Abstract
A total of 738 colonies from 64 localities along the African continent have been analysed using the DraI RFLP of the COI–COII mitochondrial region. Mitochondrial DNA of African honeybees appears to be composed of three highly divergent lineages. The African lineage previously reported (named A) is present in almost all the localities except those from north-eastern Africa. In this area, two newly described lineages (called O and Y), putatively originating from the Near East, are observed in high proportion. This suggests an important differentiation of Ethiopian and Egyptian honeybees from those of other African areas. The A lineage is also present in high proportion in populations from the Iberian Peninsula and Sicily. Furthermore, eight populations from Morocco, Guinea, Malawi and South Africa have been assayed with six microsatellite loci and compared to a set of eight additional populations from Europe and the Middle East. The African populations display higher genetic variability than European populations at all microsatellite loci studied thus far. This suggests that African populations have larger effective sizes than European ones. According to their microsatellite allele frequencies, the eight African populations cluster together, but are divided in two subgroups. These are the populations from Morocco and those from the other African countries. The populations from southern Europe show very low levels of ‘Africanization’ at nuclear microsatellite loci. Because nuclear and mitochondrial DNA often display discordant patterns of differentiation in the honeybee, the use of both kinds of markers is preferable when assessing the phylogeography of Apis mellifera and to determine the taxonomic status of the subspecies.
Similar content being viewed by others
Introduction
Biodiversity of the honeybee Apis mellifera was first assessed using morphometrics. Based on an extensive sample collection and multivariate analyses, Ruttner et al. (1978) proposed a classification which was long considered as definitive. These authors hypothesized that north-eastern Africa and the Near East would be the centre of origin of Apis mellifera. They proposed that the species invaded Africa and Europe in three distinct branches, a South and Central African branch (A), a North African and West European branch (M) and a North Mediterranean branch (C). This classification was further refined by the addition of a fourth evolutionary branch, called O, which included the Near and Middle Eastern subspecies (Ruttner, 1988). Data on mitochondrial DNA have confirmed the presence of three lineages A, M, C in Africa, western Europe and south-eastern Europe (Smith & Brown, 1988; Garnery et al., 1992, 1993). The existence of a fourth mitochondrial lineage (O) in the Middle East has recently been confirmed (Arias & Sheppard, 1996; Franck et al., 2000b; Palmer et al., 2000). The same general structure of the species also emerged from microsatellite surveys (Estoup et al., 1995; Franck et al., 2000b). The main discrepancy provided by molecular markers was a clear genetic disruption between the branches M and A in the Iberian Peninsula (Smith et al., 1991; Garnery et al., 1995; Franck et al., 1998). Consequently, honeybees from northern Africa and from western Europe are considered to have followed separate evolutionary histories.
The introduction of the African subspecies A. m. scutellata in Brazil (Kerr, 1957), its subsequent hybridization with previously imported subspecies (Michener, 1975), and the rapid spread of this new hybrid strain (Africanized honeybee) throughout the Neotropics of America (Kerr, 1992) demonstrates how humans can greatly modify the genetic architecture of honeybee populations. Although most honeybee races present in Americas before the Africanization were native to Europe (Smith & Brown, 1988; Hall & Muralidharan, 1989; Hall & Smith, 1991; Sheppard et al., 1991; McMichael & Hall, 1996), there are some clues that African honeybees from other subspecies than scutellata had been introduced into different American regions (Schiff & Sheppard, 1993; Sheppard et al., 1999). It appears necessary to characterize the genetic diversity of honeybee races in their native range (Africa, Europe, and the Near-East). Only with this information will it be possible to accurately identify intrusion of unwanted genotypes and to develop coherent policies for conservation of local honeybee races. The present article focuses on African honeybees.
