Abstract
One to three accessions of 22 Alstroemeria species, an interspecific hybrid (A. aurea × A. inodora), and single accessions of Bomarea salsilla and Leontochir ovallei were evaluated using the AFLP-marker technique to estimate the genetic diversity within the genus Alstroemeria. Three primer combinations generated 716 markers and discriminated all Alstroemeria species. The dendrogram inferred from the AFLP fingerprints supported the conjecture of the generic separation of the Chilean and Brazilian Alstroemeria species. The principal co-ordinate plot showed the separate allocation of the A. ligtu group and the allocation of A. aurea, which has a wide range of geographical distribution and genetic variation, in the middle of other Alstroemeria species. The genetic distances, based on AFLP markers, determined the genomic contribution of the parents to the interspecific hybrid.
Similar content being viewed by others
Introduction
The genus Alstroemeria includes approximately 60 described species of rhizomatous, herbaceous plants, with Chile and Brazil as the main centres of diversity (Uphof, 1952; Bayer, 1987; Aker & Healy, 1990). The Chilean and Brazilian Alstroemeria are recognized as representatives of different branches of the genus. The family of Alstroemeriaceae, to which Alstroemeria belongs, includes several related genera, such as Bomarea Mirbel, the monotype Leontochir ovallei Phil. and Schickendantzia Pax (Dahlgren & Clifford, 1982; Hutchinson, 1973).
The species classification in Alstroemeria is based on an evaluation of morphological traits of the flower, stem, leaf, fruit and rhizome (Bayer, 1987). The available biosystematic information on Alstroemeria species is restricted to the Chilean species, as described in the monograph of Bayer (1987). Little is known about the classification of the Brazilian species (Meerow & Tombolato, 1996). Furthermore, morphology-based identification is rather difficult because morphological characteristics can vary considerably in different environmental conditions (Bayer, 1987).
The immense genetic variation present in the genus Alstroemeria offers many opportunities for the improvement and renewal of cultivars. Therefore, identification of genetic relationships at the species level could be very useful for breeding in supporting the selection of crossing combinations from large sets of parental genotypes, thus broadening the genetic basis of breeding programmes (Frei et al., 1986). The species used in the study reported here are commonly used in the breeding programme of Alstroemeria for cut flowers and pot plants.
Molecular techniques have become increasingly significant for biosystematic studies (Soltis et al., 1992). RAPD markers were used for the identification of genetic relationships between Alstroemeria species and cultivars (Anastassopoulos & Keil, 1996; Dubouzet et al., 1997; Picton & Hughes, 1997). In recent years a novel PCR-based marker technique, AFLP (Vos et al., 1995), has been developed and used for genetic studies in numerous plants including lettuce (Hill et al., 1996), lentil (Sharma et al., 1996), bean (Tohme et al., 1996), tea (Paul et al., 1997), barley (Schut et al., 1997), and wild potato species (Kardolus et al., 1998). These studies indicated that AFLP is highly applicable for molecular discrimination at the species level. The technique has also been optimized for use in species such as Alstroemeria spp., which are characterized by a large genome size (2C-value: 37–79 pg) (Han et al., 1999).
In this study, we produced AFLP fingerprints of 22 Alstroemeria species, one interspecific hybrid (A. aurea × A. inodora) and the distantly related species Bomarea salsilla and Leontochir ovallei, and we analysed their genetic relationships. The interspecific hybrid was included in our study in order to investigate the possibility of identifying the parental genotypes.
Materials and methods
Plant material
Seeds and plants of 22 Alstroemeria species were obtained from botanical gardens and commercial breeders. The collection has been maintained for many years in the greenhouse of Unifarm at the Wageningen Agricultural University. When available, three accessions were selected for each Alstroemeria species, and both B. salsilla and L. ovallei were chosen as outgroups. One interspecific hybrid (A. aurea × A. inodora) was obtained from earlier research (Buitendijk et al., 1995) (Table 1). All accessions were identified according to their morphological traits (Uphof, 1952; Bayer, 1987).
