Abstract
Hirschsprung disease is a congenital disorder clinically characterized by the absence of colonic ganglia and genetically by extensive heterogeneity. Genes involved include RET, GDNF, EDNRB and EDN3. Mutations of these genes may give dominant, recessive, or polygenic patterns of inheritance. In particular in the case of missense mutations, it is therefore far from easy to assess whether a given mutation will contribute to the phenotype. We discuss criteria for such an assessment and pay special attention to functional assays. The interpretation of mutations as contributing to a disease phenotype or as merely representing a rare polymorphism has direct clinical consequences. Hirschsprung disease with major and modifying sequence variants in a variety of genes might well serve as a model for the many complex disorders for which the search for genes involved has only just been initiated.
Similar content being viewed by others
Hirschsprung disease (HSCR) or colonic aganglionosis is a congenital disorder characterized by intestinal obstruction due to an absence of intramural ganglia along variable lengths of the colon. The incidence of HSCR is 1 per 5,000 live births. Both genetic and environmental factors are thought to contribute to the development of the disease phenotype. In some families a clear dominant pattern of inheritance is observed, while in others a recessive form or a sex-modified multifactorial pattern of inheritance seems to apply. Moreover, HSCR is found associated with several syndromes and dysmorphic features. HSCR seems therefore to be a heterogeneous disease in which mutations in some genes give a dominant pattern of inheritance, mutations in other genes lead to a recessive mode of inheritance, and further genes contribute to the phenotype according to a polygenic/multifactorial model [1]. The genetic heterogeneity of HSCR is obvious from the occurrence of mutations in different genes. A dominant pattern of inheritance is found associated with mutations in the RET gene [2, 3], which codes for a receptor tyrosine kinase (fig. 1). The effect of mutations in EDNRB [4], coding for the endothelin-B receptor (fig. 2), and in EDN3 [5, 6], coding for one of the ligands of the endothelin-B receptor (fig. 3), seems to be dosage dependent, since homozygous mutations in EDNRB and EDN3 have both been found associated with a combined Waardenburg type 2 and HSCR phenotype (Shah-Waardenburg syndrome), while patients with heterozygous mutations, present either a HSCR-only phenotype or certain symptoms of the Waardenburg type 2 syndrome [4–10]. Mutations in a fourth gene, GDNF [11, 12], the gene coding for the ligand of the RET receptor (fig. 4), have been suggested to be insufficient to cause HSCR, but to contribute to the phenotype in combination with mutations in other genes.
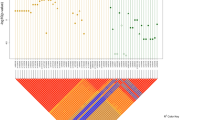
Schematic representation of the RET mRNA, the encoded protein, and the mutations found to date [16, 22–24]. Indicated are the positions of the mutations in both the protein and the mRNA (exons). Missense mutations are shown above the schematic representation of the mRNA, truncating mutations are shown under it.
Schematic representation of the EDN3 mRNA, the encoded protein preproendothelin, and the mutations found to date [5, 6, 10, 18]. Indicated are the mature peptide EDN3, which is the product of proteolytic cleavage of the preproendothelin by furin and the endothelin converting enzyme 1, the ET3-like domain, and a signal peptide domain.
Schematic representation of the GDNF mRNA, the encoded protein, and the mutations found to date [11, 12, 19]. Indicated in the figure are the potential secretion signal (S), the consensus sequence for proteolytic processing (P), the conserved peptide domains (C), the furin cleavage site (FCS), and the endothelin converting enzyme cleavage sites (ECE1CS), and the positions of the mutations reported to date.
In HSCR as well as in many other disorders, the involvement of a number of genes, either independently or in association, makes mutation detection and interpretation of the mutations — in particular missense mutations — far from easy. The question whether a single mutation contributes to the disease phenotype or merely represents a rare polymorphism is constituting a major problem with obvious direct clinical consequences.
Several criteria are generally applied, usually in combination, to assess the possible pathogenicity of a missense mutation: (1) de novo appearance of a mutation; (2) segregation of the mutation with the disease within pedigrees; (3) absence of the mutation in control individuals; (4) a change of amino acid polarity or size in the encoded peptide; (5) occurrence of the amino acid change in a domain which is evolutionarily conserved between species and/or shared between proteins belonging to the same protein family; (6) effect of the mutation in a functional in vitro system or in an animal model; (7) previous inclusion of the mutation in the increasing number of disease-specific mutation databases. Application of these criteria is not without difficulties, as will be evident from a number of examples in HSCR.
