Abstract
Genetic diversity and differentiation were studied in Corrigiola litoralis L., an annual plant species growing on seasonally flooded river banks. Plant species that are restricted to river systems may consist of highly isolated populations. For this species, pronounced genetic differentiation among rivers was expected. Plants were sampled from the river Loire (France) representing subcentral populations and the rivers Rhine, Weser and Elbe (Germany) representing peripheral ones. Allozyme electrophoresis revealed 17 putative loci in 11 enzyme systems. At the species level, percentage polymorphic loci, mean number of alleles, observed heterozygosity and expected heterozygosity were P=29%, A=1.5 ± 0.2, Ho=0.007 ± 0.005 and He=0.065 ± 0.035, respectively. Peripheral populations were smaller in number and showed decreased levels of genetic diversity relative to central populations. Corrigiola litoralis was highly inbreeding as indicated by a mean FIS of 0.755. Genetic differentiation among populations was high with a mean FST-value of 0.585. Hierarchical F-statistics revealed that genetic variability was partitioned at 57% among sites, 52% among countries and 11% among sites within countries. Genetic distances between French and German populations were 0.08, indicative of considerable differentiation at the intraspecific level. The overall low level of allozyme diversity is attributed to the breeding system and to habitat conditions homogenized by regular flooding. The decrease in diversity from subcentral to peripheral populations is considered to be a result of drift and founder effects during postglacial recolonization. Peripheral populations were characterized by a single fixed allele at locus IDH, thus representing an evolutionarily significant unit.
Similar content being viewed by others
Introduction
The spatial structure of genetic variability has long been studied, both on local and large geographical scales (Brussard, 1984; Lesica & Allendorf, 1995; Linhart & Grant, 1996). The spatial structure of allozyme diversity in plant species is influenced by a variety of species-specific traits like life form, seed dispersal or geographical range, but most importantly by the breeding system (Hamrick & Godt, 1990, 1996). In contrast to outbreeding species, selfers or species with mixed mating systems show a substantial proportion of allozyme diversity partitioned among populations. The general patterns of allozyme diversity as influenced by species traits, however, may be affected by site-specific constraints. Whether a given population is located centrally or peripherally in the species range may exert an influence on its genetic diversity. At the periphery of the species range populations are often smaller and more isolated and selection could be more intense than in the centre of the species range, possibly leading to genetic differentiation. Thus, peripheral populations of selfing species with short generation time have been identified as potentially valuable for conservation of genetic diversity (Lesica & Allendorf, 1995).
Because species ranges are dynamic, historical range changes may also have influenced current genetic structure. In central Europe, postglacial recolonization may have played an important role in shaping the genetic structure of plant populations (Hewitt, 1996). A decrease in genetic diversity of central European populations relative to glacial refugia has been found in several European tree species (e.g. Lagercrantz & Ryman, 1990; Konnert & Bergmann, 1995; Demesure et al., 1996; Dumolin-Lapègue et al., 1997). In general, wind-pollinated outbreeding tree species show high levels of allozyme diversity and low levels of spatial differentiation (Hamrick & Godt, 1990). Thus, for species with lower levels of gene flow like selfers, comparatively large effects of historical range expansions on genetic diversity and differentiation can be expected.
The level of isolation of local populations will influence the genetic structure of a species, predominantly in species restricted to small and spatially isolated habitats. Rivers, flood plains and river margins are habitats with strong spatial structure. River systems exert two major influences: strong isolation between watersheds and potentially high connectivity along the river caused by running water, both of which will affect gene flow and genetic differentiation of organisms restricted to these habitats. The varying degree of isolation among and within watersheds has been shown for several animal species to lead to strong genetic differentiation and functional isolation-by-distance, assuming gene flow along the drainage system (e.g. Arter, 1990; Hänfling & Brandl, 1998). In aquatic plants, however, very low levels of genetic diversity and differentiation are often found and the homogeneity of aquatic habitats is stated as a possible explanation (Barrett et al., 1993). There are only a few studies that have addressed genetic differentiation of plants growing on river margins. It has been shown that gene flow occurring mainly within rivers leads to considerable differentiation among rivers (Ritland, 1989; Akimoto et al., 1998; Liao & Hsiao, 1998). In this paper genetic differentiation of Corrigiola litoralis L., a plant species mainly restricted to river margins (Coker, 1962; Vogel, 1997) is addressed.
