Abstract
Root-knot nematodes (RKN) are sedentary endoparasites causing severe damage to a wide variety of crops, including tomato. Among them, the parthenogenetic species Meloidogyne arenaria, M. incognita and M. javanica are of particular economic importance. The genetic diversity and relationships of 17 populations belonging to these three major species, either avirulent or virulent against the tomato Mi resistance gene, were investigated in order to determine whether (a)virulence of the nematodes could be related to their molecular fingerprints. Genomic polymorphisms between populations were assessed by using amplified fragment length polymorphism (AFLP) markers, and data were treated by means of a multiple correspondence analysis. A total of 1550 polymorphic amplified DNA fragments were identified and used to compute the relationships between the populations. As expected, the three RKN species were clearly distributed into distinct groups, but combination of data for virulence phenotypes and DNA markers showed that clustering of populations was not associated with their (a)virulence against the tomato Mi resistance gene. Such a lack of correlation indicates that most of the observed DNA polymorphism is independent of virulence, which is presumably under host selection. This result demonstrates that virulent populations do not share a common origin, and strongly suggests that they might have appeared late after the establishment of these clonal lineages, as the result of independent mutational events.
Similar content being viewed by others
Introduction
Root-knot nematodes (RKN) belonging to the genus Meloidogyne cause extensive damage to a wide variety of economically important crops (Sasser, 1980). Among them, M. incognita (Kofoid & White) Chitwood, M. javanica (Treub) Chitwood and M. arenaria (Neal) Chitwood, are considered the three major species on the basis of both their worldwide geographical distribution and very large host range (Lamberti, 1979). Of special interest also is the fact that these three Meloidogyne species reproduce exclusively by mitotic parthenogenesis (Triantaphyllou, 1985), thus theoretically leading to genetically identical offspring, apart from random distribution of mutational events and multiple ploidy levels.
The availability of many resistance genes in both annual and perennial crops has contributed to making plant resistance the most effective and environmentally safe method to control RKN (Roberts, 1992). However, the occurrence of virulent populations, able to reproduce on resistant plants, constitutes a severe threat to this strategy. This is particularly true for tomato, where resistance to M. incognita, M. javanica and M. arenaria is thought to be controlled by a single dominant gene designated Mi (Gilbert & McGuire, 1956), and for which resistance-breaking populations have recently been reported from many places in the world (Kaloshian et al., 1996; Tzortzakakis & Gowen, 1996; Eddaoui et al., 1997). Thus, identification of additional resistance genes and their incorporation into domestic tomato is urgently needed for the development of more durable resistant cultivars.
Because breeding for resistance strongly depends on the differential virulence of the populations used to evaluate the plant materials to be tested, such information is obviously needed for Meloidogyne species. However, the genetic nature of the relationships among RKN populations that are avirulent and virulent against the Mi gene remains poorly understood. It is still unknown whether the virulent populations share some common genetic background or, conversely, if they are dispersed at random among an array of isolated and clonal avirulent genotypes. This issue is of both theoretical and practical importance. In the first instance, this system may provide new insights into the evolution of (a)virulence within a parthenogenetic organism. From a practical point of view, understanding the relationships between populations with different parasitic abilities would contribute to the development of new resistant cultivars and new strategies for managing RKN (i.e. monitoring of virulence in field populations). For the study reported here we assembled a collection of M. arenaria, M. incognita and M. javanica populations, either avirulent or virulent against the tomato Mi resistance gene, and their molecular diversity was assessed by AFLP analysis. The objectives of this research were to study the molecular diversity of these populations in order to determine whether some association could be found between the molecular fingerprints and the (a)virulence phenotypes in RKN.
Materials and methods
Nematode populations
Five populations of M. arenaria, six populations of M. incognita and six populations of M. javanica were tested for their genomic variation (Table 1). All the nematode populations used in this study were sampled from heavily infested fields or glasshouses, in areas where resistant tomatoes reportedly have not been cultured. From each population, a clonal line was raised starting from the progeny of a single female, in order to eliminate the potential within-population heterogeneity. Prior to multiplication, each species was identified according to its isoesterase electrophoretic pattern (Dalmasso & Bergé, 1978).