For EDNRB, most of the mutations observed are missense mutations (fig. 2). One of these, Gly57Ser, was considered pathogenic as it occurred in a highly conserved region and was not present in 130 control chromosomes [9]. Moreover, the mutation changes a nonpolar amino acid into a polar one. Occurrence of this mutation was independently confirmed in 3 of 40 HSCR patients, but also seen in 7 of 90 control individuals (180 chromosomes) [5]. Although the arguments for pathogenicity [9] looked solid, the later findings make the variant more likely a nonpathogenic polymorphism. A comparable example is provided by a specific RET mutation. The C to T transition in exon 19 of RET, leading to the substitution of a nonpolar cysteine for a polar arginine at codon 982, i.e. near the end of the functionally important tyrosine kinase domain, was initially found in 3 out of 80 unrelated HSCR patients and not observed among 100 control chromosomes. It was therefore assumed to be a causative mutation at the time of presentation of the data in a ‘RET Gene Mutations’ workshop at the annual meeting of the American Society of Human Genetics in 1994 [13]. Subsequently, the same mutation was found to occur twice in 180 control chromosomes [14]. Furthermore, a functional test did not demonstrate any measurable effect [15]. Thus, causative as the mutation seemed a first sight, it is more likely that it is not associated with HSCR [16].
Regarding the ‘functional test’ argument, one should be cautious: small effects on the disease phenotype may easily be missed, notably when the functional test, as in the above-mentioned case, makes use of an expression vector with a strong promoter. Moreover, it cannot be ruled out that effects of an in vivo test, such as (in)activation of cell-specific signaling pathways, would be different from those of an in vitro test. An overview of functional assays for the genes involved in HSCR is given in figure 5. Interpretation of functional assays may cause more problems. If a mutation results in a truncated protein, testing activation by RET of different cellular substrates [29, 30] (in particular Shc [31, 32], Grb2 [32], Grb10 [33], PLCγ [34], Crk [35], Nck [35]). EDNRB functioning can be measured via ligand-induced transient Ca2+ levels in cells after transfection with mutant EDNRB cDNA [4]. To examine the functional significance of an EDN3 mutation at the protein level secretion of EDN3 can be tested after transfecting a mutant EDN3 cDNA into an ECE-1 expressing cell line (CHO/ECE1 cells). As the protein encoded by EDN3, preproendothelin, is enzymatically cleaved by furin and ECE1 into the mature peptide EDN3, secretion of the peptide into the culture medium can be measured [18]. If the mutation is present in the mature peptide the functioning of EDNRB, which is activated by EDN3, might be tested. its phenotypic effect in a functional assay might seem unnecessary. Nonetheless, a recent breast cancer study [17] revealed a noncausative BRCA2 variant leading to a stop codon in the 3′ part of the gene. A mutation of EDN3 provides another example. A single nucleotide insertion was found as a heterozygous mutation in a patient with congenital central hypoventilation syndrome (CCHS), a disease which in rare cases is found associated with HSCR, although not in this patient [18]. The mutation leads to truncation of a part of the protein which is normally cleaved off during processing of the precursor to the mature product. Although it was not excluded that this mutation might represent a rare polymorphism, it was actually presented mainly as evidence for a common molecular pathogenesis of HSCR and CCHS. Still, when the mutant cDNA was transfected into Chinese hamster ovary cells, the mature peptide was produced in normal quantities. In the same in vitro test system, the EDN3 missense mutation found in an HSCR patient with Waardenburg type 2 and leading to an amino acid substitution elsewhere in a precursor domain outside the mature peptide [5] did cause near-complete absence of mature EDN3 product [M. Yanagisawa, personal commun.].
