Abstract
Previous studies have shown that individuals with a deletion of 15q26.1→qter, which includes the insulin-like growth factor I receptor (IGFIR) gene, may exhibit phenotypic characteristics similar to those individuals with Silver-Russell Syndrome (SRS). Thirty-three SRS probands, with normal karyotypes, and their parents were investigated for the presence of both copies of IGFIR by gene dosage analysis of Southern blot hybridisation. All 33 SRS probands have both copies of IGFIR. Tetranucleotide repeat marker analysis for three locations on 15q also ruled out other deletions in these regions for those markers that were informative. Two important functional regions of IGFIR were also investigated for DNA mutations, using single-stranded conformational polymorphism analysis. No mutations were found in the cysteine-rich region involved in ligand binding (exon 3) or the ATP binding region (exon 16) which could contribute to the SRS phenotype. However, a silent mutation in the third position of one of the codons in the ATP region (3174 G→A, 1013 Glu→Glu) was found.
Similar content being viewed by others
Introduction
Silver-Russell syndrome (SRS) is a clinical disorder where intrauterine growth retardation (IUGR) and poor postnatal growth lead to short adult stature [1–3]. In addition there is often a characteristic triangular facies, skeletal asymmetry and digital anomalies, including clinodactyly. Recently, evidence of a cognitive deficit has also been associated with SRS [4].
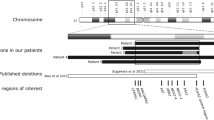
Although detailed clinical reports for SRS are available, there is still little known about its aetiology. No consistent Mendelian or chromosomal basis of inheritance has been established for SRS and most cases are sporadic [5]. The most commonly reported aetiological finding involves either deletion for distal 15q or ring chromosome 15, with 18 documented cases [6–11]. Patients with ring chromosome 15 have IUGR, microcephaly, triangular face, hypertelorism, variable mental retardation and speech delay [8, 12]. Of primary interest for SRS is the insulin-like growth factor I receptor (IGFIR) gene, localised to 15q25–26. The r(15) patients exhibit hemizygosity for 15q distal markers at 15q26.3, 15q26.2 and/or 15q26.1 and hemizygosity for IGFIR [12, 13]. These findings are consistent with an SRS patient with a terminal deletion of 15q26.1→qter, with hemizygosity for IGFIR [8]. However, 5 SRS patients who were diploid for distal 15q markers have also been reported suggesting that the loss of one copy of IGFIR may not contribute to the clinical manifestations in all cases of SRS [11].
In a cohort of 37 patients, there were 3 documented cases of SRS associated with maternal uniparental disomy (UPD) of chromosome 7 (mUPD 7) [14]. MUPD 7 has also been associated in 4 out of 35 patients with SRS or primordial dwarfism from several centres in Europe [15]. For the remaining SRS patients with copies of chromosome 7 from both parents it still remains possible that chromosome 7 may be implicated. Smaller deletions in imprinted regions may be involved, as has been found for many of the cases in Prader-Willi and Angelman syndrome, where mUPD 15 and pUPD 15 play only a small part (25 and 2% respectively) [16–18]. There are an additional 6 SRS individuals with other documented structural chromosomal abnormalities [19–24]. However, this still leaves a large majority of the patients with unexplained aetiology.
The insulin-like growth factor family is composed of insulin, IGFI and IGFII (insulin-like growth factors I and II respectively), their corresponding receptors and at least six binding proteins [25]. The ligands, receptors and binding proteins play a pivotal role in the regulation of growth and development, in both fetal and postnatal life [26, 27].
IGFIR mediates the action for both IGFI and IGFII, but having a much higher affinity for IGFI. IGFII and IGFIIR are expressed at the two-cell stage while the insulin receptor and IGFIR have been detected at the eight-cell stage in preimplantation mouse and human embryos [28, 29]. IGFIR is widely expressed after implantation especially in the developing nervous system and muscle [30], but decreases dramatically during postnatal development [31].
It is interesting to note that mice knockout experiments have shown that hemizygosity at Igf1r locus does not have any effect on growth in mice, although severe growth retardation (55% reduction in size compared to wild type mice), as well as developmental delay in ossification, CNS abnormalities and hypoplasia is observed for homozygous null mice [32, 33]. Both Igf1 and Igf2 have been shown to utilise Igf1r in early embryonic development, and the complete absence of the receptor would severely compromise the functioning of these ligands [33, 34].
