ABSTRACT
The myeloperoxidase (MPO) is an important microbicidal protein present at high concentration in the primary granule of mature granulocyte and its expression is regulated in both myeloidcell-lineage and premyelocytic-stage-specific manners. A better understanding of the underlying control mechanisms should provide insights into the temporal and co-ordinate regulation of the gene expression during granulopoiesis. We have identified its promoter by mapping the start(s) of transcription using various molecular approaches together with demonstrating the promoter function of the relevant DNA segment in a transient transfection reporter assay. Besides the major start of transcription mapped at G residue, 11 nucleotide upstream of the 3′ end of exon 0, the usage of that is specific to the MPO expressing cell lines, we have shown that irrespective of the MPO-expression status of the hematopoietic cells, transcription occurs broadly within a two kb region upstream of the 5′ proximity of the gene, and is largely terminated in intron 2. These data support a model of the premyelocytic-stage-specific MPO expression, the control of which is operated at initiation as well as elongation levels of transcription.
Similar content being viewed by others
INTRODUCTION
During hematopoiesis, both lymphoid and myeloid progenitor cells are derived from pluripotent stem cells. Myeloid progenitors further commit to one of six cell lineages and eventually form mature differentiated cells, i.e., erythrocytes, platelets (megakaryocytes), macrophages, neutrophils, eosinophils and basophils. This complex but ordered processes requires both temporal and co-ordinate control of the expression of both cell lineage-and differention-stage-specific genes, control of that is predominantly operated at the level of transcription. Understanding the control mechanisms as such demands insight into the control of individual lineage-specific 'marker' genes for each lineage. Within the hematopoienc lineages, extensive studies have described a complex interplay of regulatory cis-elements and binding nuclear proteins, in particular for the β-globin enhancer 5′ locus control region1 and the immunoglobulin heavy chain gene J-C intronic enhancer2. In this respect, there is a marked deficit of information on the granulocytic lineage, a major pathway of non-erythroid myeloid cell development.
Expression of the myeloperoxidase (MPO) gene which encodes an abundant microbicidal protein in mature neutrophilic granulocyte, is restricted to a narrow window extending from the late myeloblastic to the premyelocytic stag of granulopoiesis3. Both termination of transcription and reduction of the stability of mRNA are responsible for the switch-off of the MPO expression4. Although the onset of the MPO expression may begin with an initiation of transcription, whether the control at other levels, such as elongation of transcription is also involved remains to be determined.
In a broad term, two types of key cis-elements, i.e. enhancer and promoter participate in the transcriptional control of the protein-coding genes in eukaryotes5. We have previously identified an enhancer element at −3.4 kb upstream of the mouse MPO gene, by virtue of its hypersensitivity to DNase I in chromatin as well as potent activity in a MPO-expressing myelomonocytic cell line, WEHI 3 BD+, but not in lymphoid cell lines where the MPO gene does not express6. This enhancer consists of three elements which were initially identified as WEHI 3 BD+ specific in vitro DNase I footprints (FTI to III) and work synergistically for the maximum enhancer function7. An active participation of some novel transcription factors in the enhancer function were suggested by both sequence uniqueness of these elements and a drastic enrichment of their cognate binding proteins in WEHI 3BD+ cells in comparison to lymphoid cells. Substantiation of this notion was subseqeuntly obtained from a detail characterization of element FTI and its cognate transcription factor8. Nevertheless, others are favor of a predominant involvement in this process, of the several known transcription factors, including pu-1 and CEBP, expression of that is not restricted to the granulocytic lineage9,10. Furthermore, the potential enhancer elements elsewhere may also play a role in the control of the lineage-specific expression of the MPO gene11, 12.
Probably reflecting the complex nature of the control over the transcription initiation of the mouse MPO gene, little is known on its promoter. The cDNA cloning effort resulted in a number clones varying in their 5′ ends of the cDNA insert13. Friedman et al., concluded from their mapping exercises involved using the primer extension assay with a single primer in a MPO expressing cell line 32D Cl3 along with a non-hematopoietic cell line, that there are two starts of MPO transcription specific to the MPO-expressing myeloid cells14. One start was mapped at −210 bp upstream of the gene, whereas whether the second start was at the 3' end of exon 0 or within intron 1 remains undetermined. Besides the technical concern, a reservation has been raised, on the myeloil lineage-specific nature of the conclusion which was obtained from an experiment excluding any MPO non-expressing hematopoietic cell line. In this connection, the DNA fragment encompassing the start 1 was later found devoid of any promoter function in the stable transfected 32D Cl3 cells12.
To understand the transcriptional control of the mouse MPO gene, its myeloid-specific expression through granulopoiesis in particular, we used a combination of molecular approaches to map the start(s) of transcription in two MPO expressing along with six MPO non-expressing hematopoietic cell lines, representing various hematopoietic lineages. The reporter gene construct where the luciferase gene was driven by the DNA fragment encompassing the major start of MPO at its 3′ end, was found possessing a promoter function in all the hematopoietic cell lines tested. The ranscription activity of this fragment is also approximately 45 fold more potent in a MPO expressing than several MPO non-expressing hematopoietic cells. Finally, we have also shown an active participation of the regulation at the elongation level of transcription in the control over the differential expression of the MPO gene in granulopoiesis.
MATERIALS AND METHODS
General methods for molecular genetic manipulations described in Molecular Cloning Manual15 were used in this study.
Cell cultures
WEHI 3 BD+4,16, 18.817, EL-418, 416B19, NFS-6020, MEL21 and J774.222 cells were cultured as previously described9 (Tab 1).
RNA preparation and Northern analysis
RNA was prepared directly from, or from both nuclear and cytoplasmic fractions of cells by the guanidium thiocynanate/acid phenol method23. Northern blotting analysis was carried out according to Lehrach's protocol24 and probed with 32P labeled oligonucleotide probes at the conditions suggested by Henderson et al25. After an overnight hybridization at 60–65 °C, the membranes were finally washed at 55–60 °C in 0.1 × SSPE/0.5 % SDS. The size of the hybridized RNA band was estimated by taking as size markers, both acridine orange stained 18S and 28s RNA bands and hybridized band of the glyceraldehyde-3-phosphate dehydrogenase cDNA (approximately 0.8 kilonucleotide in size).