Functional tests for the proteins involved in HSCR. The figure shows a schematic representation of the proteins involved in HSCR. Based upon the different functions of the proteins, specific functional assays can be applied. RET, a receptor tyrosine kinase, is normally activated (phosphorylated) by its ligand GDNF in combination with GDNFR-α. No functional tests for GDNF have been described yet. Conceivably, the effect of mutant GDNF cDNA, transiently expressed and secreting its product into the culture medium, might be assessed by determining the activity of transiently expressed wild type RET. Activation of the transiently expressed (mutant) RET cDNA, in a cell line expressing GDNFR-α and stimulated by GDNF, can be measured via the phosphorylation status of the protein [15, 25–27], its transforming capacity [15, 25–27], the transport of the protein to the plasma membrane [28], or the binding to or
The interpretation of mutations in HSCR patients has become even more complicated with two reports [11, 12] each describing patients with a different RET mutation (one de novo) accompanied by the same GDNF missense mutation, Arg93Trp, thus suggestive of a digenic mode of inheritance. Both reports concluded that in these cases the mutated GDNF protein most probably cannot cause HSCR on its own, but presumably contributes to the phenotype in combination with mutant RET [11, 12]. A similar combined effect might also apply to another GDNF mutation, Asp150Asn, accompanied by a trisomy 21 [12]. The same mutation was found once (in a sporadic HSCR patient) when we screened 100 HSCR patients for GDNF mutations. A comprehensive mutation scanning of all known HSCR genes by DGGE failed to detect any further mutation in this patient (our own unpublished results). A third GDNF mutation, Pro21Ser, was present homozygously in a healthy father of two affected heterozygous individuals [12]. Whether GDNF should only be seen as a modifier gene can be debated, as in a sporadic HSCR patient a de novo mutation of GDNF has been found [19]. This might point to a major contribution of GDNF to the phenotype of that patient. That HSCR most probably is or can be polygenic has been demonstrated more convincingly by haplotype analysis in a large Mennonite kindred [4, 20]. Interpretation of mutations on the assumption of a di/polygenic mode of inheritance is difficult. Consider for example the GDNF mutation Asp150Asn, mentioned before, which has been found in HSCR patients both in combination with a trisomy 21 and on its own. When we screened 150 control individuals (300 chromosomes) we found the mutation once. On the assumption of di/polygenicity, finding the same mutation in a patient and in a control individual no longer excludes the fact that the mutation can contribute to the disease phenotype. It may be noted that all but one of the patients reported with a GDNF missense mutation inherited their mutation from a healthy parent. One would expect, however, that the frequency of such mutations in the normal population would be lower than that in the patient population. The Asp150Asn mutation is caused by a G to A transition leading to the substitution of a neutral polar amino acid residue for an acidic polar one. Comparison of GDNF with members of the TGF-β super-family [21] shows three highly conserved sequence domains, a close alignment of the seven cysteine residues of GDNF and conservation of these amino acid residues among all TGF-β superfamily members. The Asp150Asn mutation is at the very border of the first conserved domain and immediately next to one of the cysteine residues. Thus, there are several arguments favoring the idea that this might represent a pathogenic mutation. When a polygenic mode of inheritance is assumed, functional assays will also have their limitations. It might well be that a mutation will only have an effect in the context of (mutant) modifier-genes (protein-protein interactions) or even external factors interfering with specific mutant proteins.
As discussed, there is a variety of arguments that can help to decide whether or not a given variant should be considered as a causative mutation. In a polygenic model it might nevertheless be virtually impossible to conclude whether a given mutation in one of the genes supposed to be involved will contribute to the disease phenotype or not. In this respect, HSCR with its likely polygenic etiology, and with major and modifying sequence variants, might serve as a model for the many complex genetic disorders for which the search for genes involved has only just been initiated.
References
Badner JA, Sieber WK, Garver KL, Chakravarti A: A genetic study of Hirschsprung disease. Am J Hum Genet 1990;46:568–580.
Romeo G, Ronchetto P, Yin L, Barone V, Seri M, Ceccherini I, Pasini B, Bocciardi R, Lerone M, Kδδriδinen H, Martucciello G: Point mutations affecting the tyrosine kinase domain of the RET proto-oncogene in Hirschsprung’s disease. Nature 1994;364:377–378.
Edery P, Lyonnet S, Mulligan LM, Pelet A, Dow E, Abel L, Holder S, Nihoul-Fékété C, Ponder BAJ, Munnich A: Mutations of the RET proto-oncogene in Hirschsprung’s disease. Nature 1994;367:378–380.
Puffenberger EG, Hosoda K, Washington SS, Nakao K, de Wit D, Yanigisawa M, Chakravarti A: A missense mutation of the endothelin-B receptor gene in multigenic Hirschsprung’s disease. Cell 1994;79:1257–1266.
Hofstra RMW, Osinga J, Tan G, Wu Y, Kamsteeg E-J, Stulp RP, van Ravenswaaij-Arts C, Angrist M, Chakravarti A, Meijers C, Buys CHCM: A homozygous mutation in the human endothelin-3 gene associated with a combined Waardenburg type 2 and Hirschsprung phenotype (Shah-Waardenburg syndrome). Nat Genet 1996;12:445–447.