According to a recent review of the intraspecific nomenclature of Apis mellifera (Engel, 1999), 10 valid subspecies are recognized in Africa: A. m. intermissa (Maa, 1953) and A. m. sahariensis (Baldensperger, 1932) in Maghreb, A. m. lamarckii (Cockerell, 1906) and A. m. jemenitica (Ruttner, 1976) in north-eastern Africa, A. m. monticola and A. m. litorea (Smith, 1961) in south-eastern Africa, A. m. unicolor (Latreille, 1804) in Madagascar, A. m. scutellata (Lepeletier, 1836) and A. m. capensis (Eschscholtz, 1822) in southern Africa, and A. m. adansonii (Latreille, 1804) in western Africa. Hepburn & Radloff (1998) showed that these 10 subspecies have clearly separated morpho-clusters and they precisely delineated their geographical distribution. They also indicated geographical zones of high morphological variance within and between colonies, thus identifying hybrid zones among subspecies. Most African bees currently analysed belong to the A mitochondrial lineage (Smith, 1991; Garnery et al., 1992, 1993; Arias & Sheppard, 1996; De la Rùa et al., 1998). Only two colonies from Egypt have been recognized as belonging to lineage O (Arias & Sheppard, 1996; Franck et al., 2000b). Microsatellite loci are extremely polymorphic in African populations compared to European honeybee populations and this has been interpreted as a consequence of larger effective population sizes in Africa (Estoup et al., 1995; McMichael & Hall, 1996; Franck et al., 1998). African populations would have been less influenced by quaternary ice episodes which are considered to be the main cause of honeybee subspecies differentiation in Europe (Ruttner, 1988). However, the question of a common evolutionary origin of the honeybees from Africa remains unanswered.
In this paper, several new African honeybee populations are investigated. In light of these new data, the genetic variability of the honeybees from the African continent is reviewed by combining analyses of mtDNA and microsatellite polymorphisms.
Materials and methods
Sampling and DNA extraction
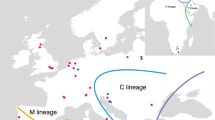
The origin of the samples is reported in Fig. 1. All African subspecies represented by a total of 738 honeybee colonies (254 colonies newly investigated) were sampled for use in this study. These colonies come from 64 locations in 21 African countries. In most cases the colonies have been morphometrically identified and several locations have been sampled within the endemic range of each subspecies (see Hepburn & Radloff [1998] for a review of original morphological data). A sample of 622 additional colonies from diverse locations in Europe, the Near East and America was used for comparison. Honeybee samples from each colony were brought back to the laboratory in vials containing 95% ethanol. The DNA was extracted from one leg with a chelex-based protocol (Estoup et al., 1996).
Distributions of the COI–COII honeybee mitotypes in 21 African countries and nine other countries used as reference. Pie charts indicate the frequencies of the five mitochondrial lineages M, C, O, Y and A. In the A lineage, the three groups of mitotypes – AI, AII and AIII – are distinguished (see Fig. 4 for the details of mitotypes included within each lineage or group). Values under country names are the numbers of locations (right) and colonies (left) analysed per country (see http://www.ensam.inra.fr/URLB for a complete description of mitotypes observed at each location). The stars refer to the countries in which populations have been analysed using microsatellite markers. The right up corner map refers to the natural distribution of Apis mellifera subspecies studied.
Mitochondrial DNA
The mtDNA region including the tRNAleu gene, the COI–COII intergenic region and the 5′ end of the COII subunit gene was PCR-amplified according to a protocol detailed elsewhere (Garnery et al., 1993). A fraction of the PCR product was run on a 1% agarose gel for total size determination and the remaining was restricted with DraI prior to electrophoresis on 7.5% polyacrylamide gel. The COI–COII mitotypes were determined for 404 new colonies. The newly found mitotypes were sequenced for further characterization as in Franck et al. (2000a). Additional data from De la Rùa et al. (1998), Garnery et al. (1993, 1995, 1998, Franck et al. (1998, 2000a, b) and Moritz et al. (1994) were included in the present study.
Microsatellite loci
Eight African populations for which a sufficient number of nonrelated honeybee individuals were available were scored at six microsatellite loci (A113, A43, A28, A24, A88 and B124). These came from Morocco (Al-Hoceima and Kenitra, A. m. intermissa; Tiznit, A. m. sahariensis), Guinea (Nimba, A. m. adansonii), Malawi (Chelinda, A. m. monticola), and South Africa (Johannesburg and Pretoria, A. m. scutellata; Cape Town, A. m. capensis). They were compared to eight additional populations from France (Valenciennes, A. m. mellifera, branch M), Sweden (Umeo, A. m. mellifera, branch M), Spain (Sevilla, A. m. iberiensis, branch M), Portugal (Porto, A. m. iberiensis, branch M), Italy (Forli, A. m. ligustica, branch C), Greece (Chalkidiki, A. m. macedonica, branch C), Sicily (Favara, A. m. siciliana, branch C), and Lebanon (El-Hermel, A. m. syriaca, branch O). The protocol followed and the original population data are reported in Estoup et al. (1995) and in Franck et al. (1998, 2000b).