Normal phenotype in mice hemizygous for Igf1r points to species difference in compensatory mechanisms for gene hemizygosity, suggesting that there may be up-regulation of the remaining Igf1r, which does not appear to be the case in humans. SRS is not a constant phenotype in deletion of 15qter, pointing to a heterogeneous disorder. Hemizygosity of IGFIR may be only a part of IUGR/SRS in deletion of 15qter as the loss of one copy of IGFIR was accompanied by loss of flanking chromosomal material, which may include other functional genes that play a role in growth and development. However, based on its pivotal role in embryonic and post-natal growth and differentiation, it is an ideal candidate for any growth retardation phenotype, and hence SRS.
Here we report findings on 33 SRS probands. These are a subset of the 37 patients reported by Preece et al. [14]; 4 were excluded from this report due to lack of DNA/blood. Since the studies were conducted in parallel, the 3 probands subsequently shown to have mUPD 7, were part of this investigation. Hemizygosity for IGFIR was investigated by quantitative analysis of Southern hybridisation. Tetranucleotide markers localised to the distal portion of 15q, were used to support Southern hybridisation data. Finally, SSCP analysis was undertaken to screen for any point mutations in two exons known to be critical to IGFIR function.
Materials and Methods
Subjects
33 SRS probands and both parents were included in this study. 28 probands fulfilled at least three of the following diagnostic criteria: low birth weight (>2 SDS below mean); short stature at the time of diagnosis (>2 SDS below mean; the range of heights is not given here, as at the time of this investigation, a number of the probands had undergone growth hormone treatment and hence were of almost normal height); characteristic facial features; and facial, trunk or limb asymmetry. The remaining 5 probands had consistent postnatal growth pattern and facial features but slightly higher birth weights (2.58–3.11 kg). Blood was obtained from the families and genomic DNA isolated [35]. Ethical approval was obtained for this study by the Joint Research Ethics Committee of Great Ormond Street Hospital and the Institute of Child Health (approval No. 1278).
The cohort consisted of 16 females and 17 males, ranging in age between 0.83 and 34.3 years at the time of investigation. Classical facial features, as described by Russell [1], were seen in 22, whilst 11 had a milder facial phenotype. Limb asymmetry of ≥ 1.0 cm associated with facial asymmetry were present in 12 individuals, and clinodactyly was seen in 24 individuals. At the time of birth of the probands, mean maternal age was 27.8 (range 17.8–37.8) years and mean paternal age was 30.6 (range 18.8–44.8) years.
Cytogenetic Studies
In probands karyotype was normal (performed at the North East Thames Regional Cytogenetic Centre).
Southern Blot Hybridisation
4 µg of total genomic DNA were digested with 24 u of HindIII (GIBCO BRL) for 6 h at 37°C and electrophoresed on 0.8% agarose gels overnight. Southern blotting and hybridisation were carried out by standard methods [36].
Filters were simultaneously probed with a 0.7-kb cDNA EcoRI fragment of the human IGF1R (IGF-1-R.8; American Type Culture Collection, Rockville, Md.) and W21G, a 1.6 kb-HindIII fragment located on 22q11.1–q11.2 (HGMP UK DNA Probe Bank).
Quantitative Analysis of Southern Hybridisation
For each filter, incorporation of radioactivity in each band was measured for each individual by volumetric analysis using a Phosphorlmager (Model 400; Molecular Dynamics). If the average reading for IGF1R and W21G of all parents on a filter are represented as IC and WC respectively, and the value of IGF1R and W21G for each proband represented as IP and WP respectively, then the ratio of IGF1R to W21G for the proband was calculated using the following formula:
A ratio of 1.0 indicates that for each copy of IGF1R there is one copy of W21G and hence diploid number of the receptor, whereas a ratio of 0.5 indicates that there is only one IGF1R copy for two W21G copies and hence hemizygosity of IGF1R. For each proband two readings were obtained. Since there were no significant differences in the readings obtained from the first nine families using duplicate filters, and then reprobed filters, all subsequent readings were obtained by stripping filters in boiling 0.1 % SDS for 30 min and reprobing as described, and the average of two readings taken.
Tetranucleotide Marker Analysis
Reaction conditions for the three sets of primers are given below:
(94° C, 4 min) × 1: (94° C, 1 min; AT° C, 1 min; 72° C, 1 min) × 25: (72° C, 10 min) × 1, where AT is the annealing temperature for D15S642 (66° C), D15S657 (63° C) and D15S816 (59° C).