Primer extension
Primer extension was essentially performed according to McKnight's protocol26 with following modifications: 1, the hybridization was carried out overnight rather than for 2 h at 60-65 °C, 2, the hybridization buffer is at pH 6.8 rather than 7.4. After a 2 h incubation with reverse transcriptase (Supertranscript TM, Gilbco), the extended products were ethanol precipitated and analyzed by electrophoresis on a 5 % sequencing gel. The size markers were provided by both the sequencing ladders with suitable primers on the relevant plasmid DNA templates and the 32P labeled Msp I digests of PBR322 DNA.
S1 nuclease protction assay
For mapping the transcripts initiated within the region extending from −954 to +11 (Here refers the major start of transcription as +1), in Fig 4, the plasmid DNA containing the relevant fragment in bluescripts SK II vector (Stratagene) was used as template for 15 cycles of linear PCR with the biotin tagged T3 primer. The single-strand biotin labeled template was magnetically isolated by using Dynabead DS M-280 Strata divin beads according to the manufacturer's instruction. The single-strand 5′ end labeled probe was made on the isolated template by T7 DNA polymerase with 4 pmole 32P labeled primed [exon 0, Tab 2] at 42 °C for 30 min. For Fig 6, the S1 probe was made as described above except for the following details: 1, the Xba II linearised plasmid DNA containing a 600 bp fragment covering the 5′ end of cDNA was used as template; 2, the biotin tagged T7 primer was used as the first primer; and 3, 32P labeled primer Da1 (exon 2, Tab 2) was used as the second primer.
The transcription occurs within the region extending from −954 to +11 of the mouse MPO gene, assyaed by both primer extension and S1 mapping from the 5′ end of primer d.Various cellular RNA was either subjected to the primer extension with primer d (+ (11 to −12), Tab 2); or S1 mapping with a single stranded probe made with primer d from a relevant template. Size markers were made by the 32P labele d Msp I fragments of PBS 322 and a sequencing ladder with labeled primerd on a relevant template. Lanes: 1 and 9: 5 μg WEHI 3 BD + cytoplasmic RNA; 2 and 10: 5 mg WEHI 3 BD+ total RNA; 8: 20 μg yeast tRNAs; 3 and 11: NFS-60; 4 and 12: WEHI 3 BD-; 5 and 13: 416 B; 6 and 14: 18.8; 7 and 15: EL-4 cells, respectively. The remaining cases contain 10 μg total RNA from.
Mapping the major start of the MPO transcripts which correlates with the differentiation-stage-specific expression of the mouse MPO gene by both primer extension and S1 mapping from the same end. 32P-kinase labeled primer Da 1 was used both for primer extension experiment [lanes 1-8] and for making the single stranded probe of S1 mapping [lanes 9-16], which covers the first 608 bp of the cDNA of the mouse MPO gene. Other experimental details were same as the previous figures. Lanes: M, 32P labeled Msp I digest of PBR 322 plasmid DNA; P, S1 probe; 1 and 16: 18.8 RNA; 2 and 15: EL-4; 3 and 14: WEHI 3 BD-; 4 and 13: J77 4.2; 5 and 12: MEL RNA; 6 and 9: NFS-60 RNA; 7 and 10: cytoplasmic RNA from WEHI 3 BD+ cells; 8 and 11: total RNA of WEHI 3 BD+ cells. For WEHI 4 BD+ cells 5 μg RNA was used, while for others, 1 0 μg RNA was used in this experiment.
After ethanol precipitation, the RNA was hybridized to the S1 probe as described for the primer extension. S1 nuclease digestion of the hybrids was initiated by adding 300 μ1 of S1 nuclease solution, i.e., 100 units of S1 nuclease in 0.28 M NaCl, 50 mM sodium acetate, pH 4.5, and 4.5 mM Zinc sulfate, followed by an incubation at 37 °C for 60 min. The protected products were extracted with phenol/chloroform, ethanol precipitated and analyzed by electrophoresis on a 5 % sequencing gel, followed by antoradiographing.
Multiple S1 nuclease protection assay
A set of oligonucleotides varying in length from 14 to 34 mers were individually kinase labeled at their 5′ end with 32P-γATP. A mixture of the labeled primers at an equal molar ratio were hybridized with RNA, followed by S1 nuclease digestion. The protected labeled oligos were analyzed by electrophoresis on a 20 % sequencing gel.
The promoter construct and the reporter gene assay
By PCR cloning with pfu enzyme (Stratagene), a DNA fragment extending from −947 to +14 of the MPO gene was cloned at a promoterless luciferase based reporter vector, pGL-2-basic (Promega), to create construct ini-OH. 1 mg ini-OH was electroporated into cells in duplicate by using a Bio-Rad Gene Pulser, at the following condition: at 250 volts, and 960 μC with 0.4 ml cell suspension at a concentration of from 0.5 to 5 × 106 cells per ml. After an overnight incubation, the luciferase activity of the combined extracts of each duplicate was measured by using Promega luciferase assay system (E1500) and the single photon mode in a PACKARD 2500 TR Liquid scintillation analyser. The mean ± SD was calculated from the results of a minimum two independent experiments, and the strength of the MPO promoter was described as ratio of its luciferase activity over that of the control plasmid Pt-109 (in which the luciferase gene is controlled by the thymidine kinase (TK) promoter (−109 to +52,) of the Herpes Simplex virus.