Statistical and phylogenetic analysis
Unbiased estimates of gene diversity for microsatellite loci were calculated according to Nei (1978). Exact tests for genetic structure were computed using the GENEPOP package version 3.1 (Raymond & Rousset, 1995). The genetic differentiation between populations was computed using unbiased estimates of FST values provided by GENEPOP and the (dμ)2 microsatellite distance (Goldstein et al., 1995).
A maximum parsimony tree of COI–COII mitotypes was constructed using the PHYLIP package version 3.5c (Felsenstein, 1993). Presence/absence of base pair substitutions and insertions/deletions along the COI–COII intergenic sequence were coded as 1/0. When a component was missing, the corresponding characters were coded as missing data (e.g. insertions/deletions and substitutions within the P sequence in lineage C). A neighbour-joining tree from population samples was calculated from microsatellite data using the chord distance of Cavalli-Sforza & Edwards (1967). Bootstrap values were computed over 2000 replications (Hedges, 1992) re-sampling individuals within population.
Results
Mitochondrial DNA
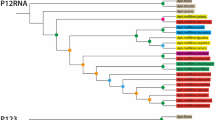
DraI RFLP of the COI–COII intergenic region provided a total of 42 different mitotypes among the 1359 colonies assayed (Fig. 2). Five mitotypes (A4′, A25, A26, A27, Y1, Y2) were not reported in previous analyses (Garnery et al., 1993, 1995, 1998; De la Rùa et al., 1998; Franck et al., 1998, 2000a, b; Palmer et al., 2000). Their restriction maps, the length of their restriction fragments, and their P sequences are given in Figs. 2 and 3.
Restriction maps (left) and restriction fragment sizes (right) of the 42 COI–COII mitotypes. The maps are deduced from DraI restriction pattern and sequences of the COI–COII intergenic mitochondrial region. Restriction sites are numbered from 1 to 10. Deletions and insertions are numbered preceded by characters d and i, respectively (see also Fig. 3). Exponent number refers to the number of equal size fragments.
Sequences of the P region of COI–COII mitotypes indicating substitution sites (small characters), DraI restriction sites (bold characters) and insertion/deletion (dashes). Four diagnostic sequences are defined: the supposed ancestral P0 sequence without deletion characterizing most of mitotypes of lineages A and O; the sequence P with the deletion d characterizing mitotypes of lineage M; the sequence P1 with the deletion d1 characterizing mitotypes of lineage A from the Atlantic coast; the sequence P2 with the deletion d2 characterizing mitotypes of lineage Y. Mitotypes C have no sequence P.
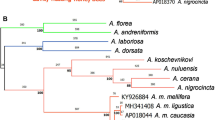
A total of 50 characters provided by COI–COII restriction patterns were used to assess the phylogeny of COI–COII mitotypes (Fig. 4). The samples from Ethiopia were characterized by mitotypes which clearly belong to a fifth lineage that we have called Y. All other mitotypes are clearly assigned to their previously described lineages A, M, C, or O (Franck et al., 2000b). The Y lineage has diverged from other lineages by around 2% (Table 1). Within lineage A, most of the mitotypes (group AI) are not differentiated in the phylogenetic analysis (Fig. 4). Only two sublineages are identified. The first one is characterized by mitotypes displaying only restriction site 1 (group AII). The second sublineage is characterized by mitotypes displaying the P1 sequence (group AIII).
Consensus tree (level 95%) of the COI–COII mitotypes over the equally parsimonious trees. The trees were established for 50 characters including restrictions sites, insertions and deletions (see Fig. 2). A, M, C, O and Y refer to the five major mitochondrial lineages. AI, AII and AIII refer to the three different groups of mitotypes within the lineage A.