Edery P, Attie T, Amiel J, Pelet A, Eng C, Hofstra RMW, Martelli H, Badaud C, Munnich A, Lyonnet S: Mutation of the endothelin-3 gene in the Waardenburg-Hirschsprung disease (Shah Waardenburg syndrome). Nat Genet 1996;12:442–444.
Kusafuka T, Wang Y, Puri P: Novel mutations of the endothelin-B receptor gene in isolated patients with Hirschsprung’s disease. Hum Mol Genet 1996;5:347–349.
Auricchio A, Casari G, Staiano A, Ballabio A: Endothelin-B receptor mutations in patients with isolated Hirschsprung disease from a non-inbred population. Hum Mol Genet 1996;5: 351–354.
Amiel J, Attie T, Jan D, Pelet A, Edery P, Bidaud C, Lacombe D, Tam P, Simeoni J, Flori E, Nihoul-Fekete C, Munnich A, Lyonnet S: Heterozygous endothelin receptor B (EDNRB) mutations in isolated Hirschsprung disease. Hum Mol Genet 1996;5:355–357.
Bidaud C, Pelet A, van Camp G, Salomon R, Attie T, Eng C, Bonduelle M, Nihoul-Fekete C, Willems P, Munnich A, Lyonnet S: Endothelin-3 gene mutations in isolated and syndromic Hirschsprung disease. Eur J Hum Genet, in press.
Angrist M, Bolk S, Halushka M, Lapchak PA, Chakravarti A: Germline mutations in glial cell line-derived neurotrophic factor (GDNF) and RET in a Hirschsprung disease patient. Nat Genet 1996;14:341–344.
Salomon R, Attie T, Pelet A, Bidaud C, Eng C, Amiel J, Sarnacki S, Goulet O, Ricour C, Nihoul-Fekete C, Munnich A, Lyonnet S: Germline mutations of the RET ligand GDNF are not sufficient to cause Hirschsprung disease. Nat Genet 1996;14:345–347.
Angrist M, Bolk S, Chakravarti A: Mutation detection in autosomal dominant Hirschsprung disease: SSCP analysis of the RET pro-to-oncogene. Am J Hum Genet 1994;55:A209.
Hofstra RMW, Cheng NC, Hansen C, Stulp RP, Stelwagen T, Clausen N, Tommerup N, Caron H, Westerfeld A, Versteeg R, Buys CHCM: No mutations found by RET mutation scanning in sporadic and hereditary neuroblastoma. Hum Genet 1996;97:362–364.
Pasini B, Borrello MG, Greco A, Bongarzone I, Luo Y, Mondellini P, Alberti L, Miranda C, Arighi E, Bocciardi R, Seri M, Barone V, Radice MT, Romeo G, Pierotti MA: Loss of function effect of RET mutations causing Hirschsprung disease. Nat Genet 1995;10:35–40.
Angrist M, Bolk S, Thiel B, Puffenberger EG, Hofstra RMW, Buys CHCM, Chakravarti A: Mutation analysis of the RET receptor tyrosine kinase in Hirschsprung disease. Hum Mol Genet 1995;4:821–830.
Puget N, Healey CS, Gayther SA, Mangion J, Stratton MR, Lynch HT, Goldgar DE, Ponder BAJ, Lenoir GM: A polymorphic stop codon in BRCA2. Nat Genet 1996;14:253–254.
Bolk S, Angrist M, Xie J, Yanigisawa M, Silvestri JM, Weese-Mayer DE, Chakravarti A: Endothelin-3 frameshift mutation in congenital central hypoventilation syndrome. Nat Genet 1996;13:395–396.
Ivanchuk SM, Myers SM, Eng C, Mulligan LM: De novo mutation of GDNF ligand for the RET/GDNFR-a receptor complex in Hirschsprung disease. Hum Mol Genet 1996;5:2033–2036.
Chakravarti A: Endothelin receptor-mediated signaling in Hirschsprung disease. Hum Mol Genet 1996;5:303–307.
Burt DW: Evolutionary grouping of the transforming growth factor-β superfamily. Biochem Biophys Res Commun 1992;184:590–595.