RESULTS
Transcription initiates broadly within a wide region upstream of the mouse MPO gene, disregarding the MPO expressing status of hematopoietic cells
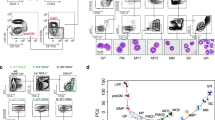
To address the premylocytic-stage-specific nature of the MPO gene expression we included in this study, six MPO non-expressing along with two MPO expressing cell lines: WEHI 3 BD+ cell where the MPO gene expressed at a high level and NFS-60 cell line in which there is only a moderate level of the MPO expression. Apart from two distant lymphoid cell lines, EL-4 (pre-T) and 18.8 (pre B), there are two cell lines in other myeloid lineages, i.e. MEL in erythrocytic and J744.2 in the monocytic lineage toward macrophage, and two cell lines in the same lineage as WEHI 3 BD+, i.e., WEHI 3 BD- and 416 B (Tab 1 and Fig 1).
Hematopoietic cell lineages and the mouse cell lines used in this study A simplest view of the hematopoiesis is presented. Both myeloid and lymphoid progenitors are derived from pluripotent stem cells. Myeloid progenitors diversify, forming one of four different precursors, which subsequently develop to one of eight types of mature cells, as indicated. Lymphoid progenitors commit themselves further to form either T or B lymphoid precursors which eventually produce mature T and B lymphocytes, respectively. The mouse cell lines used for this study and their approximate positions in hematopoiesis are also indicated. Two MPO-expressing cell lines: WEHI 3 BD+ and NFS-60 are under lined.
Under-representation of the 5 part of mRNA is common in the conventionally constructed cDNA libraries, probably as a result of the inherent tendency of the reverse transcription reaction to stop at the G-C rich region of mRNA during the first strand synthesis. The 5 terminus of the longest cDNA clones is more likely to locate at downstream of the real 5 end of the mRNA in vivo. Therefore, we searched for the major start(s) of the transcription at the upstream of the 5 end of the MPO gene, represented by the 5 end of the cDNA insert of the longest clone. Three independent primer extension experiments were carried out with primers 1, 2, 3, the locations of which are at −667 and −208 upstream and +24 bp downstream of the 5 end of the exon 0, respectively [Tab 2]. Suprisingly, the pattern of the extended bands of the MPO expressing differs very little from the non-expressing cells (Fig 2 for primer 1, and data not shown). There were two predominant bands, * and ** which are approximately 610 and 240 nucleotides in size, and potentially represent the starts of transcription at −1210 and −920 bp upstream of the 5′ end of the MPO gene (Fig 2). Since the end of exon 0 is actually at −490 upstream of the major start of transcription which would be described in the later sections, the location of both bands ** and * are also described in relation to the major start, as bands −1700** and −1430*, respectively. As band −1430* is equally dense in the lanes associated with 416 B, EL-4 and WEHI 3 BD+ cellular RNA, and relatively fainter in the lane with 18.8 RNA, we paid more attention to band −1700** that seems specific to WEHI 3 BD+ RNA. The primer extension also tends to stop at the regions rich in secondary structures, of the RNA template, we carried out an primer extension experiment with primer B, which is 349 bp up stream of primer 1 to reassess the start of transcription, represented by band −1700** in Fig 2.However, the corresponding band was present in all the lanes without any significant variation in its density, associated with both WEHI 3 BD+, and each of five MPO non-expressing cell's RNA (Fig 3). Moreover, there is an addition band, i.e., band −1800***, representing the start at 100 nucleotides further upstream exists in all lanes at a very similar density. We further performed a primer extension experiment with primer X which 297 bp upstream of primer B. In addition to a confirmation of the real existence of the sites represented by band −1700** and band −1800*** in Fig 3, there were other two bands further upstream which share the same pattern (data not shown). In view of the fact that the extended product from a closer primer more likely reflects the real 5end of RNA in cell than that from a distal primer, the pattern of band −1700** in Fig 2 is likely artificial.
The upstream transcription activity of the mouse MPO gene demonstrated by the primer-extension assay with primer 1. 10 mg total RNA from the following cell lines were annealed with 0.1 pm 32P labeled primer 1 at its 5 end, followed by an extension reaction with reverse transcriptase. The extended products were electrophoretically separated on a 7.0 M urea/5% poly acrylamide gel, then an autoradiograph of which was made. The location of the potential starts of transcription, reflected by the mobility of the extended bands were indicated both side to the autorad and in the drawing underneath. The size markers are provided by both sequencing ladder with 32P labeled primer 1 on the relevant template and labeled Msp I digested fragments of PBR 322 plasmid DNA.
The upstream transcription activity of the mouse MPO gene demonstrated by the primer extension assay with primer B. 10 μg total RNA of several cell lines (indicated at the top of the autorad) were subjected to the primer extension analysis with primer B. The size markers were provided by the sequencing ladder of a relevant plasmid DNA, primed with 32P labeled primer B on a relevant template.
Since all these cell lines have been well-documented at aspects of their MPO expressing status, which contradicts to the implication of the observation (Fig 2, 3 and data not shown), namely, the upstream initiated transcription does not correlate with the lineage specific expression of the MPO gene in hematopoiesis, we have to eliminate the possibility of our data being an experimental artifact by the following arguments. First, the yeast tRNA does not lead to any extended products with several primers (not shown). Second, the experiments have been carried out in a most stringent manner, for instance, the hybridization is carried out at 60-65 °C. Finally, the existence of the key starts of transcription have been concluded by the consensus among, at least, two independent primer extension experiments with different primers [Fig 2 and 3 and not shown]. In this connection, an alternative interpretation has also been considered, that the transcription initiated at the MPO gene region in WEHI 3 BD+ cell may come from different gene loci in other hematopoietic cell lineages, if the involved gene loci share both sequence of the relevant primers and distance between the initiation sites to the relevant primers. This is very unlikely either. There is no precedent for a significant sequence similarity to this extent at the upstream region of even among the members with in a single gene family. The requirement, for the transcriptions to be initiated at the same distance away from the sites represented by the primers on different gene templates among these cell lines are even more difficult to be met.