The distribution of COI–COII mitotypes per country is given in Fig. 1. African colonies display mainly mitotypes A1 and A4 (group AI). Both mitotypes are also observed in high proportion in the Africanized bees from Mexico. In the populations from the Indian Ocean islands (A. m. unicolor), the proportion of mitotypes A1 is almost 100%. In continental African populations, the proportion of mitotypes A1 decreases progressively from Guinea toward south-eastern Africa and is replaced by mitotypes A4 in A. monticola, A. m. scutellata and A. m. capensis. From Guinea toward northern Africa, A1 is replaced by mitotypes A8, A9 and A10 (group AII) in A. m. sahariensis and A. m. intermissa. Most of the mitotypes from northern Africa are also observed in A. m. siciliana and A. m. iberiensis populations from southern Europe. Nevertheless, these populations display a high proportion of mitotypes A which are extremely rare in continental Africa. Mitotypes A2 and A3 (group AI) are principally observed in Spain and Sicily, and mitotypes of the AIII group are the most frequent within A. m. iberiensis populations from Portugal and the Canary Islands. Note that the populations from the Canary Islands also contain mitotypes C1 due to recent introduction of A. m. ligustica queens as in A. m. mellifera populations (Fig. 1). In north-eastern Africa, mitotypes belonging to three different lineages are identified (Fig. 1). Mitotypes A27 are observed only in A. m. litorea and they are highly divergent from other A mitotypes (Fig. 2). Mitotypes O and Y are observed only in A. m. lamarckii and A. m. jemenitica, respectively (Fig. 1).
Microsatellite loci
Gene diversity estimates per population and microsatellite locus range from 0.671 (A24, Kenitra) to 0.922 (A88, Chelinda). Mean gene diversities per population are given in Table 2. The samples from Africa display gene diversity higher than those from Europe and Middle East (Mann–Whitney’s U-test, P < 10 −6). Among the African samples, the Moroccan populations show the lowest values (Mann–Whitney’s U-test, P=4 × 10−5).
Fisher exact tests for genic differentiation indicate that all pairs of populations are significantly differentiated (Pmultilocus < 10−5). Among African populations FST values range from 0.0052 (Chelinda/Nimba) to 0.1239 (Kenitra/Cape Town) and (dμ)2 values range from 1.4535 (Pretoria/Nimba) to 12.1512 (Tiznit/Cap Town) (Table 3).
In the neighbour-joining tree (Fig. 5) the populations are separated into four groups {Al-Hoceima, Kenitra, Tiznit, Cape Town, Johannesburg, Pretoria, Nimba and Chelinda}, {Seville, Porto, Valenciennes and Umea}, {Favara, Forli and Chalkidiki}, {El-Hermel} corresponding to the four branches A, M, C and O of Ruttner’s classification (Ruttner, 1988). Within branch A, two sub-branches are identified. North-western African populations (Al-Hoceima, Kenitra and Tiznit) representative of the subspecies A. m. intermissa and A. m. sahariensis are clustered in the first sub-branch. The second sub-branch is composed of A. m. adansonii (Nimba) and A. m. monticola (Chelinda) populations in Equatorial Africa and A. m. capensis (Cape Town) and A. m. scutellata (Johannesburg, Pretoria) populations in southern Africa. Although A. m. iberiensis (Porto, Seville) and A. m. siciliana (Favara) populations display numerous A mitotypes, according to their morphological classification they cluster within branches M and C, respectively (Ruttner, 1988).
Discussion
The results can be summarized as follows. (1) Honeybees from north-eastern Africa contain three highly divergent mitochondrial lineages A, O, and Y, the third being newly characterized. (2) In the other parts of Africa, honeybees carry only mitotypes of lineage A. Nevertheless, honeybees from north-western Africa and from tropical and southern Africa highly diverge when mitochondrial and microsatellite data are considered. (3) southern European and American populations also display several A mitotypes. In Spain, Portugal and Sicily they have several possible origins, presumably resulting from successive ‘Africanizations’. In Mexico, the mitotypes A observed correspond to those displayed by A. m. scutellata populations from South Africa. (4) Microsatellite data indicate a high level of polymorphism in African populations compared to European ones.
The coexistence of three mitochondrial lineages in north-eastern Africa provides support for Ruttner et al.’s (1978) hypothesis that this area is the probable centre of origin for Apis mellifera. Note, however, that the Horn of Africa and the Rift Valley are the main channels of colonization from Asia to Africa. The presence of the mitochondrial lineage O in Egypt and Somalia and Y in Ethiopia may also result from successive honeybee invasions of Africa from the Middle East.