Corrigiola litoralis (Illecebraceae, Caryophyllales) is an annual herb primarily growing on sandy or gravelly alluvial river bank soils at the flood-limit of rivers that are regularly flooded in winter. It produces several decumbent radiating shoots, 10–60 cm in diameter and reproduces entirely by seeds that lack specific dispersal devices but may be transported by water. Concerning the mode of pollination, statements range from ‘insect and self pollination’ (Frank & Klotz, 1990), ‘pollination may be entomophilous, but cleistogamy occurs under unfavourable conditions’ (Coker, 1962) to ‘self pollination’ (Oberdorfer, 1983; Vogel, 1997). Corrigiola litoralis has an atlantic–subatlantic–meridional–littoral distribution and occurs naturally in Portugal, France, Benelux, S. England, W. and N. Germany, W. Poland, locally along Mediterranean coasts, and in S. and E. Africa (Meusel et al., 1965; Field, 1994; Vogel, 1997). It finds optimal habitat conditions along the Loire (France), an unregulated river with extensive sandbanks (Wisskirchen, 1995). In Germany, C. litoralis has its largest populations along the river Elbe where it is regularly found. It reaches its continental limit of distribution near the border of the Czech Republic (Fig. 1). Along the rivers Weser and Rhine, only small and ephemeral populations are present. Along German rivers plants remain rather small compared to the Loire populations (Wisskirchen, 1995). Because of water level regulation C. litoralis is declining or already extinct at many places, e.g. in western Germany it is now absent from half of all previously occupied grid squares (Haeupler & Schönfelder, 1988). Based on the species range, population sizes, and individual plant size, we refer to German populations as peripheral, whereas French populations are subcentral.
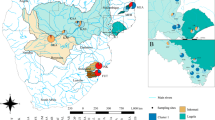
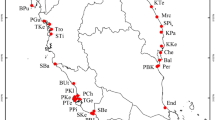
Sampling sites and species range (shaded area, after Field, 1994) of Corrigiola litoralis. For abbreviation of sites, see Table 1.
This study was undertaken to characterize levels of genetic diversity and genetic differentiation of C. litoralis within and among rivers. The rivers investigated represent subcentral and peripheral positions in the range of C. litoralis. It is hypothesized that: (i) small populations are depauperate in genetic diversity; (ii) peripheral populations show higher levels of homozygosity and inbreeding because of unfavourable conditions; (iii) genetic differentiation occurs mainly between and not within rivers, because gene flow predominantly occurs by seed dispersal and is mainly restricted to a single catchment area.
Methods
Plants were sampled from natural populations on sandy or stony river banks. In October 1997 between nine and 19 plants were sampled randomly from 15 populations along the rivers Elbe, Weser, Rhine and Loire (including Allier) (Fig. 1, Table 1). In large populations whole plants were sampled, whereas in small populations (e.g. at the Bonn site the total population size was n=9) only single branches were sampled from plants remaining on the site. Samples were immediately frozen in liquid nitrogen, transferred to the laboratory and stored at −42°C until analysis. Population size was obtained by direct counting or by estimation from habitat size and density.
Genetic variation was assessed using standard horizontal starch gel electrophoresis of isozymes from homogenized branches (including stems, leaves and flowers; Tris-HCl grinding buffer, Soltis et al., 1983). Staining recipes followed Wendel & Weeden (1989). A total of 11 enzyme systems gave clearly resolvable banding patterns: ACOH, ADH, AAT, GPI, IDH, CAP, MDH, PGDH, PGM, SkDH, MNR (Table 2).
The number of multilocus genotypes (Ngt), percentage of polymorphic loci (P), observed mean number of alleles per locus (Ao), mean observed heterozygosity (Ho), and mean expected heterozygosity under Hardy–Weinberg equilibrium (He) were calculated using Nei’s (1978) unbiased heterozygosity. The number of multilocus genotypes, Ngt, is sensitive to sample size per population, but as correlation between them was nonsignificant, Ngt was used in further analyses. Means of the diversity statistics were obtained by averaging population estimates within countries or over all populations. Species-level estimates were obtained by pooling all individuals across populations for calculation. Because sample sizes per population were very small for performing chi-squared tests, significance of deviations from Hardy–Weinberg equilibrium expected under panmictic conditions was tested with the conservative ‘Probability-test’ of GENE-POP (Raymond & Rousset, 1995). Tests were pooled across sites and across loci using Fisher’s method. Significance of FST values was assessed with a probability test using a Markov chain method (Raymond & Rousset, 1995).