Yin L, Barone V, Seri M, Bolino A, Bocciardi R, Ceccherini I, Pasini B, Tocco T, Lerone M, Cywes S, Moore S, van der Winden JM, Abramowicz MJ, Kristofferson U, Larsson LT, Hamel BCJ, Silengo M, Martucciello G, Romeo G: Heterogeneity and low detection rate of RET mutations in Hirschsprung disease. Eur J Hum Genet 1994;2:272–280.
Attie T, Pelet A, Edery P, Eng C, Mulligan LM, Amiel J, Boutrand L, Beldjord C, Nihoul-Fekete C, Munnich A, Ponder BAJ, Lyonnet S: Diversity of RET mutations in Hirschsprung disease. Hum Mol Genet 1995;4:1381–1386.
Hofstra RMW, Osinga J, Stulp RP, Scheffer H, Meijers C, Buys CHCM: Mutations in three genes are found associated with the development of Hirschsprung disease: RET, EDNRB and EDN3. Am J Hum Genet 1996;59:A263.
Santoro M, Carlomagno F, Romano A, Bottaro DP, Dathan NA, Grieco M, Fusco A, Vecchio G, Matoskova B, Kraus MH, Di Fiore PP: Activation of RET as a dominant transforming gene by germline mutations of MEN 2A and MEN 2B. Science 1994;267:381–383.
Borrello MG, Smith DP, Pasini B, Bongarzone I, Greco A, Lorenzo MJ, Arighi E, Miranda C, Eng C, Alberti L, Bocciardi R, Mondellini P, Scopsi L, Romeo G, Ponder BAJ, Pierotti MA: RET activation by germline MEN 2A and MEN 2B mutations. Oncogene 1995; 11:2419–2427.
Carlomagno F, De Vita G, Berlingieri MT, de Franciscis V, Melilo RM, Colantuoni V, Kraus MH, Di Fiore PP, Fusco A, Santoro M: Molecular heterogeneity of RET loss of function in Hischsprung’s disease. EMBO J 1996; 15: 2717–2725.
Iwashita T, Murakami H, Asai N, Takahashi M: Mechanism of RET dysfunctioning by Hirschsprung mutations affecting its extracellular domain. Hum Mol Genet 1996,5:1577–1580.
Songyang Z, Carraway KL III, Eck MJ, Harrison SC, Feldman RA, Mohammdi M, Schlessinger J, Hubbard SR, Smith DP, Eng C, Lorenzo MJ, Ponder BAJ, Mayer BJ, Cantley LC: Catalytic specificity of protein tyrosine kinases is critical for selective signalling. Nature 1995; 373:536–539.
Liu X, Vega QC, Decker RA, Pandey A, Worby CA, Dixon JE: Oncogenic RET receptors display different autophosphorylation sites and substrate binding specificities. J Biol Chem 1996;271:5309–5312.
Asai N, Murakami H, Iwashita T, Takahashi M: A mutation at tyrosine 1062 in MEN2A-RET and MEN2B-RET impairs their transforming activity and association with Shc adaptor protein. J Biol Chem 1996;271:17644–17649.
Borello MG, Pelicci G, Arighi E, De Filippis L, Greco A, Bongarzone I, Rizeti MG, Pelicci O, Pierotti MA: The oncogenic versions of the RET and TRK tyrosine kinases bind Shc and Grb2 adaptor proteins. Oncogene 1994;9: 1661–1668.
Pandey A, Duan H, Di Fiores PP, Dixit VM: The RET receptor protein tyrosine kinase associates with the SH2-containing adaptor protein Grb10. J Biol Chem 1995;270:21461–21463.
Borello MG, Alberti L, Arighi E, Bongarzone I, Battistini C, Bardelli A, Pasini B, Piutti C, Rizzetti MG, Mondellini P, Radice MT, Pierotti MA: The full oncogene activity of Ret/ptc2 depends on tyrpsine 539, a docking site for phospholipase Cγ. Mol Cell Biol 1996;16: 2151–2163.
Bocciardi R, Mograbi B, Pasini B, Borello MG, Pierotti MA, Bourget I, Fisher S, Romeo G, Rossi B: The MEN2B mutation switches the specificity of the RET kinase towards cellular substrates that are susceptible to interact with Crk and Nck. Abstr Eur Soc Hum Genet 29th Annu Meeting, Med Genet 1997;9:16.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hofstra, R.M.W., Osinga, J. & Buys, C.H.C.M. Mutations in Hirschsprung Disease: When Does a Mutation Contribute to the Phenotype. Eur J Hum Genet 5, 180–185 (1997). https://doi.org/10.1007/BF03405914
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03405914