AFLP protocol
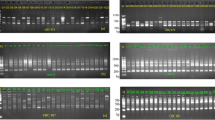
Genomic DNA was isolated from young leaves of greenhouse-grown plants using the cetyltrimethy-lammonium bromide (CTAB) method according to Rogers & Bendich (1988). The AFLP technique followed the method of Vos et al. (1995) with modifications of selective bases of pre- and final amplifications (Han et al., 1999). To assess interspecific variation, autoradiograms comprising the AFLP fingerprints of a mixture of three accessions per species were analysed by pooling 5 μL of the final selective amplification products according to Mhameed et al. (1997). The low level of variation between individual samples showed that pooling accessions was justified. Three primer combinations (E + ACCA/M + CATG, E + ACCT/M + CATC and E + AGCC/M + CACC) were selected from a test of 96 primer combinations, and these produced 272, 211 and 233 bands, respectively (Table 2). The choice of the primers used in the study was based upon the visual clarity of banding patterns generated and a preferably low fingerprint complexity. The complexity of the banding pattern is a major limiting factor for scoring AFLP fingerprints of large-size genomes.
Data analysis
Positions of unequivocally visible and polymorphic AFLP markers were transformed into a binary matrix, with ‘1’ for the presence, and ‘0’ for the absence of a band at a particular position. The genetic distance (GD) between species was based on pair-wise comparisons and calculated according to the equation: GDxy=1 − [2Nxy/(Nx + Ny)], where Nx and Ny are the numbers of fragments to individuals x and y, respectively, and Nxy is the number of fragments shared by both (Nei & Li, 1979). Genetic distances were computed by the software package TREECON (v. 1.3b) (Van De Peer & De Wachter, 1993). The dendrogram of the 22 Alstroemeria species, the interspecific hybrid, Bomarea and Leontochir was generated based on the GD matrix by using cluster analysis, the UPGMA (unweighted pair group method using arithmetic averages) method with 1000 bootstraps (Sneath & Sokal, 1973; Felsenstein, 1985) (Fig. 1). Principal co-ordinate analysis was performed to access interspecies relationships based on the Nei & Li (1979) coefficient [2Nxy/(Nx + Ny)] using the NTSYS-PC program (Rohlf, 1989).
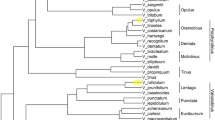
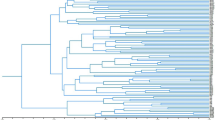
Dendrogram of 22 Alstroemeria species, Bomarea salsilla and Leontochir ovallei resulting from a UPGMA cluster analysis based on Nei’s genetic distances obtained from 716 AFLP bands. The bootstrap analysis was conducted using TREECON (v. 1.3b) with 1000 bootstrap subsamples of the data matrix. Percentage values for those branches occurring in at least 60% of the bootstrap topologies are shown.
Results and discussion
The average genetic distance among species excluding Bomarea, Leontochir, the interspecific hybrid and A. umbellata was 0.65 GD (a table showing the genetic distances between all the species studied is available from the authors on request). Alstroemeria umbellata was excluded because the accessions used were found to be highly related and possibly wrongly classified as different from A. pelegrina. The average GD among accessions within a species was 0.32 GD (data not shown). In addition, the average GD between Brazilian species (GD: 0.27) and between Chilean species (GD: 0.33) was not significantly different.