Patterns of genetic variation within and among populations were analysed using hierarchical F-statistics (Wright, 1965, 1978) with sites, countries (Elbe + Weser + Rhine populations=Germany; Loire populations=France) and the total as hierarchical levels. Additionally, differentiation of populations was assessed by UPGMA-cluster analysis of genetic distances (Nei, 1972) between sites. Population genetic analyses were performed with GENEPOP (Raymond & Rousset, 1995) and BIOSYS (Swofford & Selander, 1989). Cluster analysis was performed with NTSYS 1.80 (Rohlf, 1983). Outcrossing rate was calculated as t=(1 − F)/(1 + F), where F is Wright’s fixation index (Nei & Syakundo, 1958), assuming that populations are in inbreeding equilibrium, i.e. F is constant over generations. Only populations and loci that showed polymorphism were used in the calculation of outcrossing rate. The effects of population size and country on genetic diversity statistics were tested with ANCOVA using countries (Germany, France) as groups and population size as a covariate. In this analysis, population size was log-transformed and percentage polymorphic loci was arcsin-transformed.
Results
Genetic diversity
A total of 187 individuals was analysed for electrophoretic variability of 11 enzymes encoded by 17 putative genetic loci (Tables 2 and 3 At the species level C. litoralis showed a low level of genetic diversity, with only five out of 17 (29%) loci being polymorphic: GPI-2, PGM, IDH-1, SkDH and ACOH-1 (Table 3). Applying a 0.99 or 0.95 criterion, only three loci (18%) were polymorphic. Species-wide estimates of mean number of alleles per locus and genetic diversity were A=1.5 ± 0.2 (SE), and He=0.065 ± 0.035 (SE), respectively.
French and German populations showed marked differences in their levels of genetic diversity. Along the Loire, all populations were polymorphic and all alleles but one of the polymorphic loci were detected. In contrast, in Germany only four downstream populations of the Weser and the Elbe were polymorphic, each at one single locus (GPI-2). In France a total of 16 multilocus genotypes was found in 77 individuals, whereas only four multilocus genotypes were detected in 111 individuals in Germany. Average gene diversity was He=0.059 ± 0.006 for French and He=0.007 ± 0.007 for German populations.
Corrigiola litoralis was highly homozygous: 109 out of 111 plants from nine German populations were homozygous at all 17 loci analysed. Only two plants from different populations were heterozygous at locus GPI-2 in German sites (Ho=0.001). In the Loire populations, a higher level of heterozygosity was detected (Ho=0.015, Table 1). Corrigiola is far from being at Hardy–Weinberg equilibrium, because out of 18 single-locus tests for deviation from equilibrium performed from all populations, 12 were significant (data not shown). Only one test is expected to be significant by chance at the 5% significance level. All combined tests, either combining across populations or across loci, were significant. Mean FIS of polymorphic loci was 0.755, indicating a low level (14%) of outcrossing (Table 4). Calculation of outcrossing rate for countries revealed 16% in French and 3% in German populations.
Measures of genetic diversity, Ngt, A, %P, Ho and He, were significantly correlated with each other (r ≥ 0.658, P ≤ 0.008). Ngt, A, %P and He were significantly correlated with population size (r ≥ 0.477, P ≤ 0.036). However, ANCOVA revealed that the country level had significant effects on all genetic diversity statistics, whereas population size had no significant effect after controlling for country (Table 5). This indicated that decreased genetic diversity in small populations was mainly a result of overall reduced genetic diversity in German populations.
Genetic differentiation
Population subdivision is high as shown by four of five FST values being significantly different from zero and reaching 0.98 at locus IDH (Table 4). Mean FST was 0.585. As no differentiation was found between the German rivers Rhine, Weser and Elbe, they were grouped together as Germany vs. France (Loire). The country level was used as an additional level in a hierarchical analysis. Hierarchical F-statistics revealed that genetic variability was partitioned at 57% among sites, 52% among countries and 11% among sites within countries (Fsite/total=0.568, Fsite/country=0.105, Fcountry/total=0.517). Differences between countries were attributable to two alleles of PGM and one allele of GPI-2, which only occurred in Loire populations, and the a-allele and b-allele of IDH-1, which occurred nearly exclusively in French and German sites, respectively (Table 3).