The quaternary climatic changes including desertification and vegetation shifts are considered quite important in explaining faunal distribution within the tropics (Potts & Behrensmeyer, 1992). They were probably responsible for honeybee subspecies diversification in Africa. The mitochondrial and microsatellite divergences between honeybee subspecies from northern and southern sides of the Sahara (mainly A. m. intermissa and A. m. adansonii) may have occurred during the late Pleistocene (around 15 000 years BP) when the Sahara extended largely southwards into the present Sahelian zone and the extreme north-west of Africa displayed favourable moist conditions (Mediterranean-like vegetation) for honeybees (Hooghiemstra et al., 1992; Lioubimsteva, 1995) (Figs. 1, and 5). In the Middle Holocene (around 8000 years BP), African climates became moister and the Sahara desert almost completely disappeared (Gasse et al., 1990; Hooghiemstra et al., 1992). This change has probably facilitated gene flow between honeybees from the Maghreb and the Sahel. This hypothesis is further strengthened by contact zones along both sides of the Atlas range evidenced by mitochondrial DNA data (Garnery et al., 1995). Another putative refuge for honeybees was the woodland and coastal forest along the borders of the Indian Ocean which persisted during the last arid period (around 15 000 years BP) when eastern Africa was largely characterized by open vegetation and the disappearance of forests at the top of eastern African mountains (Hamilton, 1982; Lovett, 1993a). This may explain the mitochondrial divergence observed between A. m. scutellata and A. m. litorea (Fig. 2) which was also recently reported by Meixner et al. (2000). The montane forest belt from the eastern African arc is rich in endemic species of plants (Lovett, 1993b). However the ecologically isolated subspecies A. m. monticola cannot be differentiated from A. m. scutellata using mitochondrial DNA markers, although some differences can be detected using isoenzyme analyses (Meixner et al., 1994, 2000). This may reflect recurrent introgression between both subspecies in the high tableland from northern Malawi studied here (N. Koeninger, personal communication). The honeybee subspecies from tropical and southern Africa (A. m. adansonii, A. m. monticola, A. m. scutellata, A. m. capensis and A. m. unicolor) are only slightly differentiated at the molecular level. The main difference among these five African subspecies arises from the variation in the proportion of mitotypes A1 and A4. It is noteworthy that both mitotypes only differ in the number of Q sequences (Fig. 2). Thus these differences are probably not evolutionarily significant across a Quaternary time scale. Consequently, the absence of endemic mitotypes and the absence of mitochondrial polymorphism in the A. m. unicolor populations could point to a recent colonization of the Indian Ocean islands by an extremely reduced number of colonies. Compared to A. m. unicolor, the ‘Africanized honeybee’ from Mexico displays both African mitotypes A1 and A4 whereas around 50 A. m. scutellata colonies have been introduced in Brazil in 1956.
The main explanation for the low molecular differentiation among African subspecies is probably a result of their highly migratory behaviour (absconding, swarming), specifically of A. m. adansonii and A. m. scutellata (Fletcher, 1978; Hepburn & Radloff, 1998). Direct genetic evidence of such behaviour is the high level of polymorphism at nuclear markers observed in all African populations assayed thus far (Table 2; Hall, 1992, 1998; Estoup et al., 1995; McMichael & Hall, 1996; Franck et al., 1998). This is also an explanation for the diffusion of mitotypes A outside the African continent into the Americas, the Iberian Peninsula and the Mediterranean Islands (Fig. 1 Hall & Muralidharan, 1989; Smith et al., 1989, 1991; Hall & Smith, 1991; Garnery et al., 1993, 1995; Sheppard et al., 1997; Sinacori et al., 1998). Interestingly, the diffusion of the African mitochondrial genome into southern Europe and the Americas does not necessarily correspond to the ‘Africanization’ of the nuclear genome (Lobo & Krieger, 1992, 2000a; Franck et al., 1998). Conversely, whereas European subspecies such as A. m. mellifera and A. m. ligustica have been repeatedly introduced into northern Africa (Second, 1975), not a single M or C mitotype has been found in north-continental Africa (but see Fig. 1 and De la Rùa et al., 1998 for the Canary Islands). Nevertheless, some evidence, including the sequences of microsatellite alleles (Franck, 1999), suggests nuclear introgression of Moroccan honeybees by some European alleles.
Consequently, mitochondrial data alone are probably insufficient to infer taxonomic and genetic status of honeybee colonies. Extending microsatellite analysis to honeybee subspecies from eastern Africa and the Middle East will be useful in the future for understanding the phylogeography of Apis mellifera and resolving relationships among all the African subspecies.