Virulence test
Two cultivars of Lycopersicon esculentum were used: Saint Pierre, a susceptible control and the near-isogenic Piersol, which carries the Mi gene for RKN resistance (Laterrot, 1975). All experiments were conducted in a climate-controlled room at a mean temperature of 20°C and 16 h day length. The reproduction of each nematode population was evaluated on both susceptible and resistant tomato cultivars using a previously described test tube culture and assay (Castagnone-Sereno et al., 1993). Each tomato plant was inoculated with 25 second-stage juveniles of each nematode genotype. Plants were arranged in a randomized complete block design with 10 replicates for each nematode × plant combination tested and the experiment was repeated once.
Eight weeks after inoculation, the washed root systems were placed in cold yellow eosin solution (0.1 g/L of water) and stirred for 30 min to stain the egg masses. The numbers of egg masses per root system were counted and a reproductive index (Ri) was calculated according to the following ratio: Ri = number of egg masses obtained/number of juveniles inoculated. Theoretically, Ri ranges from 0 (no reproduction at all) to 1 (each juvenile inoculated gave one egg mass). A two-way analysis of variance (ANOVA) was performed on the data, with Meloidogyne populations and tomato cultivars as factors and Ri values as variable. Means were separated with Fisher’s protected least significant difference test (PLSD, at P =0.05) to compare rates of reproduction between populations. All the statistical work was carried out using the STATVIEW software.
Genomic DNA isolation
Total DNA was extracted from 200 to 300 μL of nematode eggs of each population that had been stored at −80°C before use. Eggs were ground in liquid nitrogen with a mortar and pestle. The DNA was purified from the resulting powder by a phenol-chloroform extraction (Sambrook et al., 1989). Following ethanol precipitation, DNA was resuspended in TE buffer (0.01 M Tris pH 8.0, 0.001 M EDTA).
AFLP procedure
The AFLP procedure was performed essentially as previously described (Semblat et al., 1998), but including a number of minor modifications.
DNA restriction and ligation of adapters. Adapters used in this study were obtained from Eurogentec (Herstal, Belgium). Their sequences were as follows:
HindIII-adapter: 5′-CTCGTAGACTGCGTACC-3′ 3′-CATCTGACGCATGGTCGA-5′
MseI-adapter: 5′-GACGATGAGTCCTGAG-3′ 3′-TACTCAGGACTCAT-5′
One μg of total genomic DNA was digested at 37°C for 3 h with 10 U of HindIII and 10 U of MseI in RL buffer (10 mM Tris pH 7.5, 10 mM MgAc, 50 mM KAc, 5 mM DTT, 50 ng μL–1 BSA) in a final volume of 80 μL. Then, 50 μL of the digestion was mixed with 10 μL of a solution containing 5 pmoles of the HindIII-adapter, 50 pmoles of the MseI-adapter, 1 U of T4 DNA ligase, 1 mM ATP in RL buffer and incubated for 3 h more at 37 °C. After ligation, the reaction mixture was diluted tenfold in TE buffer and stored at −20°C.
Selective preamplification. Primers used in selective preamplification were 5′-GACTGCGTACCAGCTTA-3′ (H + A) and 5′-GATGAGTCCTGAGTAAA-3′ (M + A), which are, respectively, complementary to the core sequence of the HindIII-adapter and the MseI-adapter. Five μL of the ligated fragments were amplified with 75 ng of H + A and 75 ng of M + A primers, 0.2 mM of all four dNTPs, 0.5 U of Taq DNA polymerase in 1 × Taq buffer (Appligene, Illkirch, France) in a final volume of 50 μL. Amplification was performed for 28 cycles according to the following cycle programme: 1 min at 94°C, 1 min 20 s at 60°C, 1 min at 72°C. Preamplification products were diluted fivefold in TE buffer.