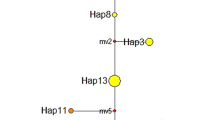
Genetic distances according to Nei (1972) among populations ranged from 0.000 to 0.102. Mean genetic distance was 0.010 and 0.001 within France and Germany, respectively, and 0.080 between France and Germany. The UPGMA cluster analysis of genetic distances (Fig. 2) emphasizes the geographical pattern of differentiation. Populations from the Loire and those from German rivers formed two distinct clusters. However, populations were not clustered according to geographical position along the respective rivers.
Discussion
Genetic diversity at the species level
Corrigiola litoralis is highly inbreeding, which was confirmed by seed-set in plants in a growth chamber (Durka, pers. obs.). Even in subcentral populations allowing optimal growth, only few heterozygotes were found. Thus the notion of Coker (1962) that cleistogamy occurs under unfavourable conditions must be extended in that autogamy is the rule in C. litoralis even under favourable conditions. In three German populations showing some allozyme polymorphism, outcrossing rate was lower than in French populations, supporting the hypothesis that in peripheral populations inbreeding is increased.
Although selfing leads to individual homozygosity, selfing species are not necessarily poor in allelic variants (Allard et al., 1968). Corrigiola litoralis, however, had low levels of overall diversity compared to other selfing species. The percentage of polymorphic loci (population level: 7.6%; species level: 29%) is far lower than the mean of selfing species (population level: 20%; species level: 42%; Hamrick & Godt, 1990). Limited allozyme variation may reflect restricted habitat range or homogeneity of the environment (Hedrick et al., 1976). Very low levels of genetic variability have been recorded in aquatic taxa (Barrett et al., 1993). Triest (1991) suggests that this may partly reflect the generally more homogeneous substrates in water and wetlands compared to strictly terrestrial habitats. All populations investigated in this study originated from similar primary habitats on sandy or gravelly banks of rivers flooded regularly in winter. In time and space this is a very narrow niche in which genetic homogeneity is plausible. On the other hand, within-habitat heterogeneity with respect to soil water is very high on sandbanks and riverbanks varying from 0 to 2 m above water level. Additionally, soil substrate along the river varies from gravel in the upper reaches to sand in the lower reaches. Thus, there is considerable heterogeneity in the habitat of C. litoralis, which could give rise to genetic variability. However, habitat dynamics may override any small-scale habitat structure. Corrigiola litoralis is faced with regular dramatic perturbations of the habitat through flooding which disrupts any small-scale structure of, for example, topography. Additionally, although seeds of C. litoralis lack special dispersal devices and would, without flooding, stay beneath the dying plant, flooding leads to mixing of seed genotypes, thus preventing the build-up of small-scale genetic structure.
Although there is considerable heterogeneity of habitat conditions, both within and between populations, regular flooding may promote genetic homogeneity because all individuals of a population may face the same environment. This may be a general feature of the plant community in which C. litoralis grows. Several of these annual species (e.g. Chenopodium spp., Polygonum lapathifolium, Xanthium spp.) are reported to display very low levels of genetic variability, at least as colonizers (Barrett & Shore, 1989). No perennial species is found together with Corrigiola. Interestingly, the plants from river margins for which considerable genetic variation has been reported, were outbreeding, insect-pollinated perennials (Ritland, 1989; Liao & Hsiao, 1998). Thus, the cgeneral observation that allozyme variability is lower in annual selfers than in perennial outcrossers (Hamrick & Godt, 1990) is confirmed for plants of river margins as well.