References
Arias, M. C. and Sheppard, W. S. (1996). Molecular phylogenetics of honey bee subspecies (Apis mellifera L.). Mol Phyl Evol, 5: 557–566.
Baldensperger, P. J. (1932). Variétés d’abeilles en Afrique du nord. 5th Congress International d’Entomologie, Paris.
Cavalli-Sforza, L. and Edwards, A. W. F. (1967). Phylogenetic analysis: models and estimation procedures. Am J Hum Genet, 19: 233–257.
Cockerell, T. D. A. (1906). New rocky mountain bees, and other notes. Can Entomol, 38: 160–166.
Cornuet, J. M., Garnery, L. and Solignac, M. (1991). Putative origin and function of the intergenic region COI and COII of Apis mellifera L. mitochondrial DNA. Genetics, 128: 393–403.
de La Rùa, P., Serrano, J. and Galliàn, J. (1998). Mitochondrial DNA variability in the Canary Islands honeybees (Apis mellifera L.). Mol Ecol, 7: 1543–1547.
Engel, M. S. (1999). The taxonomy of recent and fossil honey bees (Hymenoptera: Apidae; Apis). J Hym Res, 8: 165–196.
Eschscholtz, J. F. (1822). Entomographien. Reimer, Berlin.
Estoup, A., Garnery, L., Solignac, M. and Cornuet, J. M. (1995). Microsatellite variation in honey bee (Apis mellifera L.) populations: hierarchical genetic structure and test of the infinite allele and stepwise mutation models. Genetics, 140: 679–695.
Estoup, A., Largiadèr, C. R., Perrot, E. and Chourrout, D. (1996). Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol Mar Biol Biotech, 5: 295–298.
Felsenstein, J. (1993). PHYLIP (Phylogenetic inference package). Departement of Genetics, University of Washington, Seattle.
Fletcher, D. J. C. (1978). The African bee, Apis mellifera adansonii, in Africa. Ann Rev Entomol, 23: 151–171.
Franck, P. (1999). Approche génénique des questions évolutives associées à la sociobiologie et à la phylogéographie de l’abeille domestique (Apis mellifera L.). PhD Thesis, Ecole Nationale Superieure Agronomique, Montpellier.
Franck, P., Garnery, L., Solignac, M. and Cornuet, J. M. (1998). The origin of west European subspecies of honeybees (Apis mellifera): New insights from microsatellite and mitochondrial data. Evolution, 52: 1119–1134.
Franck, P., Garnery, L., Celebrano, G., Solignac, M. et al (2000a). Hybrid origins of the Italian honeybees, Apis mellifera ligustica and A. m. sicula. Mol Ecol, 9: 907–923, 10.1046/j.1365-294x.2000.00945.x.
Franck, P., Garnery, L., Solignac, M. and Cornuet, J. M. (2000b). Molecular confirmation of a fourth lineage in honeybees from Middle-East. Apidologie, 31: 167–180.
Garnery, L., Cornuet, J. M. and Solignac, M. (1992). Evolutionary history of the honey bee Apis mellifera inferred from mitochondrial DNA analysis. Mol Ecol, 1: 145–154.
Garnery, L., Solignac, M., Celebrano, G. and Cornuet, J. M. (1993). A simple test using restricted PCR-amplified mitochondrial DNA to study the genetic structure of Apis mellifera L. Experientia, 49: 1016–1021.
Garnery, L., Mosshine, E. H., Oldroyd, B. P. and Cornuet, J. M. (1995). Mitochondrial DNA variation in Moroccan and Spanish honey bee populations. Mol Ecol, 4: 465–471.
Garnery, L., Franck, P., Baudry, E., Vautrin, D. et al (1998). Genetic biodiversity of the West European honeybee (Apis mellifera mellifera and Apis mellifera iberica). I. Mitochondrial DNA. Génét Sél Évol, 30: 31–47.
Gasse, F. R., Tehet, A., Durand, A., Gilbert, E. et al (1990). The arid–humid transition in the Sahara and Sahel during the last glaciation. Nature, 346: 141–146.
Goldstein, D. B., Ruiz Linares, A., Cavalli-Sforza, L. L. and Feldman, M. W. (1995). Genetic absolute dating based on microsatellites and origin of modern humans. Proc Natl Acad Sci USA, 92: 6723–6727.