Selective amplification. Primers used in the selective amplification step were H + AAA and H + AAC in combination with M + AAA, M + AAC, M + AAG, M + AAT, M + ACA, M + ACC, M + ACG, M + ACT, M + AGA, M + AGC, M + AGG, M + AGT, M + ATA, M + ATC, M + ATG, M + ATT. Five ng of the Hind primers were labelled using 1 μCi of 33P-dATP and 0.2 U of T4 polynucleotide kinase for 45 min at 37°C and 15 min at 70°C. Amplification of 1 μL of diluted products from the selective preamplification was performed as described above using 5 ng of labelled primer, 30 ng of Mse primer and 0.2 U of Taq in a final volume of 20 μL. The temperature profile used was: 1 min at 94 °C, 1 min 20 s at 65 °C and 1 min at 72 °C. The annealing temperature was decreased each cycle by 0.7 °C for 12 cycles and was continued at 56 °C for the remaining 24 cycles.
Gel analysis. The amplification products were mixed with an equal volume of formamide dye (98% formamide, 10 mM EDTA, 0.025% each of xylene cyanol and bromophenol blue) and denatured for 3 min at 90°C. Five μL were loaded on a 5% polyacrylamide gel and electrophoresed in 1 × TBE buffer for 2 h at 50 W. Following transfer on Whatmann 3MM paper, gels were dried and X-ray films were exposed for 2–3 days at room temperature.
Data analysis
Bands were scored as present (1) or absent (0) directly from the autoradiographs. Experiments were repeated at least once, and only DNA fragments consistently present or absent between repeats were recorded and considered as binary characters. DNA fingerprints from each population were converted to a 0–1 matrix, and these data were treated by a multiple correspondence analysis (MCA) (Benzecri, 1973; Greenacre & Hastie, 1987), using the PROC CORRESP procedure in the SAS/STAT package (SAS Institute Inc., 1990). This method consists of a weighted principal component analysis of a multiway contingency table. It is used to find a low-dimensional graphical representation of the association between rows (nematode populations) and columns (AFLP data) of the 0–1 matrix in a Euclidian space. In the current study, two populations sharing close coordinates for one axis will be closely related based on AFLP signals contributing to the determination of that axis.
Results
Virulence variation
Both susceptible (Saint Pierre) and Mi-resistant (Piersol) tomato cultivars were inoculated with the 17 RKN field populations, and the resulting egg masses present on the roots were counted 2 months later. Because no significant differences were observed between the repeated experiments, results were pooled together. The results of the ANOVA (Table 2) showed the existence of a strong influence of both the nematode population (P< 0.0001) and the tomato cultivar (P< 0.0001) on the numbers of egg masses observed. Moreover, a significant interaction occurred between the two factors (P< 0.0001), which indicated a differential reproductive ability of the nematode populations tested on each tomato genotype. The global reproduction data are reported in Table 3. As expected, all Meloidogyne populations were able to reproduce on the susceptible tomato Saint Pierre, with Ri values ranging from 0.736 to 0.988. Conversely, Ri values obtained on the resistant cultivar Piersol separated populations which were virulent against the Mi gene (Ri values ≥ 0.785) and populations which were avirulent (Ri values ≤ 0.020). The virulence of two M. arenaria populations (ma1 and ma3), three M. incognita populations (mi2, mi4 and mi6), and three M. javanica populations (mj3, mj5 and mj6) was demonstrated by their strong ability to reproduce on the resistant cultivar, with Ri values in the ranges 0.798–0.855, 0.884–0.962 and 0.785–0.965, respectively. The remaining avirulent populations appeared largely unable to reproduce on Piersol, with Ri values ranging from 0 to 0.020.
DNA variation revealed by AFLPs
The specific status of each nematode population was determined on the basis of isoesterase banding patterns, and no infraspecific variation could be found within any of the three RKN species tested (Fig. 1).
Isoesterase phenotypes of the 17 Meloidogyne populations used in this study. Population codes are given in Table 1.