Buitendijk & Ramanna (1996) suggested that the Chilean and Brazilian species form distinct lineages. The genetic diversification of Alstroemeria species as detected by the AFLP technique revealed three main clusters with 99% bootstrap values: the Chilean species, the Brazilian species and the outgroup (Fig. 1). This finding would support an early divergence of these groups and is consistent with the occurrence of interspecific crossing barriers between the Chilean and Brazilian species (De Jeu & Jacobsen, 1995; Lu & Bridgen, 1997). The variance of the first three principal co-ordinates accounted for 34.9% of the total variation, differentiated effectively among the species and reflected the main clustering of the dendrogram. From the principal co-ordinate plot, four groups were clearly demarcated: (i) the Brazilian group; (ii) the Chilean group; (iii) the A. ligtu group; and (iv) the outgroup (Fig. 2). The Brazilian species (A. brasiliensis, A. psittacina and A. inodora) were consistently assigned to one cluster with 98% bootstrap values, whereas the Chilean species were rather weakly clustered with 62% bootstrap values containing several subgroups within the Chilean group (Fig. 1 and 2). The dispersion of the Chilean species on the principal co-ordinate plot reflected a wider genetic variation than the Brazilian species. However, the narrow variation of the Brazilian species might be caused by the limited number of species investigated.
Relationships among 22 Alstroemeria species, the F1 hybrid, Bomarea salsilla and Leontochir ovallei by principal co-ordinate analysis using Nei and Li coefficients. The three principal co-ordinates accounted for 34.9% of the total variation. PC1, PC2 and PC3: first, second and third principal co-ordinates. See Table 1 for species names.
Buitendijk & Ramanna (1996) described the similarities between C-banding patterns of A. inodora and A. psittacina; in our study these species clustered strongly, reinforcing this finding (Fig. 1). The similarity between A. psittacina and A. inodora was also revealed by allozyme analysis (Meerow & Tombolato, 1996) and by a study using species-specific repetitive probes (De Jeu et al., 1995). These findings are also supported by the fact that A. inodora and A. psittacina are easily crossed (De Jeu & Jacobsen, 1995).
In addition, the Chilean species A. aurea was positioned between three subgroups (Fig. 2). The unique position of A. aurea, and the observation that this species has a wide geographical spread, suggest that other Chilean species may have evolved from A. aurea ecotypes. Alstroemeria aurea is indeed a widespread inhabitant in the regions with higher rainfall at the more southern latitudes between 33 and 47°S in Chile (Bayer, 1987; Buitendijk & Ramanna, 1996). It is not found in Brazil, although A. aurea plants are found on both sides of the Andes mountains in Argentina, supporting the possibility that A. aurea ecotypes were also the ancestors of the Brazilian species (A.F.C. Tombolato, personal communication).
Alstroemeria pelegrina and A. umbellata were assigned as sister species with a GD of 0.26 showing a remarkable genetic similarity (data available on request). The species we coded under the name A. umbellata actually seemed to be an A. pelegrina species that did not flower for many years. Alstroemeria haemantha was assigned to a group together with A. ligtu ssp. ligtu, A. ligtu ssp. incarnata and A. ligtu ssp. simsii (Figs 1 and Fig. 2) (Aker & Healy, 1990; Ishikawa et al., 1997). Bayer (1987) suggested the synonymous name of A. ligtu ssp. ligtu for A. haemantha Ruiz and Pavon. Our results support this hypothesis. Alstroemeria exserens was positioned between the Chilean group and the A. ligtu group (Fig. 2). Alstroemeria andina and A. angustifolia ssp. angustifolia, and A. hookeri ssp. cumminghiana and A. hookeri ssp. hookeri were clustered together with 95% and 93% bootstrap values, respectively.
The interspecific hybrid (A1P2–2) was included in our study in order to investigate the possibility of the identification of the parental genotypes. The F1 hybrid A1P2–2 showed a 0.45-GD value with A. inodora and 0.59 GD value with A. aurea showing genomic contribution of both parents (data available on request). It indicated the feasibility of the AFLP technique as a tool for the identification of parental genotypes (Sharma et al., 1996; Marsan et al., 1998). Bomarea and Leontochir showed the mean GD value of 0.83 as the outgroup, thus showing large genetic distances within the Alstroemeriaceae family.