Hall, H. G. (1992). Further characterization of nuclear DNA RFLP markers that distinguish African and European honeybees. Arch Insect Biochem Physiol, 19: 163–175.
Hall, H. G. (1998). PCR amplification of a locus with RFLP allele specific to African honey bees. Biochem Genet, 36: 351–361.
Hall, H. G. and Muralidharan, K. (1989). Evidence from mitochondrial DNA that African honey bees spread as continuous maternal lineages. Nature, 339: 211–213.
Hall, H. G. and Smith, D. R. (1991). Distinguishing African and European honeybee matrilines using amplified mitochondrial DNA. Proc Natl Acad Sci USA, 88: 4548–4552.
Hamilton, A. C. (1982). Environmental History of East Africa: A Study of the Quaternary. Academic Press, London.
Hedges, S. B. (1992). The number of replication replications needed for accurate estimation of the bootstrap P-value in phylogenetic studies. Mol Biol Evol, 9: 366–369.
Hepburn, H. R. and Radloff, S. E. (1998). Honeybees of Africa. Springer, Berlin.
Hooghiemstra, H., Stalling, H., Agwu, C. O. C. and Dupont, L. M. (1992). Vegetation and climatic changes at the northern fringe of the Sahara 250,000–5,000 years BP. Rev Palaeobot Palynol, 74: 1–53.
Kerr, W. E. (1957). Introdução de abelhas Africanas no Brasil. Bras Apic, 8: 211–213.
Kerr, W. E. (1992). Abelhas Africanas su introduction y expansion en el continente Americano; Subspecies y ecotipas Africanos. Indust Apic, 13: 12–21.
Latreille, P. A. (1804). Notice des espèces d’abeilles vivant en grande société, ou abeilles proprement dites, et description d’espèces nouvelles. Ann Mus Natl Hist Nat, 5: 161–178.
Lepeletier, A. (1836). Histoire Naturelle des Insectes. Suites à Buffon. Hymenoptères. Roret, Paris.
Lioubimsteva, E. U. (1995). Ladscape changes in the Saharo–Arabian area during the last glacial cycle. J Arid Environ, 30: 1–17.
Lobo, J. A. and Krieger, H. (1992). Maximum likelihood estimates of gene frequencies and racial admixture in Apis mellifera L. (Africanized honeybees). Heredity, 68: 441–448.
Lovett, J. C. (1993a). Climatic history and forest distribution in eastern Africa. In: Lovett, J. C. and Wasser, S. K. (eds) Biogeography and Ecology of the Rainforest of Eastern Africa, pp. 23–29, Cambridge University Press, Cambridge.
Lovett, J. C. (1993b). Eastern arc moist forest flora. In: Lovett, J. C. and Wasser, S. K. (eds) Biogeography and Ecology of the Rainforest of Eastern Africa, pp. 33–55, Cambridge University Press, Cambridge.
Maa, T. C. (1953). An inquiry into the systematics of the tribus Apidini or honeybees (Hymenoptera). Treubia, 21: 525–640.
Mcmichael, M. and Hall, H. G. (1996). DNA RFLPs at a hightly polymorphic locus distinguish European and African subspecies of the honey bee Apis mellifera L. and suggest geographical origins of New World honey bees. Mol Ecol, 5: 403–416.
Meixner, M. D., Sheppard, W. S., Dietz, A. and Krell, R. (1994). Morphological and allozyme variability in honey bees from Kenya. Apidologie, 25: 188–202.
Meixner, M. D., Arias, M. C. and Sheppard, W. S. (2000). Mitochondrial DNA polymorphism in honeybee subspecies from Kenya. Apidologie, 31: 181–190.
Michener, C. D. (1975). The Brazilian bee problem. Ann Rev Entomol, 20: 399–416.
Moritz, R. F. A., Cornuet, J. M., Kryger, P., Garnery, L. et al (1994). Mitochondrial DNA variability in South African honeybees (Apis mellifera L.). Apidologie, 25: 169–178.
Nei, M. (1978). Estimation of average heterozygoty and genetic distance from a small number of individuals. Genetics, 89: 145–163.
Palmer, M. R., Smith, D. R. and Kaftanoglu, O. (2000). Turkish honeybees: genetic variation and evidence for a fourth lineage of Apis mellifera mtDNA. J Hered, 91: 42–46.