AFLP fingerprinting of the 17 Meloidogyne populations with 32 primer combinations revealed a total number of 2104 reproducible amplified DNA fragments, of which 1550 were polymorphic (on the basis of their presence/absence) among all RKN populations. Depending on the primer combination and the template used, 33 to 114 bands were observed, with the number of polymorphic fragments ranging from 19 to 53. The size of the amplified DNA fragments ranged from 50 to 600 bp. Typical examples of AFLP fingerprints for the 17 populations are shown in Fig. 2, the first one with primer M + AGG (Fig. 2a) and the second one with primer M + AGT (Fig. 2b), both in combination with primer H + AAA.
AFLP fingerprints of 17 Meloidogyne populations after selective amplification with primer combination H + AAA/M + AGG (a) or H + AAA/M + AGT (b). Population codes are given in Table 1.
Each primer combination used in this study unambiguously allowed the differentiation of the three species tested. Among the polymorphic fragments, 197 appeared as species-specific characters: 20, 122 and 55 for M. arenaria, M. incognita and M. javanica, respectively. However, notable differences in polymorphism were observed within each of the three species. In M. arenaria, 798 amplified fragments were polymorphic. Significantly fewer polymorphic bands were amplified for both M. incognita and M. javanica, with 128 and 111 polymorphic fragments, respectively. Conversely to M. arenaria, most of these bands (112 and 99, respectively) were shared by at least two populations within one species. Using appropriate primer combination(s), each of the 17 RKN populations could be easily differentiated from all the others.
Results of MCA
Because we did not have any information about the allelic state of each band scored, we could not rule out the hypothesis that some markers are linked. For that reason, we analysed the AFLP data by the descriptive MCA method, the pertinence of which has been established previously for polymorphism studies (Erselius & de Valavieille, 1984; Colas et al., 1998).
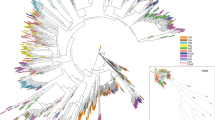
The distribution of the 17 Meloidogyne spp. populations in the factorial planes [1,2] and [3,4] is presented in Fig. 3. Each of the first four axes explained 41.7, 15.3, 11.8 and 8.6% of the total variability, respectively. Because they resulted together in an assessment of 77.4% of the total variability, the analysis was therefore limited to the four first axes.
Distribution of 17 Meloidogyne populations according to the multiple correspondence analysis (MCA) conducted on AFLP data. Both factorial planes [1,2] (a) and [3,4] (b) are represented. ▵ = M. arenaria; ○ = M. incognita; □ = M. javanica. White symbols correspond to avirulent populations, black symbols correspond to virulent populations. Population codes are given in Table 1.
The three RKN species clearly belong to distinct groups in the factorial plane [1,2], which explained 57.0% of the total variability: coordinates on axis 1 separated M. incognita populations from those belonging to the two other species, whereas coordinates on axis 2 separated M. arenaria from M. javanica populations (Fig. 3a). Conversely to M. arenaria populations, which appeared weakly clustered together, populations from M. incognita on one side, and from M. javanica on the other side, generally shared very close coordinates in the factorial plane [1,2]. In the factorial plane [3,4], which explained 20.4% of the total variability, all the populations were concentrated around the origin of both axes, except for M. arenaria population ma5, M. incognita population mi6 and M. javanica population mj6 (Fig. 3b).
Within and among each of the three RKN species, virulent populations were not clearly differentiated from the avirulent ones, but rather appeared randomly distributed in the factorial planes [1,2] and [3,4]. In a large number of occurrences, avirulent and virulent populations even shared very close coordinates, and therefore no subgroups could be distinguished in the Euclidian space (Fig. 3). From these data, it can be concluded that no correlation could be achieved between (a)virulence of the populations and the AFLP markers resolved.