6 µl of PCR reaction were denatured and electrophoresed on 6% denaturing Polyacrylamide gel (National Diagnostics).
Single-Stranded Conformational Polymorphism Analysis
Radiolabelled PCR reactions for single-stranded conformational polymorphism (SSCP) analysis were carried out as described above. SSCP was used to study two regions. Primer sequences for exon 3 (bp 685–996) (CF and CR) and for exon 16 (bp 3001–3228) (AF and AR), and their reaction conditions are shown below (exon sequences are shown in bold, and intron sequences are shown in normal type) [38, 39]:
CF 5′-CTCTCCACAGTGTGCCCAAG-3′
CR 5′-ATACCTCTGGCTGCCGTTGC-3′
(94°C,4 min) × 1: (94° C, 1 min; 65° C, 1 min; 72° C, 1 min) × 30:(72°C, 10 min) × 1
AF 5′-TCTTCTCCAGTGTACGTTCC-3′
AR 5′-GGAACTTTCTCTTACCACATG-3′
(94° C, 4 min) × 1: (94° C, 1 min; 57° C, 1 min; 72° C, 1 min) × 30:(72° C, 10 min) × 1
7 µl of the PCR reactions were denatured and electrophoresed on 6% nondenaturing Polyacrylamide gel (easigel, Scotlab), below 10° C for 16–20 h at 280–320 V.
Sequencing
Sequencing was carried out on a model 373 ABI sequencer.
Results
Quantitative Analysis of Southern Hybridisation
Filters containing HindIII digests of proband and parent genomic DNA were hybridised simultaneously with IGFIR and W21G probes, giving two distinct bands of approximately 1.7 and 3.5 kb respectively (fig. 1). The ratio of IGFIR to W21G for the probands are summarised in figure 2, showing that the ratios of IGFIR to W21G follows a normal distribution with ratios close to one. The range of average readings obtained was 0.84–1.28, with a mean of 1.04 (SD 0.10). These data indicate no evidence of hemizygosity of IGFIR in this group of 33 SRS probands.
Southern hybridisation of four families. Total genomic DNA was digested with HindIII and electrophoresed on 0.8% agarose gel and blotted. Filters were probed simultaneously with IGFIR (1.7 kb, lower band) and W21G (3.5 kb, upper band) and exposed to X-ray film overnight. Order: mother, proband and father.
Tetranucleotide Marker Analysis
All 33 families were informative for at least one of the tetranucleotide markers, although only one family was informative for all the markers. In all cases where the markers were informative, Mendelian inheritance was demonstrated for the families, supporting no deletion of the region around IGFIR (fig. 3).
SSCP Analysis of the Cysteine-Rich Region
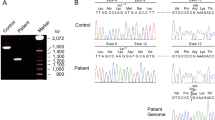
PCR amplification of the cysteine-rich region gave a band of 327 bp which corresponds to the 312-bp exon 3 plus flanking intron nucleotides incorporated into the primers [38, 39].
The three families in figure 4 are representative of all 33 probands’ and their parents’ banding pattern for this region. The lack of any band shift in the PCR products indicates no mutations in this region [40].
SSCP analysis of exon 3 of the IGFIR gene. Lane 1, non-denatured control sample; lanes 2–10, 3 of 33 families investigated, order mother, proband and father. The upper two bands represent the two denatured alleles for this region and the third lowest band is an alternative conformation of one of these alleles. Note that all individuals have identical banding patterns.
SSCP Analysis of the ATP Region
A band of 255 bp was observed corresponding to the 228-bp exon 16 and flanking intron sequence incorporated into the primer design. 28 of the 33 families had identical banding patterns [38, 39].
Band shifts were observed for SSCP in this region, where individuals in five families had one or two bands (fig. 5). Sequence analysis showed that individuals with single bands were of two types. The first group were homozygous G at position 3174, in agreement with the published sequence [38, 39]. The second group of individuals with a single band had a base transition (G → A) at this position, and were homozygous for this change. Individuals with two bands were heterozygous; the transition was in the third base of a codon, GAG → GAA, where both code for glutamine.
Discussion
Previous reports for IGFIR hemizygosity as a possible aetiological cause for SRS have all been supported by visible cytogenetic deletions [6–11]. There is no report of SRS individuals with normal karyotype who are hemizygous for IGFIR. In this investigation, the largest to date for hemizygosity of IGFIR, both chromosomes 15 were cytogenetically normal. This did not rule out the possibility that IGFIR was deleted, as conventional karyotyping methods have a range limit of 1–2 Mb and IGFIR covers only 100 kb [38]. Quantitative analysis of Southern hybridisation data together with polymorphic PCR markers, point to the presence of two copies of IGFIR for all 33 SRS probands, thereby ruling out the possibility that hemizygosity at this locus could contribute to the observed phenotype in this group.