Intraspecific patterns of genetic differentiation
Genetic differentiation was found mainly among countries at a level of Nei distances of 0.08. This value corresponds to distances found between local races or subspecies, which typically range from 0.00–0.05 and 0.02–0.07, respectively (e.g. Levin, 1978; Helenurm & Ganders, 1985; Nei, 1987). Considering the geographical distribution of C. litoralis (Fig. 1), two subranges can be distinguished, namely (i) a north-eastern part including the Rhine, Weser and Elbe systems characterized by the b-allele of IDH-1 and (ii) a south-western part including the Loire characterized by the a-allele of IDH-1 and multiple alleles at GPI-2 and PGM. Large-scale geographical patterns of intraspecific races fixed for alternative alleles have previously been documented in other selfing annual species, e.g. Chenopodium fremontii (Crawford & Wilson, 1977). The b-allele of IDH-1, which occurs throughout the German populations, was detected in a single individual along the Loire. Although sample sizes in the Loire populations were restricted to 10–19 individuals and may therefore underestimate rare alleles, in a total of 77 individuals the b-allele only occurred once. Thus, the b-allele of IDH-1 is extremely rare in France and it might be argued that it is not identical to that detected in the German populations and represents a homoplasy.
Genetic differentiation in C. litoralis between French and German populations was caused by loss of alleles at two loci and fixation of different alleles at another locus. Several mechanisms may have led to the observed differentiation: (i) selection, (ii) genetic drift, or (iii) founder-effects in postglacial colonization history.
(i) Peripheral populations are under strong directional selection. Selection may act on enzyme loci or on genes linked to them (Clegg, 1983). In the context of the overall level of genetic diversity, habitat disturbance was discussed above as a potential selective force leading to genetic uniformity. However, different levels of diversity in subcentral and peripheral populations are not interpreted as an effect of differential intensities of habitat disturbance. The habitat of C. litoralis has to be maintained by physical perturbation equally in subcentral and peripheral populations. Selective forces limiting the atlantic–subatlantic range of C. litoralis may be connected to features of continentality like decreasing minimum temperatures, lengthening of frost periods or frequency of late frosts. In fact, the range of C. litoralis seems to be limited by low temperature, because in Germany no populations are found below the −2°C mean January isotherm (Vogel, 1997). Thus, low temperature might select specific genotypes in peripheral populations. However, it is not clear how low temperature can affect C. litoralis, because it survives frost periods as seed and its growing season is limited by the time of subsiding and rising floods.
(ii) Peripheral populations are at higher risk of genetic drift than more central populations. The rate of genetic drift is specified by the population size and the selfing rate (Gregorius, 1991). Peripheral populations of C. litoralis were smaller and had a higher inbreeding rate than subcentral populations. Thus, genetic drift may have led to decreased genetic variability in peripheral German populations of C. litoralis.
(iii) Peripheral populations frequently go extinct and recolonization may take place by only a few individuals comprising a low number of genotypes. Declining genetic diversity towards the periphery of the species ranges has been found in many species (Lesica & Allendorf, 1995). For European species a decline in genetic diversity was found, for example, in Alnus glutinosa (King & Ferris, 1998), Arabidopsis thaliana (Kuittinen et al., 1997), Calluna vulgaris (Mahy et al., 1997), Picea abies (Lagercrantz & Ryman, 1990) and Quercus (Dumolin-Lapègue et al., 1997). The decline of diversity in these species parallels south–north or east–west postglacial colonization routes and can be interpreted as the effect of repeated bottlenecks during range expansions (Hewitt, 1996). A similar scenario could be invoked for C. litoralis to explain the loss of alleles in German populations. Postglacial recolonization of the Rhine, Weser and Elbe river systems may have started from only a few genotypes originating from French rivers. This is supported by the fact that at the two loci GPI-2 and PGM, the allele observed in Germany is the most common allele in France. However, the change of one fixed allele into the other at locus IDH-1 requires the additional assumption of mutation at that particular locus since the time of colonization. Alternatively, a different source population for the current German populations might be postulated. To address this issue further, an extensive investigation of the IDH-1 locus and other markers at the DNA level needs to be undertaken in the whole range of C. litoralis.
In recent decades, C. litoralis has increasingly been found on secondary habitats like railway embankments, industrial dumps and water reservoirs with changing water level (Vogel, 1997). In the Netherlands, C. litoralis is currently only known from railway embankments (Westhoff, 1968). However, populations from such secondary habitats were not included in this study. It would be of interest to know to what extent these populations resemble the genotypes from primary habitats along river margins. As C. litoralis is a declining species, secondary habitats might play a role in conservation of the species (Vogel, 1997).