Discussion
The recent emergence of virulent nematode populations on resistant cultivars is becoming a cause of concern, because they could significantly reduce the durability of the resistance genes used. The purpose of this work was to study the genetic diversity and potential relationships of M. arenaria, M. incognita and M. javanica avirulent and virulent populations, in order to investigate the origin of their virulence against the tomato Mi resistance gene. Genomic fingerprints of the 17 populations tested were carried out using AFLP markers, and the data were analysed with a MCA. Clearly, our results showed that clustering of RKN populations was not associated with their (a)virulence against the tomato Mi resistance gene. Within each specific group, avirulent and virulent populations were not distributed into separate subgroups. Thus, it seems unlikely that virulent populations share a common origin, either within RKN as a whole, or within any of the three species tested here. This lack of correlation between groups determined by molecular markers and virulence indicates that most of the observed DNA polymorphism is largely independent of virulence, which is under host selection. Independence between virulence and phenetic relationships has been well documented for other plant pathogens (e.g. fungi; Chen et al., 1993; Balardin et al., 1997), but, to our knowledge, this is the first report of such a result in the case of plant-parasitic nematodes.
The three RKN species studied here reproduce exclusively by apomictic (i.e. mitotic) parthenogenesis, where meiosis is suppressed and the egg undergoes a single mitotic maturation (Triantaphyllou, 1985). Although they theoretically clone themselves from generation to generation, parthenogenetic animals have been shown to display some degree of genetic variation, and it has been recently demonstrated in a number of cases that such diversity results from either mutation (Fu et al., 1998) or multiple (i.e. hybrid) origin (Sites et al., 1990). The lack of correlation observed here between (a)virulence of the populations and their AFLP fingerprints strongly suggests that the virulent phenotypes appeared late after the establishment of these clonal lineages. Therefore, it seems reasonable to hypothesize that they might be the result of independent mutational events, which probably occurred in parallel with the development of agricultural practices. Also, it is well known that, during its evolution, the genome of RKN has undergone extensive cytological modifications involving, among others, polyploidy, aneuploidy, and chromosome rearrangements (e.g. translocations, breakages without reassociation, etc.). Because of the holocentric nature of Meloidogyne chromosomes, such chromosomal modifications would not have been lost in subsequent generations, and would surely have contributed to the emergence of new clones (Triantaphyllou, 1981). From that point of view, and although it remains to be demonstrated, the hypothesis that virulent populations arose from avirulent ones through some unusual cytogenetical events is to be considered.
In the potato cyst nematode Globodera rostochiensis, close agreement was found between AFLP-based dendrograms and the current pathotype classification of populations (Folkertsma et al., 1996). This international pathotype scheme classifies potato cyst nematode populations on the basis of their multiplication rates on eight different potato lines carrying different resistance genes, and recognizes five pathotypes within G. rostochiensis (Kort et al., 1977). Conversely, the clustering of populations of the closely related species G. pallida revealed limited resemblance with their pathotype classification, maybe because of the structure of the genetic variation between them (Folkertsma et al., 1996). In fact, the level of G. pallida intraspecific variation was high compared to G. rostochiensis, and a considerable portion of the polymorphic DNA fragments proved to be scattered, which hampered an unambiguous classification of the populations analysed (Folkertsma et al., 1996). In cultivated tomato cultivars, resistance is conferred by the Mi gene only, although a number of other RKN resistance genes have recently been identified, mainly in accessions of the wild species Lycopersicon peruvianum (Yaghoobi et al., 1995; Veremis & Roberts, 1996a,b). The populations used here have been characterized for their (a)virulence against the Mi gene only, and our data for virulence phenotypes and DNA markers suggest that, within each species, virulent field-populations are not structured in congruence with their host genotype, i.e. presence or absence of Mi. Of interest would be the analysis of a wider collection of Meloidogyne biotypes showing differences in their (a)virulence to a range of additional tomato resistance genes, in order to test the generalization of our conclusions. Moreover, our work included only populations considered to be ‘naturally virulent’, i.e. that have been isolated from field populations thought not to have been selected by the Mi gene in recent agricultural time, although previous exposure(s) in evolutionary time may have occurred. Therefore, the absence of correlation between (a)virulence and molecular patterns could also be related, at least in part, to this apparent lack of coevolution between the plant and nematode genotypes.
