and orange shadows represent, respectively, direct and inverted match between the reference and the sequenced allele. Lack of shadow at the reference locus represents a deletion in the sequenced allele.
Supplementary Figure 3 Structure of recovered alleles from PigA-deficient, mouse ES cell clones.
(A) Positions of primer pairs used to screen PigA-deficient clones. In grey, primers used for diagnostic PCRs on alleles that could not be recovered. Other display conventions as in Fig. 3. (B-D) Recovered alleles. (B) 5' intronic guide, (C) 3' intronic guide, (D) exonic guide. Display conventions as in Fig. 4. Alleles are sorted by total length.
Supplementary Figure 4 Analysis of the diversity of editing outcomes at PigA locus.
PigA locus was edited using 5’ intronic guide (#15) in biological quadruplicate. Duplicate PCR reactions spanning the cut site performed on DNA extracted from the bulk of PigA negative cells and resolved on an agarose gel (product size 3500bp, 1F/1R primer pair, Fig. S2A). (A) Experimental considerations. (B) Deletion fingerprints. Cell line: JM8 – the original mouse embryonic stem cell line (transfected with gRNA and Cas9 PiggyBac vectors); CAS – subclone of JM8 expressing Cas9 from a single-copy lentiviral transgene (transfected with only gRNA PiggyBac vector). Ladder scale is in kilobases.
Supplementary Figure 5 Genotyping of GFP-deficient progenitor cells.
(A) Positions of primer pairs and the gRNA. Display conventions as in Fig. 3A. (B) Agarose gel pictures. Asterisks indicate clones with at least one deletion allele. Red asterisks indicate eight clones, whose deletion alleles were verified by Sanger sequencing (Fig. S6A). Ladder scale is in kilobases.
Supplementary Figure 6 Structure of recovered deletion alleles from GFP-deficient clones and gating strategy for Cd9 cell sorting.
(A) Display conventions as in Fig. S3. (B) Cells were edited with various intronic and exonic gRNAs, sorted for “low”, “medium” and “high” classes and single cell cloned (Table S5; sorting was performed twice for guides #1 and #86 and once for guides #35 and #80; flow cytometric analysis was replicated independently 4-7 times, see also Table S1).
Supplementary Figure 7 Overlap between unique PigA alleles derived using different methods.
“Single cell” and “PacBio” refer to alleles shown in Fig. S2B-D and Fig. 2A. “Plasmid” alleles were derived in the same experiment by Sanger sequencing of 3.5kb long PCR fragments cloned into plasmid vectors (primer pair 1F/1R, Fig. S2A). “RNP” alleles were derived by single cell cloning in an independent experiment only using guide #15 (Fig. S1A “Electroporation exp2”; Fig. S1B).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1632 kb)
Supplementary Table 1
Flow cytometry results and gRNA sequences. (XLSX 8 kb)
Supplementary Table 2
Automatic annotation of all PigA alleles in the study (XLSX 35 kb)
Supplementary Table 3
Detailed description of PigA alleles recovered from single cell clones. (XLSX 15 kb)
Supplementary Table 4
Diagnostic PCRs at the PigA locus. (XLSX 5 kb)
Supplementary Table 5
Summary of PCR genotyping experiments (XLSX 7 kb)
Supplementary Table 6
PCR primers. (XLSX 7 kb)
Supplementary Data 1
PigA alleles (TXT 820 kb)
Supplementary Data 2
Barcodes (ZIP 1 kb)
Supplementary Note
Supplementary Note 1 (PDF 169 kb)
Rights and permissions
About this article
Cite this article
Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat Biotechnol 36, 765–771 (2018). https://doi.org/10.1038/nbt.4192
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4192
This article is cited by
-
Improving CRISPR–Cas9 directed faithful transgene integration outcomes by reducing unwanted random DNA integration
Journal of Biomedical Science (2024)
-
An integrated toolkit for human microglia functional genomics
Stem Cell Research & Therapy (2024)
-
Novel insights into TCR-T cell therapy in solid neoplasms: optimizing adoptive immunotherapy
Experimental Hematology & Oncology (2024)
-
Decoding the complexity of on-target integration: characterizing DNA insertions at the CRISPR-Cas9 targeted locus using nanopore sequencing
BMC Genomics (2024)
-
HiHo-AID2: boosting homozygous knock-in efficiency enables robust generation of human auxin-inducible degron cells
Genome Biology (2024)