Nevertheless, to consolidate the observations in Fig 2 and 3, we, in parallel with a compatible primer extension, used S1 nuclease protection assay, which requires 100 % sequence complementarility between the target RNA and the single strand probe to analyze the transcription activity within the region extending from −954 to +11 (Fig 4). A 5′ end labeled antisense single strand probe was made and hybridized to the total RNA from seven cell lines. Following S1 digestion of the hybrids, the protected single strand probes were separatedy electrophoresis. The pattern of the extended products by d primer which are characteristic with many bands shorter than 150 nucleotide in length (lanes 1-7, Fig 4), differs drastically from the pattern of S1 mapping (lanes 9-15, Fig 4), where the predominant band is the full length probe together with many faint bands equally distributed across the lanes. Since all such bands are absent in yeast tRNA lane (8, Fig 4), we should conclude that transcription initiates at many sites in a comparable frequency within the region extending from −947 to +11, between the MPO-expressing and non-expressing cell lines. It is also probably correct to conclude that transcription initiated further upstream shown in Fig 2 and 3 are indeed real in all the hematopoietic cells tested, in view of both a slight overlap between the regions covered in Fig 4 and Fig 2, and the consensus among the primer extension experiments involved with different primers.
Northern analysis with oligo nucleotide probes confirms the myeloid-lineage specific nature of the MPO gene transcription
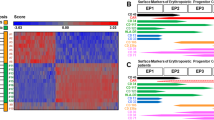
Expression of increasing numbers of the eukaryotic genes are found to be mainly controlled at the level of elongation28, 29, 30. Differential termination of the upstream initiated transcription may be resporisible for the lineage-specific expression of the mouse MPO expression. Therefore, we first asked whether the upstream initiated transcripts constitute any part of the mature MPO transcripts ? The Northern blots of 20 mg total RNA from WEHI 3 BD+ and three MPO-non-expressing cell lines: WEHI 3 BD-, 18.8 and E1-4 were sequentially probed with the 32P labelled oligonucleotides (Tab 2). As shown in Fig 5, a band representing the full length mature MPO mRNA (approximately 2.7 kilo nucleotides in size), along with two bigger but lighter bands were only detected in the lane containing WEHI 3 BD+ RNA by exon 1 (oligo f), exon 2 (oligo Da 1) and exon 3 (oligo E1) as well as the cDNA probe (panel 5, 7, 8 and 2, Fig 5). Therefore, despite of that upstream transcription occur equally among all the hematopoietic cells tested (Fig 2, 3, 4), the steady state MPO RNA is only present in the WEHI 3 BD+ cells but not others, as previously documented. None of three bands were hybridisable by oligo 3a, which is −137 upstream of the major start (panel 4, Fig 5), indicating that any participation of the upstream initiated transcripts across this region, in the formation of the steady state MPO mRNA, is not likely. Therefore, the two big faint bands are not MPO RNA precursors, the products of the complete transcription of the MPO gene. Instead, they are probably a result of inadequate denaturation of RNA which is often associated with the form aldehyde-agarose based Northern analysis. This is indeed supported by the compatible hybridzation pattern with gyceraldehyde-3-phosphate dehydrogenase gene probe, a ubiquitously expressed house keeping gene (panel 1, Fig 5).
Northern blot analyses with the oligo nucleotide primers 2 0 μg total RNA of following cell lines: WEHI 3 BD+, WEHI 3 BD-, 18.8, and EL-4, were analyzed by Northern blotting assay with various 32P labeled probes: Panels: 1, the cDNA probe of the mouse glyceral dehyde-3-phosphate; 2, the cDNA probe of the mouse MPO gene; 3, primer e [intron 1]; 4, 3a [exon 0]; 5, primer f [exon 1]; 6, primer g [intron 2]; 7, primer Da 1 [exon 2] and 8, primer E1 [exon 3]. The relevant information on primers is in Tab 2 and the line drawing underneath the autorads. The size markers were provided by either acridine orange stained bands of 28s and 18s rRNA revealed by UV illumination and the hybridband of the glyceraldehyde-3-phosphate dehydrogenase gene.
Finally, the undetectability of any MPO bands with either intron 1(e) or 2(g) oligo probes suggests that splicing out of both intron 1(e) and 2(g) is likely to be an operationally inseparable event from the formation of the MPO primary transcripts.
The usage of the start of transcription, at the G residue, 11 bp from the 3′ end of exon 0 is specfic to the MPO-expressing cells
Failing to find any correlation of the starts described in Fig 2, 3, 4, with the lineage-specific expression of the mouse MPO gene, together with the absence of the exon 0 in the steady status of MPO RNA (Fig 5) prompted a search, within the downstream region for the major start(s) of the MPO transcription which may link directly to the myeloid-lineage-specific expression of the MPO gene. Both S1 mapping and primer extension experiment from the site represented by the 5′ end of the exon 2 oligo, Da 1, were carried out. As shown in Fig 6, except for all the documented MPO-non-expressing cell lines, primer extension with Da 1 primer revealed three bands in both total and cytoplasmic RNA fractions of WEHI 3 BD+ cell (lanes 7 and 8, Fig 6). A weak top band was also detected in the total RNA from NFS-60 cells (lane 6, Fig 6). A comparable result was obtained from the S1 protection experiment with a probe which sequence corresponds to the region from −490 to +118 of the cDNA or from −490 to +679 devoid of both intron 1 and 2 of the MPO genomic sequence (lanes 9–16, Fig 6). By comparing the sequencing ladders from the cDNA template with Da 1 oligonucleotide, the ends of these three bands, representing the starts of the transcription are mapped at the underlined nucleotides in the following sequence: TTGACAGTGTCAG||AGATGAA. The G residue, mapped at 11 nucleotides upstream of the 3′ end of exon 0 is the major start of transcription which were detected in both WEHI 3 BD+ and NFS-60 cells, although it was much weaker in the later, probably reflecting the relative level of the MPO expression in cell. The minor start, mapped at G, 7 nucleotides from the 3′ terminus of exon 0, was absent in the lane associated with NFS-60 RNA. Finally, it is not clear as to if the A residue, mapped at the first residue of exon 1, may represent the real start of transcription or the end of a processed transcripts in WEHI 3 BD+ cells.