Potts, R. and Behrensmeyer, A. K. (1992). Late Cenozoic terrestrial ecosystems. In: Behrensmeyer, A. K., Damuth, J. D., Di Michele, W. A. et al. (eds) Terrestrial Ecosystems Through Time: Evolutionary Paleoecology of Terrestrial Plants and Animals, pp. 419–541, University of Chicago Press, Chicago.
Raymond, M. and Rousset, F. (1995). GENEPOP (version 1.2): population genetics software for exact test and ecumenism. J Hered, 86: 248–250.
Ruttner, F. (1976). Les races d’abeille de l’Afrique. In: XXV Congrès International d’Apiculture, pp. 347–367, Grenoble.
Ruttner, F. (1988). Biogeography and Taxonomy of Honeybees. Springer-Verlag, Berlin, Heidelberg.
Ruttner, F., Tassencourt, L. and Louveaux, J. (1978). Biometrical–statistical analysis of the geographic variability of Apis mellifera L. Apidologie, 9: 363–381.
Schiff, N. M. and Sheppard, W. S. (1993). Mitochondrial DNA evidence for 19th century introduction of African honey bees into the United States. Experientia, 49: 350–352.
Second, G. (1975). L’apiculture dans les pays d’afrique du nord, deuxième partie. Bull Techn Apic, 2: 9–20.
Sheppard, W. S., Soares, A. E. E. and de Jong, D. (1991). Hybrid status of honey bee populations near the historic origin of the Africanization in Brazil. Apidologie, 22: 643–652.
Sheppard, W. S., Arias, M. C., Grech, A. and Meixner, M. D. (1997). Apis mellifera ruttneri, a new honey bee subspecies from Malta. Apidologie, 28: 287–293.
Sheppard, W. S., Rinderer, T. E., Garnery, L. and Shimanuki, H. (1999). Analysis of Africanized honey bee mitochondrial DNA reveals further diversity of origin. Genet Mol Biol, 22: 73–75.
Sinacori, A., Rinderer, T. E., Lancaster, V. and Sheppard, W. S. (1998). A morphological and mitochondrial assessment of Apis mellifera from Palermo, Italy. Apidologie, 29: 481–490.
Smith, F. G. (1961). The races of honeybees in Africa. Bee World, 42: 255–260.
Smith, D. R. (1991). African bees in the Americas: insights from biogeography and genetics. Trends Ecol Evol, 6: 17–21.
Smith, D. R. and Brown, W. M. (1988). Polymorphisms in mitochondrial DNA of European and Africanized honeybees (Apis mellifera). Experientia, 44: 257–260.
Smith, D. R., Taylor, O. R. and Brown, W. M. (1989). Neotropical Africanized honeybees have African mitochondrial DNA. Nature, 339: 213–215.
Smith, D. R., Palopoli, M. F., Taylor, B. R., Garnery, L. et al (1991). Geographical overlap of two mitochondrial genomes in Spanish honeybees (Apis mellifera iberica). J Hered, 82: 96–100.
Acknowledgements
We gratefully thank R. Z. Ramamonjisoa, C. Kerdelué, M. Harry, D. Lachaise, E. Franck, N. Koeninger, G. Lanher, R. Crew, L. Gaume, R. Vandame, Y. Garba, S. Laouali for their help in collecting samples and J.-Y. Rasplus for useful comments on the biogeography of insects in Africa.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Franck, P., Garnery, L., Loiseau, A. et al. Genetic diversity of the honeybee in Africa: microsatellite and mitochondrial data. Heredity 86, 420–430 (2001). https://doi.org/10.1046/j.1365-2540.2001.00842.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00842.x
Keywords
This article is cited by
-
First evaluation of genetic diversity among honeybee populations in Kazakhstan
Apidologie (2023)
-
Molecular analyses of mitochondrial DNA reveal new haplotypes and lineages within Ethiopian honeybees (Apis mellifera)
International Journal of Tropical Insect Science (2023)
-
Genetic Characterizations of the Iranian Honey Bee (Apis mellifera meda Skorikov 1929) Populations Using the Microsatellite DNA Markers
Biochemical Genetics (2023)
-
Genetic diversity and population structure of the Saharan honey bee Apis mellifera sahariensis from southeastern Morocco: introgression assessment and implications for conservation
Apidologie (2023)
-
The post-capping period of the tropical honey bee subspecies Apis mellifera unicolor in La Réuion
Apidologie (2023)