IGFIR is a heterotetrameric, transmembrane glycoprotein, composed of two α- and two β-subunits linked by disulfide bonds. IGFIR shows a high degree of primary and secondary structural similarity to the insulin receptor. Ligand binding in the cysteine-rich domain [41] of the α-subunits at the extracellular surface stimulates intracellular, tyrosine-specific protein kinase activity which leads to β-subunit autophosphorylation and subsequent phosphorylation of cytoplasmic components of an IGF-I-specific signal transduction cascade [38, 42, 43].
In the two exons examined by SSCP, exon 3 and 16, no mutation was detected that would play a role in disabling IGFIR. Exon 3 is the cysteine-rich domain involved in ligand binding. It has been shown that IGFI, IGFII and at a much lower affinity, insulin, bind to this region, and the specificity of ligand binding is not only determined by the number of cysteine residues but also their distribution in the protein binding region and the flanking amino acids [41]. It was proposed that a mutation that interfered with ligand recognition would impair IGFIR functioning, so that only the non-mutated allele would be capable of ligand binding and hence activation of the tyrosine kinase domain. However, no mutations were detected.
The second region that was investigated using SSCP was exon 16, located in the tyrosine kinase domain. This is the most highly conserved region compared to the insulin receptor (80–95%; [39]) highlighting its functional importance. There are four potential ATP binding sites, Gly 976, 978 and 981 and Lys 1003, in the tyrosine kinase domain [44]. Disruption of any of these binding sites would effectively down-regulate subsequent phosphorylation of other target proteins normally receptive to IGFI/IGFII mediated signalling via IGFIR. It is unlikely that the novel mutation found at nt 3174 (G→A) has phenotypic implications. Since there is no change in the amino acid and hence the protein structure, the transition does not alter the function of this region. This is the first time that a polymorphism, showing Mendelian inheritance, has been described in the coding region of IGFIR.
Despite the lack of any evidence for hemizygosity of IGFIR being involved in a possible aetiology for SRS in our cohort of 33 individuals, it does not suggest that it should be completely excluded as a candidate gene. Other well documented cases with deletion of one copy of IGFIR show evidence of IUGR, and in a subset, the SRS phenotype. Since this receptor plays such a pivotal role in early embryonic development, it is important to include it in any investigation concerned with prenatal and postnatal growth retardation, in addition to other candidate genes involved in cell growth, proliferation, and development. Although no smaller deletions or base mutations were found in IGFIR for these individuals, only two out of the total of 21 exons encoding the functional receptor were investigated by SSCP, and mutation in the other exons could also affect receptor functioning.
References
Russell A: A syndrome of intrauterine dwarfism recognisable at birth with craniofacial dysostosis, disproportionate short arms and other anomalies. Proc R Soc Med 1954;47:1040–1044.
Silver H, Kiyasu W, George J, Deamer W: Syndrome of congenital hemihypertrophy, shortness of stature and elevated gonadotropins. Pediatrics 1953;12:356–368.
Silver HK: Silver syndrome; in Buyse ML (ed): Birth Defects Encyclopedia. Dover, Centre for Birth Defects Information Services, 1990, pp 1535–1538.
Lai KYC, Skuse D, Stanhope R, Hindmarsh PC: Cognitive abilities associated with the Silver-Russell syndrome. Arch Dis Child 1994;71: 490–496.
McKusick VA: Mendelian Inheritance in Man, ed 10. Baltimore. John Hopkins University Press, 1990, pp 1696–1697.
Kristofferson V, Heim S, Mandahl N, Sundkvist L, Szelest J, Hagerstrand I: Monosomy and trisomy of 15q24 → qter in a family with a translocation t(6; 15)(p25; q24). Clin Genet 1987;32:169–171.
Francke U, Darras BT, Foellmer BE: Loss of IGF-I receptor gene in patients with ring chromosome 15 is related to Russell-Silver like phenotype. Proc 9th Annu-David W. Smith Workshop Malformations Morphogenesis, Oakland, August 1988.
Roback EW, Barakat AJ, Dev VG, Mbikay M, Chretien M, Butler MG: An infant with deletion of the distal long arm of chromosome 15 (q26.1-qter) and loss of insulin-like growth factor I receptor gene. Am J Med Genet 1991;38: 74–79.