Selection of M. incognita virulent lines from wild-type avirulent isolates under laboratory-controlled conditions by repeated exposure to resistant tomatoes carrying the Mi gene is possible (Jarquin-Barberena et al., 1991), although isofemale-line studies demonstrated that not all the populations of this species are potentially able to respond to such a selection process (Castagnone-Sereno et al., 1994). Undoubtedly, it seems reasonable to consider that selection for virulence may also occur under agronomic conditions after repeated planting of resistant tomato (Kaloshian et al., 1996; Eddaoui et al., 1997). Therefore, to test the hypothesis that selection for virulence by resistant genotypes may play a major role in determining the genetic relationship of RKN populations, two alternative sampling strategies need to be developed, either among laboratory-selected virulent lines or in areas where large-scale monoculture of resistant cultivars is routinely used. In both cases, the highly informative AFLP fingerprinting technique would then provide information on whether selection of (a)virulence alleles may be considered as a driving force for the genetic structure of RKN populations.
References
Balardin, R. S., Jarosz, A. M. and Kelly, J. D. (1997) Virulence and molecular diversity in Colletotrichum lindemuthianum from South, Central, and North America. Phytopathology. 87: 1184–1191.
Benzecri, J. P. (1973) L’analyse des Données vol. 2, L’analyse des Correspondences.Dunod, Paris.
Castagnone-Sereno, P., Bongiovanni, M. and Dalmasso, A. (1993) Stable virulence against the tomato resistance Mi gene in the parthenogenetic root-knot nematode Meloidogyne incognita. Phytopathology. 83: 803–805.
Castagnone-Sereno, P., Wajnberg, E., Bongiovanni, M., Leroy, F. and Dalmasso, A. (1994) Genetic variation in Meloidogyne incognita virulence against the tomato Mi resistance gene: evidence from isofemale line selection studies. Theor Appl Genet. 88: 749–753.
Chen, X., Line, R. F. and Leung, H. (1993) Relationships between virulence variation and DNA polymorphism in Puccinia striiformis. Phytopathology. 83: 1489–1497.
Colas, V., Lacourt, I., Ricci, P., Vanlerberghe-Masutti, F., Venard, P., Poupet, A.et al (1998) Diversity of virulence in Phytophthora parasitica on tobacco, as reflected by nuclear RFLPs. Phytopathology. 88: 205–212.
Dalmasso, A. and Bergé, J. B. (1978) Molecular polymorphism and phylogenetic relationship in some Meloidogyne spp. application to the taxonomy of Meloidogyne. J Nematol. 10: 323–332.
Eddaoui, M., Ammati, M. and Rammah, M. (1997) Identification of the resistance breaking populations of Meloidogyne on tomatoes in Morocco and their effect on new sources of resistance. Fundam Appl Nematol. 20: 285–289.
Erselius, L. J. and Devalavieille, C. (1984) Variation in protein profiles of Phytophthora: Comparison of six species. Trans Br Mycol Soc. 83: 463–472.
Folkertsma, R. T., Rouppevandervoort, J. N. A. M., Degroot, K. E., Vanzandvoort, P. M., Schots, A., Gommers, F. J.et al (1996) Gene pool similarities of potato cyst nematode populations assessed by AFLP analysis. Mol Plant–Microbe Interact. 9: 47–54.
Fu, J., Macculloch, R. D., Murphy, R. W., Darevsky, I. S., Kupriyanova, L. A. and Danielyan, F. (1998) The parthenogenetic rock lizard Lacerta Unisexualis: an example of limited genetic variation. J Mol Evol. 46: 127–130.
Gilbert, G. C. and Mcguire, D. C. (1956) Inheritance of resistance to severe root-knot from Meloidogyne incognita in commercial type tomatoes. Proc Am Soc Hortic Sci. 63: 437–442.
Greenacre, M. J. and Hastie, T. (1987) The geometric interpretation of correspondence analysis. J Am Stat Ass. 82: 437–447.
Jarquin-Barberena, H., Dalmasso, A., Deguiran, G. and Cardin, M. C. (1991) Acquired virulence in the plant parasitic nematode Meloidogyne incognita I. Biological analysis of the phenomenon. Rev Nématol. 3: 123–134.