In conclusion, the genetic variation and the genetic relationships among Alstroemeria species were efficiently rationalized by using AFLP markers for the characterization of germplasm resources. In general, the topologies of the dendrogram and the principal co-ordinate analysis of our study were in agreement with Bayer’s views (Bayer, 1987) on the classification of the Alstroemeria species. Furthermore, this technique might be useful for the identification of parental genotypes in interspecific hybrids.
References
Aker, S. and Healy, W. (1990). The phytogeography of the genus Alstroemeria. Herbertia 46: 76–87.
Anastassopoulos, E. and Keil, M. (1996). Assessment of natural and induced genetic variation in Alstroemeria using random amplified polymorphic DNA (RAPD) markers. Euphytica 90: 235–244.
Bayer, E. (1987). Die Gattung Alstroemeria in Chile. Mitt Bot Staatsamml München, 24: 1–362.
Buitendijk, J. H. and Ramanna, M. S. (1996). Giemsa C-banded karyotypes of eight species of Alstroemeria L. and some of their hybrids. Ann Bot, 78: 449–457.
Buitendijk, J. H., Pinsonneaux, N. A. C., van Donk, M. S. and Lammeren, A. A. M. (1995). Embryo rescue by half-ovule culture for the production of interspecific hybrids in Alstroemeria. Sci Hortic, 64: 65–75.
Dahlgren, R. M. T. and Clifford, H. T. (1982). Monocotyledons A Comparative Study. Academic Press, London.
de Jeu, M. J. and Jacobsen, E. (1995). Early postfertilization ovule culture in Alstroemeria L. and barriers to interspecific hybridization. Euphytica 86: 15–23.
de Jeu, M. J., Lasschuit, J., Chevalier, F. and Visser, R. G. F. (1995). Hybrid detection in Alstroemeria by use of species-specific repetitive probes. Acta Hortic, 420: 62–64.
Dubouzet, J. G., Murata, N. and Shinoda, K. (1997). RAPD analysis of genetic relationships among Alstroemeria L. cultivars. Sci Hortic, 68: 181–189.
Felsenstein, J. (1985). Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783–791.
Frei, O. M., Stuber, C. W. and Goodman, M. M. (1986). Use of allozymes as genetic markers for predicting performance in maize single cross hybrids. Crop Sci, 26: 37–42.
Han, T. H., van Eck, H. J., de Jeu, M. J. and Jacobsen, E. (1999). Optimization of AFLP fingerprinting of organisms with a large genome size: a study on Alstroemeria spp. Theor Appl Genet, 98: 465–471.
Hill, M., Witsenboer, H., Zabeau, M., Vos, P., Kesseli, R. and Michelmore, R. (1996). PCR-based fingerprinting using AFLPs as a tool for studying genetic relationships in Lactuca spp. Theor Appl Genet, 93: 1202–1210.
Hutchinson, J. (1973). The Families of Flowering Plants. Clarendon Press, Oxford.
Ishikawa, T., Takayama, T., Ishizaka, H., Ishikawa, K. and Mii, M. (1997). Production of interspecific hybrids between Alstroemeria ligtu L. hybrid and A. pelegrina L. var. rosea by ovule culture. Breed Sci, 47: 15–20.
Kardolus, J. P., van Eck, H. J. and van Den Berg, R. G. (1998). The potential of AFLPs in biosystematics: a first application in Solanum taxonomy. Pl Syst Evol, 210: 87–103.
Lu, C. and Bridgen, M. P. (1997). Chromosome doubling and fertility study of Alstroemeria aurea × A. caryophyllaea. Euphytica 94: 75–81.
Marsan, P. A., Castiglioni, P., Fusari, F., Kuiper, M. and Motto, M. (1998). Genetic diversity and its relationship to hybrid performance in maize as revealed by RFLP and AFLP markers. Theor Appl Genet, 96: 219–227.
Meerow, A. W. and Tombolato, A. F. C. (1996). The Alstroemeria of Itatiaia. Herbertia. 51: 14–21.