Tamura T, Tohma T, Ohta T, Soejima H, Harada N, Abe K, Nikawa N: Ring chromosome 15 involving deletion of the insulin-like growth factor receptor gene in a patient with features of Silver-Russell syndrome. Clin Dysmorphol 1993;2:106–113.
Peoples R, Milatovich A, Francke U: Hemizygosity at the insulin-like growth factor 1 receptor (IGFIR) locus and growth failure in the ring chromosome 15 syndrome. Cytogenet Cell Genet 1995;70:228–234.
Rogan PK, Seip JR, Driscoll DJ, Papenhausen PR, Johnson VP, Raskin S, Woodward AL, Butler MG: Distinct 15q genotypes in Russell-Silver and ring 15 syndromes. Am J Med Genet 1996;62:10–15.
Butler MG, Fogo B, Fuchs DA, Collins FS, Dev VG, Phillips JA: Two patients with ring chromosome 15 syndrome. Am J Med Genet 1988; 9:149–154.
Wilson GN, Sauder SE, Bush M, Beitins IZ: Phenotypic delineation of ring chromosome 15 and Russell-Silver syndromes. J Med Genet 1985;22:233–236.
Preece MA, Price S, Davies V, Gough L, Stanier P, Trembath R, Moore GE: Maternal uniparental disomy 7 in the Silver-Russell syndrome. J Med Genet 1997;34(1):6–9.
Kotzot D, Schmitt S, Bernasconi F, Robinson W, Lurie I, Llynia H, Mehes K, Hamel BC, Otten BJ, Hergesberg M: Uniparental disomy 7 in Silver-Russell syndrome and primordial growth retardation. Hum Mol Genet 1995;4: 583–587.
Greger V, Woolf E, Lalande M: Cloning of the breakpoints of a submicroscopic deletion in an Angelman syndrome patient. Hum Mol Genet 1993;2:921–924.
Buiting K, Saitoh S, Gross S, Dittrich B, Schwartz S, Nicholls RD, Horsthemke B: Inherited microdeletions in the Angelman and Prader-Willi syndromes define an inprinting centre on human chromosome 15. Nat Genet 1995;9:395–400.
Erdel M, Schuffenhauer S, Buchholz B, Bart-Witte U, Kochl S, Utermann B, Duba HC, Utermann G: Routine screening for microdeletions by FISH in 77 patients suspected of having Prader-Willi or Angelman syndromes using YAC clone 273A2 (D15S10). Hum Genet 1996;97:784–793.
Chauvel PJ, Moore CM, Haslam RHA: Trisomy-18 mosaicism with features of Russell-Silver syndrome. Dev Med Child Neurol 1975; 17(2):220–224.
Christensen MF, Nielsen J: Deletion short arm 18 and Silver-Russell syndrome. Acta Paediatr Scand 1978;67:101–103.
Graham JM Jr, Hoehn H, Lin MS, Smith DW: Diploid-triploid mixoploidy: Clinical and cytogenetic aspects. Paediatrics 1981;68:23–28.
Ramirez-Duenas ML, Medina C, Ocampo-Campos R, Rivera H: Severe Silver-Russell syndrome and translocation (17;20)(q25;q13). Clin Genet 1992;41:51–53.
Midro AT, Debek K, Sawicka A: Second observation of Silver-Russell syndrome in a carrier of a reciprocal translocation with one breakpoint at site 17q25 (letter). Clin Genet 1993;44: 53–55.
Schinzel AA, Robinson WP, Binkert F, Fanconi A: An interstitial deletion of proximal 8q (q11–q13) in a girl with Silver-Russell syndrome-like features. Clin Dysmorphol 1994;3: 63–69.
Jones JI, Clemmons DR: Insulin-like growth factors and their binding proteins: Biological actions. Endocr Rev 1995;16(1):3–34.
Caiola S, Di-Biase N, Buongiorno AM: Somatomedine/fattori di crescita insolino-simili (IGFs): Caratteristiche chimiche e funzionali. Ann Ist Super Sanita 1992;28:553–561.
Milner RD, Hill DJ: Fetal growth control: The role of insulin and related peptides. Clin Endocrinol 1984;21:415–433.
Kaye PL, Bell KL, Beeb LF, Dunglison GF, Gardner HG, Harvey MB: Insulin and the insulin-like growth factors (IGFs) in preimplantation development. Reprod Fertil Dev 1992;4:353–386.