Premature termination of the upstream initiated transcription occurs mainly at intron 2
The aforementioned observations indicated that the upstream transcripts were likely to terminated at upstream of the exon 2, demarcated by Da 1 oligo. To determine the region where termination may take place, we first carried out primer extension experiments with two partially overlapped exon 1 primers i. e. Zc and f, (Tab 2). Although the results in Fig 7 corroborated the conclusion regarding to the major start in Fig 6, the band reflecting the minor start was also seen in primer extension experiment with primer f in the lanes associated with either 416 B or WEHI 3 BD- RNA (Fig 7). Since this has not been confirmed by the experiment with primer Zc which is closer to the start of transcription, the presence of such a band shown in the experiment with primer f, is likely to be an experimental artifact. There are bigger bands in all the lanes associated with the RNA from all four cell lines (Fig 7). Since neither primer extension and S1 mapping experiments covering the region upstream of exon 2, (Da 1) defected any transcription with all the MPO non-expressing cells (Fig 6), the upstream initiated transcripts in the those cells are mainly, if not entirely, terminated within the region between exon 1 and e, i.e., the 159 bp intron 2. Finally, there is no difference of the bands representing either major or minor starts between nuclear and cytoplasmic fractions of WEHI 3 BD+ RNAs, substantiating the view that the splicing out both intron 1 and 2 is tightly linked to the formation of primary transcripts.
Mapping the major start of the MPO transcripts with two partially overlapped exon 1 primers [primers Zc and f] Various RNAs [indicated at the top of the autorad] were used for primer extension with primer Zc and f, respectively. 5 μg total, cytoplasmic and nuclear RNA fractions of WEHI 3 BD+ cells, 10 μg total RNA from NFS-60, WEHI 3 BD- and 416 B cells were used in this study.
Although molecular mechanisms for the premature termination of transcription is still unclear, co-existed antisense transcripts may play a key part in the differential control of expression of the gene coding erb A, c- and N- myc, or basic fibroblastic growth factor31. Therefore, it is desirable to assess the antisense transcription around intron 2 where the termination of the upstream MPO transcription are likely to occur (Fig 5, 6, 7). Three primers for detection of the antisense transcripts (Tab 2) at this region are: 1, Alu-1, a 17 mer oligo nucleotide contain eleven nucleotides which shares only 52 % similarity with the consensus of the Alu I repeat, and a six nucleotide insertion. Therefore, this primer should be specific to the 180 bp Alu repeat starting from 29 bp upstream of exon 1, at the antisense orientation. 2, Za 1-, a 25 nucleotide primer which is located at 5' part of exon 1. And 3, Zb-, is 20 nucleotide oligo nucleotide which starts 9 bp down stream of the 3′ end of exon 1. As shown in Fig 8, while no bands specific to oligos Alu- and Za- are detectable, the Zb- band is obvious in the RNA from both MPO-expressing and non-expressing cell lines. By taking the β-actin band as a loading control, the Zb- antisense transcript appears more abundant in EL-4, WEHI 3 BD- and 416 B cells than WEHI 3 BD+ and NFS-60 as well as 18.8 cells.
Assessment of the relevant abundance of the RNA transcribed from the various regions at the 5′ part of the MPO gene by a multiple - S1/oligonucleotide assay Various RNAs were hybridized with a set of 32P end labeled oligonucleotides, the information of which is indicated left to the autorad and in the line drawing underneath. After S1 digestion of the hybrids, the remaining was electrophoretically separated and autorad of which was obtained from the dried gel.
Finally, we have noticed that the bands h (Exon 2) and f (Exon 1) in lanes 1-3 associated with the RNAs from WEHI 3 BD+ and NFS-60 cells, are lighter than band E1 (Exon 3), indicating that a partial termination of the MPO transcription occurs at between exons 2 and 3 in these two MPO expressing cells.
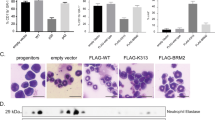
The DNA Segment containing the major start of transcription possesses a promoter function and is more potent in the cells expressing the MPO gene
Our mapping efforts has established a close correlation of the major start of transcription, mapped at the G residue, 11 nucleotide from 3′ end of exon 0, to the lineage-specific expression of the MPO gene (Fig 4, 5, 6, 7, 8). Here, we employed a conventional transient transfection assay to determine if the relevant DNA fragment possesses any promoter function and if such an activity is more potent in MPO expressing than none-expressing cells. The DNA fragment extending from −954 to +14 was put at upstream of the firefly luciferase gene to form construct ini-OH (panel A, Fig 9). As shown in Panel B, Fig 9, the luciferase activity of the cellular extracts of cells transfected by ini-OH is comparable to that by pt 109 where the luciferase gene is driven by the HSV thymidine kinase gene promoter (from −109 to +52), in all the MPO non-expressing cells including EL-4, MEL, 416B, 18.8 and WEHI 3 BD-. As expected, the promoter activity of this fragment is approximately 45 fold more potent than pt 109 in WEHI 3 BD+ cells, where the MPO gene is actively transcribed and the major start of transcription is used. This may be contributed by both an exclusive usage of the major start of transcription in WEHI 3 BD+ cells, and the presence of an enhancer element within this fragment12.
The promoter function of a relevant region [−954 to +14] of the MPO gene in cells analyzed by a transient tansfection assay Panel A, The line-drawing shows the relevant in formation of the luciferase construct (ini-OH) for assaying the MPO promoter function; Panel B, The relative luciferase activity of the cell extracts from the cells transfected with ini-OH over pt 109 along with the relevant SD value were calculated and shown.