Kaloshian, I., Williamson, V. M., Miyao, G., Lawn, D. A. and Westerdahl, B. B. (1996) Resistance-breaking nematodes identified in California tomatoes. Calif Agric. 50: 18–19.
Kort, J., Ross, H., Rumpenhorst, H. J. and Stone, A. R. (1977) An international scheme for identifying and classifying pathotypes of potato cyst nematodes Globodera rostochiensis and G. pallida. Nematologica. 23: 333–339.
Lamberti, F. (1979) Economic importance of Meloidogyne spp. in subtropical and Mediterranean climates. In: Lamberti, F. and Taylor, C. E. (eds) Root-Knot Nematodes (Meloidogyne species): Systematics Biology and Control pp. 341–357. Academic Press, London.
Laterrot, H. (1975) Séries de lignées isogéniques de tomate ne différent que par certains gènes de résistance aux maladies. Phytopath Medit. 14: 129–130.
Roberts, P. A. (1992) Current status of the availability, development, and use of host plant resistance to nematodes. J Nematol. 24: 213–227.
Sambrook, J., Fritsch, E. F. and Maniatis, T. (1989) Molecular Cloning: a Laboratory Manual2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
Sas Institute, Inc. (1990) SAS/STAT User’s Guide Release 6.07, vol. 1. SAS Institute Inc., Cary, NC.
Sasser, J. N. (1980) Root-knot nematodes: a global menace to crop production. Plant Dis. 64: 36–41.
Semblat, J. P., Wajnberg, E., Dalmasso, A., Abad, P. and Castagnone-Sereno, P. (1998) High-resolution DNA fingerprinting of parthenogenetic root-knot nematodes using AFLP analysis. Mol Ecol. 7: 119–125.
Sites, J. W. J. R., Peccinini-Seale, D. M., Moritz, C., Wright, J. W. and Brown, W. M. (1990) The evolutionary history of parthenogenetic Cnemidophorus lemniscatus (Sauria, Teiidae). I. Evidence for a hybrid origin. Evolution. 44: 906–921.
Triantaphyllou, A. C. (1981) Oogenesis and the chromosomes of the parthenogenetic root-knot nematode Meloidogyne incognita. J Nematol. 13: 95–104.
Triantaphyllou, A. C. (1985) Cytogenetics, cytotaxonomy and phylogeny of root-knot nematodes. In: Sasser, J. N. and Carter, C. C. (eds) An Advanced Treatise on Meloidogyne, Vol. I, Biology and Control pp. 113–126. North Carolina State University Graphics, Raleigh, NC.
Tzortzakakis, E. A. and Gowen, S. R. (1996) Occurrence of a resistance breaking pathotype of Meloidogyne javanica on tomatoes in Crete, Greece. Fundam Appl Nematol. 19: 283–288.
Veremis, J. C. and Roberts, P. A. (1996a) Identification of resistance to Meloidogyne javanica in the Lycopersicon peruvianum complex. Theor Appl Genet. 93: 894–901.
Veremis, J. C. and Roberts, P. A. (1996b) Relationships between Meloidogyne incognita resistance genes in Lycopersicon peruvianum differentiated by heat sensitivity and nematode virulence. Theor Appl Genet. 93: 950–959.
Yaghoobi, J., Kaloshian, I., Wen, Y. and Williamson, V. M. (1995) Mapping a new nematode resistance locus in Lycopersicon peruvianum. Theor Appl Genet. 91: 457–464.
Acknowledgements
We thank Christian Slagmulder for the photographic artwork. Support from the European Community (grant FAIR1-PL95-0896), from the Comité Technique Permanent de la Sélection (CTPS, France), and from the Bureau des Ressources Génétiques (BRG, France) is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Semblat, JP., Bongiovanni, M., Wajnberg, E. et al. Virulence and molecular diversity of parthenogenetic root-knot nematodes, Meloidogyne spp.. Heredity 84, 81–89 (2000). https://doi.org/10.1046/j.1365-2540.2000.00633.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00633.x