Lighten AD, Hardy K, Winston RML, Moore GE: Expression of the insulin-like growth factors and their receptors in human pre-implantation embryos. Mol Reprod Dev, in press.
Bondy CA, Werner H, Roberts CT JR, LeRoith D: Cellular patterns of insulin-like growth factor I (IGF-I) and type I IGF receptor gene expression in early organogenesis: Comparison with IGF-II expression. Mol Endocrinol 1990; 4:1386–1398.
Werner H, Woloschak M, Adamo M, Shen-Orr Z, Roberts CT Jr, LeRoith D: Developmental regulation of the rat insulin-like growth factor I receptor gene. Proc Natl Acad Sci USA 1989; 86:7451–7455.
Liu JP, Baker J, Perkins AS, Robertson EJ, Efstratiadis A: Mice carrying null mutations of the genes encoding insulin-like growth factor I (Igf-1) and type 1 IGF receptor (Igf1r). Cell 1993;75:59–72.
Baker J, Liu JP, Robertson EJ, Efstratiadis A: Role of insulin-like growth factors in embryonic and postnatal growth. Cell 1993;75: 73–82.
Rappolee DA, Sturm KS, Behrendtsen O, Schultz GA, Pedersen RA, Werb Z: Insulin-like growth factor II acts through an endogenous growth pathway regulated by imprinting in early mouse embryos. Genes Dev 1992;6:939–952.
Kunkel LM, Smith DK, Boyer SH, Borgaookar SD, Wachtel SS, Miller OJ, Breg R: Analysis of Y chromosome specific reiterated DNA in chromosomal variants. Proc Natl Acad Sci USA 1977;74:1245–1249.
Sambrook J, Fritsh EF, Maniatis T: Molecular Cloning: A Laboratory Manual, ed 2. Cold Spring Harbor Laboratory Press, 1989.
Malcolm S, Donlon TA: Report of the 2nd International Workshop on human chromosome 15 mapping 1994. Cytogenet Cell Genet 1994;67:1–22.
Ullrich A, Tam AW, Yang-Feng T, Tsubokawa M, Collins C, Henzel W, Le Bon T, Kathuria S, Chen E, Jacobs S, Francke U, Ramachandran J, Fujita-Yamaguchi Y: Insulin-like growth factor I receptor primary structure: Comparison with insulin receptor suggests structural determinants that define functional specificity. EMBO J 1986;5:2503–2512.
Abbott AM, Bueno R, Pedrini MT, Murray JM, Smith RJ: Insulin-like growth factor I receptor gene structure. J Biol Chem 1992;267: 10759–10763.
Orita M, Suzuki Y, Sekiyu T, Hatashi K: Rapid and sensitive detection of point mutations and DNA polymorphisms using the polymerase chain reaction. Genomics 1989;5:874–879.
Gustaffson TA, Rutter WJ: The cysteine-rich domains of the insulin and insulin-like growth factor I receptors are primary determinants of hormone binding specificity. J Biol Chem 1990;265:18663–18667.
Ullrich A: Insulin-like growth factor I receptor cDNA cloning. Methods Enzymol 1991; 198: 17–26.
LeRoith D, Werner H, Beitner-Johnson D, Roberts CT Jr: Molecular and cellular aspects of the insulin-like growth factor I receptor. EndocrRev 1995;16(2):143–163.
Kato H, Faria TN, Stannard B, Roberts CT, LeRoith D: Role of tyrosine kinase activity in signal transduction by the insulin-like growth factor-I (IGF-I) receptor. J Biol Chem 1993; 268:2655–2661.
Acknowledgements
S.A.A. and E.W. were funded by the MRC and Children Nationwide. S.A.A. is now a Dunhill Medicine Trust Fellow. S.P. is grateful for support from the Child Growth Foundation and Serono Laboratories. We would also like to thank Professor Peter Scambler (ICH) for use of the PhosphorImager and Lisa Lowry (RPMS) for sequencing the PCR products. We are grateful to Mr. L. Butler, Director of the NE Thames Regional Cytogenetics Unit, for the karyotype analyses and to our clinical colleagues who referred families for study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abu-Amero, S., Price, S., Wakeling, E. et al. Lack of Hemizygosity for the Insulin-Like Growth Factor I Receptor Gene in a Quantitative Study of 33 Silver Russell Syndrome Probands and Their Families. Eur J Hum Genet 5, 235–241 (1997). https://doi.org/10.1007/BF03405923
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03405923