DISCUSSION
Technical aspects of the mapping assays for the start of transcription of the mouse MPO gene: a paradigm
There are obvious discrepancies between our mapping data and Friedman's14. (Fig 10), which is very unlikely a result of the inherent difference between the MPO expressing cells, as Friedman et al used 32D Cl 3 cell line, whereas WEHI 3 BD+ and NFS-60 were used in our study. A subsequent observation, created further doubts on Friedman's mapping data, that the relevant DNA fragment did not display any promoter function in the stable transfected 32D Cl 3 cells12. On the contrary to Friedman's approach based on a single primer extension assay without any hematopoietic cell control, our conclusions were based on the observations from both S1 mapping as well as primer extension assay in a manner where any start of transcription have been confirmed by at least two independent assays with different primers on eight hematopoietic cell lines (Fig 2, 3, 4 and 6, 7). We found: 1, irrespective of the MPO-expressing status of the hematopoietic cells, transcription occurs broadly within a 2 kb region at the upstream of the gene; and 2, the transcription initiated at the major start mapped at the 490 bp down stream of the 5′ proximity of the exon 0 is specific to the MPO-expressing cells. Furthermore, the proven promoter activity of the fragment (−947 to +14) in all the cell lines by a transient transfection-reporter assay (Fig 9) provides an essential piece of functional evidence to the mapping data obtained by our approoach. In view of the demand for 100 % sequence complementarity between target RNA and single stranded probe, S1 mapping data should be able to stand on its own. By comparison with the S1 mapping data (Fig 4 and 7), it is possible to evaluate the observation in the context of the transcription of the mouse MPO gene from primer extension experiments, the advantage of which is sensitive and straight forward. As shown in Fig 6 and 7, three independent primer extension experiments with primer Da 1, f and Zc lead to an identical mapping result concerning the major start, which is in a full agreement with the S1 mapping data. But there are some differences. For instance, the presence of the minor site in both 416 B and WEHI 3 BD- cells shown by the primer extension with primer f, was undetectable by the primer extension with primer Da 1 and Zc, respectively (Fig 6 and 7). Therefore, it is rather premature to conclude both the presence and location of the start of transcription based on a single primer extension experiment.
Finally, we have compared the sequence similarity around the major start of transcription which is specific to the MPO expressing cell lines, WEHI 3 BD+ and NFS-60, between mouse and human [Fig 11]. There are 64.7 % identity between the −154 to +61 region of the mouse and the −146 to +68 of the human MPO. There is a TATA like sequence located at −24 and −19 upstream of the mouse and human MPO gene, respectively. Several cis-elements bound by known transcription factors are at similar locations within this segment. Considering the fact that the major start of the human MPO gene was determined by taking the end-labelled restriction fragments of plasmid DNA, PBR 32232 as the size marker, the likelihood for its having an error of a few nucleotides is quite high. Therefore, there is indeed a strong conservation in both structure and function of the MPO promoter between mouse and human.
The upstream transcription and its implication to the differential expression of the mouse MPO gene during hematopoiesis
It has been known for sometime that an alternative usage of promoters resulting in mRNA with different 5′ ends is responsible for the differential expression of a number of genes during development33. For instance, the stage-specific expression of a 1.7 kb mRNA for cytochrome c gene is produced by an initiation of transcription at an upstream rather than from the canonical site34. Another example is the human aldolase A gene, where initiation of transcription can occur at three promoters within 1.56 kb region, giving rise to the mRNA species differing at their 5′ ends and a selective use of these promoters determines its differential expression during muscle maturation35. Since none of upstream initiated transcripts constitutes any part of the mature MPO mRNA (Fig 5), the role, if any, played by the upstream transcription in the control of cell-lineage-specific expression must be different from the cases listed above.
There are precedents for upstream transcriptions at the sites lacking any features of the canonical promoter elements36, 37, 38, 39, 40, 41. It has been postulated that DNase I hypersensitive sites that reflect in the open chromatin structures may contribute to the upstream initiated transcription, based on the co-localisation of the initiation sites of upstream transcripts and DNase I hypersensitive sites in chromatin of the human epsilon globin gene42. However, Our attempts to detect such an open chromatin structure at the region where the upstream transcription actively occur has failed in both MPO-non expressing and -expressing hematopoietic cells6 (unpublished results). Other unknown mechanisms, therefore, may account for the observations described in this report.
Selective termination of transcription and its role in differential control of the MPO expression during hematopoiesis
Controls at the elongation level of transcription has been shown in a number of eukaryotic genes, including most if not all of the genes involving growth control and stress response of cell, such as, c-myc43, c-myb44, c-fos, and heat shock gene45. In all the cases, engaged RNA polymerase II machineries pause at as far as few hundred bp down stream of the start of transcription and continue the elongation process when receiving the appropriate signals28. The principal control of the gene expression in those case is concerning whether and when the initiated transcription is elonglated. The advantage of such a control may relate to the fact that activation of gene expression responds more quickly to extracellular signals by boosting the transcription rate to maximum in a shortest possible time. However, this control mechanism may not take place in the expression of mouse MPO gene, because 1, the MPO is neither a growth control nor stress-responsive gene and 2, the termination mapped at intron 2 (Fig 6, 7, 8), is as far as 3 kb from the most distant initiation site identified (Fig 2).
We have also explored the possible involvement of the antisense transcription into the elongation control or premature termination of the MPO trarscripts31 by assessing the presence of antisense transcripts at around the exon 1 region (Fig 8). Indeed, there is a loose correlation between the abundance of an antisense transcript, designated by primer Zb-, and termination of sense transcripts. By the multiple S1 oligo nucleotide in this report, we also witnessed a partial termination of the MPO transcription at downstream of exon 2 in both MPO expressing cell lines (rather than at upstream of exon 2 in all MPO non-expressing cell lines) (Fig 8). However, further investigation is required for both consolidation of the results in this report and uncovering the mechanisms behind.