Mhameed, S., Sharon, D., Kaufman, D., Lahav, E., Hillel, J., Degani, C. and Lavi, U. (1997). Genetic relationships within avocado (Persea americana Mill.) cultivars and between Persea species. Theor Appl Genet, 94: 279–286.
Nei, M. and Li, W. H. (1979). Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA, 76: 5269–5273.
Paul, S., Wachira, F. N., Powell, W. and Waugh, R. (1997). Diversity and genetic differentiation among populations of Indian and Kenyan tea (Camellia sinensis (L.) O. Kuntze) revealed by AFLP markers. Theor Appl Genet, 94: 255–263.
Picton, D. D. and Hughes, H. G. (1997). Characterization of Alstroemeria species using Random Amplified Polymorphic DNA (RAPD) analysis. HortScience 32: 482–482 Abstract: 323.
Rogers, S. O. and Bendich, A. J. (1988). Extraction of DNA from plant tissues. Plant Mol Biol Manual, 6: 1–10.
Rohlf, F. J. (1989). NTSYS-Pc Numerical Taxonomy and Multivariate Analysis System, version 1.80. Exeter Publications, New York, NY.
Schut, J. W., Qi, X. and Stam, P. (1997). Association between relationship measures based on AFLP markers, pedigree data and morphological traits in barley. Theor Appl Genet, 95: 1161–1168.
Sharma, S. K., Knox, M. R. and Ellis, T. H. (1996). AFLP analysis of the diversity and phylogeny of Lens and its comparison with RAPD analysis. Theor Appl Genet, 93: 751–758.
Sneath, P. H. A. and Sokal, R. R. (1973). Numerical Taxonomy. W.H. Freeman, San Francisco, CA.
Soltis, P. S., Soltis, D. E. and Doyle, J. J. (1992). Molecular Systematics of Plants. Chapman & Hall, New York, NY.
Tohme, J., Gonzalez, D. O., Beebe, S. and Duque, M. C. (1996). AFLP analysis of gene pool of a wild bean core collection. Crop Sci, 36: 1375–1384.
Uphof, J. C. T. (1952). A review of the genus Alstroemeria. Plant Life. 8: 37–53.
van Depeer, Y. and de Wachter, R. (1993). Treecon: a software package for the construction and drawing of evolutionary trees. Comput Applic Biosci. 9: 177–182.
Vos, P., Hogers, R., Bleeker, M., Reijans, M., Vandelee, T., Hornes, M. et al (1995). AFLP: a new technique for DNA fingerprinting. Nucl Acids Res, 23: 4407–4414.
Wilkin, P. (1997). Leontochir ovallei Alstroemeriaceae. Curtis’s Bot Magazine. 14: 7–12.
Acknowledgements
The authors would like to thank Anja G.J. Kuipers and Jaap B. Buntjer for critical reading of the manuscript and for helpful comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Han, TH., de Jeu, M., van Eck, H. et al. Genetic diversity of Chilean and Brazilian Alstroemeria species assessed by AFLP analysis. Heredity 84, 564–569 (2000). https://doi.org/10.1046/j.1365-2540.2000.00682.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00682.x
Keywords
This article is cited by
-
Cytogenetics of wild species of the Alstroemeriaceae family (Liliales)
Plant Systematics and Evolution (2021)
-
AFLP analysis of genetic variation among three natural populations of horseshoe crab Tachypleus tridentatus along Chinese coast
Chinese Journal of Oceanology and Limnology (2011)
-
Application of AFLP markers for population genetic study on half-smooth tongue sole Cynoglossus semilaevis
Chinese Journal of Oceanology and Limnology (2011)
-
Assessment of genetic diversity through RAPD, ISSR and AFLP markers in Podophyllum hexandrum: a medicinal herb from the Northwestern Himalayan region
Physiology and Molecular Biology of Plants (2010)
-
Analysis of genetic diversity in Glyptosternum maculatum (Sisoridae, Siluriformes) populations with AFLP markers
Environmental Biology of Fishes (2009)