A model of both cell-lineage-and differentiation-stage-specific control of the mouse MPO gene during hematopoiesis
Based on our results in this report and previous works6, 7, 8 we postulated a model for the control of the mouse MPO gene during granulopoiesis (Fig 12). There are, at least, four operationally defineable events concerning the cell-lineage and differentiation-stage-specific expression of the mouse MPO gene, during the hematopoiesis in which the pluripotent stem cells evolve to the mature granulocytes. The first event relates to the upstream transcription coupling with its termination at upstream of exon 2, which may start when the first major commitment of the pluripotent stem cells is made to either myeloid and lymphoid progenitors and lasts ever since, till the formation of all the mature cells. This is supported by the observation that all the cell lineages being tested in this report possess this feature, including erythroid, neutrophilic granulocytic, monocytic toward macrophage, B lymphoid as well as T lymphoid (Fig 2, 3, 4). This phenomenon appears specific to hematopoietic lineages as Friedman failed to detect any transcription of this kind in a non-hematopoietic cell control14. The second event concerns the change from the repressive to open chromatin structure of the distant enhancer (−3.4 kb), which was detected as a strong DNase I hypersensitive site in WEHI 3 BD+ cells, but not in lymphoid cells6. A relevant study with an in vitro differentiation system suggested that this DNase I hypersensitive site occurs even in the multipotential progenitor (myeloid progenitor)-stage prior to the onset of the MPO expresson which happen only in granulocyte-committed hematopoietic progenitors9. Therefore, the commitment to other myeloid lineage of the myeloid progenitor likely parallels with the disappearance of this DNase I hypersensitivity site. It seems not possible to separate the following two events: activation of the distal enhancer and the transcription from the major start. Their specific association with the expression status of the premyeloid stage-specific granulocyte has been vividly demonstrated by the sharp contrasts with regarding to the usage of the major start for transcription between MPO-expressing granulocytic cell lines, represented by WEHI 3 BD+ and NFS-60 cells, and non-expressing granuloctyic cell lines (WEHI 3 BD- and 416B) in this report (Fig 2, 3, 4, 5, 6, 7, 8, 9). It is apparent, more works need to be carried out for a better understanding of both the nature of the extracelluar signals and the relevant pathways leading to activation of the MPO expression. However, this hypothetical model should provide a useful framework for addressing these molecular details of the myeloid-lineage and premyelocytic-stage-specific expression of the gene expression in general and the mouse MPO gene in particular.
References
Orkin SH . Globin gene regulation and switching: circa 1990. Cell 1990; 63:665–72.
Staudi L, Lenardo M . Immunoglobulin gene transcription. Annu Rev Immunol 1991; 9:373–98.
Koeffler HP, Kanyard J, Pertcheck m . Myeloperoxidase: its structure and expression during myeloid differentiation. Blood 1985; 65:484.
Tobler A, Miller CW, Johnson KR, Selsted ME, Rovera G, Koeffler H ?. Regulation of gene expression of myeloperoxidase during myeloid differentiation. J Cell Physiol 1988; 136:215–25.
McKnight S, Yamamoto RK . Transcriptional regulation. Vol. 22. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York. 1992.
Zhu JD, Bennett CA, MacGregor AD, Greaves MF, Goodwin GH, Ford AM . A myeloid-lineage-specific enhancer upstream of the mouse myeloperoxidase gene. Leukaemia 1994; 8:717–23.
Zhu JD . The myeloid-lineage-specifci enahncer of the mouse myeloperoxidase gene conssists of three cis-elements defined as in vitro DNase I footprints. FEBS Letters 1994; 339:243–8.
Zhu J . FT I, a novel positive myeloid-lineage-transcription regulatory element with the mouse myeloperoxidase gene enhancer, En1. Cell Research 1995; 5:75–91.
Ford A, Bennett C, Healy L, Towatari M, Greaves M, Enver T . )Regulation of the myeloperoxidase enhancer-binding proteins PU1, C-EBP-alpha, C-EBP-beta, C-EBP-delta during granulocyte-lineage specfication. Proc Natl Aca Sci USA 1996; 93:10838–43.
Scott L, Civin CI, Rorth P, Friedman AD . A novel temporal expression pattern of three C/EBP family members in differentiating myelomonocytic cells. Blood 1992; 7.
Friedman A, Britosbray M, Sozow J . The murine myeloperoxidase gene contians a bipartite distal enhancer, including a novel region regulated by PEBP2/CBF. Leukemia Research 1996; 20:817–20.
Suzow J, Friedman -AD . The murine myeloperoxidase promoter contains several functional elements, one of which binds a cell type-restricted transcription factor, myeloid nuclear factor-I (mynf1). Mol Cell Biol 1993; 13:2141–51.
Venturelli DNS, Gemperlein I, Bittenbender S, Hudson S, Rovera G . Nucleotide sequence of cDNA for murine myeloperoxidase. Nucl Acids Res 1989; 17:5852.
Friedman AD, Krieder BL, Venturelli D, Rovera G . Transcriptional regulating of two myeloid-specific genes, myeloperoxidase and lactoferrin, during differentiation of the murine cell line 32D C13. Blood 1991; 78:2426–32.
Sambrook J, Fritsch EF, Maniatis T . Molecular cloning-A laboratory manual (2 edn). Cold Spring Harbor Laboratory Press, 1989; New York.
Warner NL, Moore MAS, Metcalf D . A transplantable myelomonocytic leukemia in BALB/c mice: cytology, karyotype, and muramidase content. J Natl Cancer Inst 1969; 43:963–82.
Alt F, Rosenberg N, Lewis S, Thomas E, Baltimore D . Organization and reorganization of immunoglobulin genes in A-MuLV-trasnformed cells: rarrangementof heavy but not light chain gene. Cell 1981; 27:381–90.
Herberman R . Serological analysis of cell surface antigens of tumours induced by murine leukaemia virus. J Natl Cancer Inst 1972; 48:265–71.
Dexter TM, Allen TD, Scott D, Teich NM . Isolation and characterisation of a bipotential haematopoietic cell line. Nature 1979; 277:471–4.
Holms KL, Palaszynski E, Fredrickson TN, Morse III HC, Ihle JN . Correlation of cell-surface phenotype with the establishment of interleukin 3-dependent cell lines from wild-mouse murine leukemia virus-induced neoplasms. Proc Natl Acad Sci USA 1985; 82:6687–91.
Slosberg E, SCHER B, SCHER W, JOSEPHSON S, WAXMAN S . Localization of tresite of action of protease in ducers of erythroid-differentiation of mouse erythroleukemia (mel) cells to the cell-surface. 1986; 14:442.
Unkeless J . Characterization of a monoclonal antibody directed against mouse macrophage and lymphocyte Fc receptors. J Exp Med 1979; 150:580–96,.
Chomczynski P, Sacchi N . Single step method of RNA,isolation by acid guanitdium thiocyanate-phenol-chloroform extraction. Anal Biochem 1987; 162:156–9.
Lehrach H, Diamond D, Wozney JM, Boedtker H . RNA molecular weight determinations by gel electrophoresis under denaturing conditions, a critical reexamination. Biochemistry 1977; 16:4743.
Henderson GS, Coonary JT, Davidson JM, Stewart SJ, House FS, McCurley TL . A reliable method for northern blot analysis using synthetic oligonucleotide probes. Biotechniques 1991; 10:190–6.
McKnight SL, Kingsbury R . Transcriptional control signals of a eukaryotic protein-coding gene. Science 1985; 217:316–24.
Nordeen SK . Luciferase reporter gene vectors for analysis of promoters and enhancers. Biotechniques 1988; 6:454–8.
Spencer CA, Groudinge M . Transcription elongation and eukaryotic gene regulation. Oncogene 1990; 5:777–85.
Richardson JP . Transcription termination. Crit Rev Biocehm Mol Biol 1993; 28:1–30.
Greenblatt J, Nodwell JR, Mason SW . Transcriptional antitermination. Nature 1993; 364:401–6.
Nellen W, Hildebrandt M, Mahal B, Mohrle A, Kroger P, Maniak M, Oberhauser R, Sadiq M . Mechanisms of gene regulation by endogenous and artificially introduced antisense RNA. Biochem Soc Trans 1992; 20:750–4.
Chang KS, Zhao SR, Wang YP, Lu JF Trujillo JM, Stass SA, Freireich EJ . Down regulation of myeloperoxidase gene associated with specific nuclease hypersensitive sites during TPA induced differentiation of HL-60. Leukemia 1991; 5:205–9.
Schibler U, Sierra F . Alterantive promoters in development gene expression. Ann Rev Genet 1997; 21:237–57.
Hake LE, Hecht NB . Utilisation of an alternative transcription initiation site of somatic cytochrome c in the mouse produces a testis-specific cytochrome c mRNA. J Biol Chem 1993; 268:4788–97.
Izzo P, Costanzo P, Lupo A, Rippa E, Paoleila G, Calvacore F . Human aldolase A gene: structural organization and tissue-specific expression by multiple promoters and alternate mRNA processing. Eur J. Biochem 1988; 174:569–78.
Geng Y, Johnson LF . Lack of an initiator element is responsible for multiple transcriptional intiatiion sites of the TATA-less mouse thymidylate synthease promoter. Mol Cell Biol 1992; 13:4894–903.
Ishii S, Nerlino GT, Patan I . Promoter region of the human Harvey ras proto-oncogene: similarity to the EGF receptor proto-oncogene promoter. Science 1985; 230:1378–81.
Ishii SX YH, Stratton RH, Roe BA, Merlino GT, Pastan I . Characterisation and sequence of the promoter region of the human epidermal growth factor recptor gene Proc Natl Acad Sci USA 1985; 82:4920–4.
Reynolds GA, Goldstein JL, Brown MS . Multiple mRNAs for 3-hydoroxy-3-methylglutaryl coenzyme A reductase determined by multiple transcription initiation sites and intron splicing sites in the 5′ untranslated region. J Biol Chem 1985; 260:10369–77.
Allan M, Lanyon WG, Paul J . Multiple origins of transcription in the 4.5 kb upstream of the epsilon-globin gene. Cell 1983; 35:187–97.
Allan M, Zhu J, Montague P, Paul J . Differential response of multiple epsilon-globin cap sites to cis-acting and trans-acting controls. Cell 1984; 38:399–407.
Zhu J, Allan M, Paul J . The chromatin structure of the human epsilon globin gene: nuclease hpersensitive sites correlate with multiple initiation sites of transcription. Nucl Acids Res 1984; 12:9191–204.
Krumm A, Meulia T, Brunvand M, Groundine M . The block to transcriptional elongation within the human c-myc gene is determined in the promoter-proximal region. Genes Dev 1992; 6:2201–13.
Watson RJ . A transcriptional arrest mechanism invovled in controlling constitutive levels of mouse c-myb mRNA. Oncogene 1988; 2:267–72.
Lee H, Kraus KW, Wolfner M, Lis JT . DNA sequence requirements for generating paused polymerase at the start of Hsp70. Genes and Dev 1992; 6:284.
Acknowledgements
This work was supported by both Cancer Research Campaign and Leukaemia Research Fund, UK.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
ZHU, J. Myeloidcell-lineage and premylocytic-stage-specific- expression of the mouse myeloperoxidase gene is controlled at initiation as well as elongation levels of transcription. Cell Res 9, 107–134 (1999). https://doi.org/10.1038/sj.cr.7290010
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290010
Keywords
This article is cited by
-
Neuronal regulation of type 2 innate lymphoid cells via neuromedin U
Nature (2017)
-
The ATF/CREB site is the key element for transcription of the human RNA methyltransferase like 1 (RNMTL1) gene, a newly discovered 17p13.3 gene
Cell Research (2002)
-
The promoter analysis of the human C17orf25 gene, a novel chromosome 17p13.3 gene
Cell Research (2002)