This study has shown that at the species level C. litoralis displays very low allozyme variability as an effect of inbreeding. This effect may be further enhanced by a narrow niche in a habitat regularly homogenized by flooding. Central European populations of C. litoralis are genetically depauperate relative to more south-western populations, probably as an after-effect of postglacial colonization history. Nevertheless, pronounced genetic differentiation caused by unique alleles makes peripheral populations of C. litoralis in central Europe important as evolutionarily significant units.
References
Akimoto, M., Shimamoto, Y. and Morishima, H. (1998). Population genetic structure of wild rice Oryza glumaepatula distributed in the Amazon flood area influenced by its life-history traits. Mol Ecol. 7: 1371–1381.
Allard, R. W., Jain, S. K. and Workman, P. L. (1968). The genetics of inbreeding populations. Adv Genet. 14: 55–131.
Arter, H. E. (1990). Spatial relationship and gene flow paths between populations of the alpine snail Arianta arbustorum (Pulmonata: Helicidae). Evolution. 44: 966–980.
Barrett, S. C. H. and Shore, J. S. (1989). Isozyme variation in colonizing plants. In: Soltis, D. E. and Soltis, P. S. (eds) Isozymes in Plant Biology, pp. 106–126. Dioscorides Press, Portland, OR.
Barrett, S. C. H., Eckert, C. G. and Husband, B. C. (1993). Evolutionary processes in aquatic plant populations. Aquat Bot. 44: 105–145.
Brussard, P. F. (1984). Geographic patterns and environmental gradients: The central–marginal model in Drosophila revisited. Ann Rev Ecol Syst. 15: 25–64.
Clegg, M. T. (1983). Detection and measurement of natural succession. In: Tanksley, S. D. and Orton, T. J. (eds) Isozymes in Plant Genetics and Breeding, Part A, pp. 241–255. Elsevier, Amsterdam.
Coker, P. D. (1962). Corrigiola litoralis L. Biological Flora of the British Isles. J Ecol. 50: 833–840.
Crawford, D. J. and Wilson, H. D. (1977). Allozyme variation in Chenopodium fremontii. Syst Bot. 2: 180–190.
Demesure, B., Comps, B. and Petit, R. J. (1996). Chloroplast DNA phylogeography of the common beech (Fagus sylvatica L.) in Europe. Evolution. 50: 2515–2520.
Dumolin-Lapègue, S., Demesure, B., Fineschi, S., Lecorre, V. and Petit, R. J. (1997). Phylogeographic structure of white oaks throughout the European continent. Genetics. 146: 1475–1487.
Field, M. H. (1994). Corrigiola litoralis in the British Middle Pleistocene. New Phytol. 126: 393–395.
Frank, D. and Klotz, S. (1990). Biologisch-ökologische Daten zur Flora der DDR. Wissenschaftliche Beiträge Martin-Luther Univ Halle-Witt. 32
Gregorius, H. -R. (1991). Gene conservation and the preservation of adaptability. In: Seitz, A. and Loeschcke, V. (eds) Species Conservation: A Population-Biological Approach, pp. 31–48. Birkhäuser, Basel.
Haeupler, H. and Schönfelder, P. (1988). Atlas der Farn- und Blütenpflanzen der Bundesrepublik Deutschland. Ulmer, Stuttgart.
Hamrick, J. L. and Godt, M. J. (1990). Allozyme diversity in plant species. In: Brown, A. H. D., Clegg, M. T., Kahler, A. L. and Weir, B. S. (eds) Plant Population Genetics, Breeding, and Genetic Resources, pp. 43–63. Sinauer Associates, Sunderland, MA.
Hamrick, J. L. and Godt, M. J. (1996). Effects of life history traits on genetic diversity in plant species. Phil Trans R Soc B. 351: 1291–1298.
Hänfling, B. and Brandl, R. (1998). Genetic differentiation of the bullhead Cottus gobio L. across watersheds in Central Europe: evidence for two taxa. Heredity. 80: 110–117.
Hedrick, P. W., Ginevan, M. E. and Ewing, E. P. (1976). Genetic polymorphism in heterogeneous environments. Ann Rev Ecol Syst. 7: 1–32.
Helenurm, K. and Ganders, F. R. (1985). Adaptive radiation and genetic differentiation in Hawaiian Bidens. Evolution. 39: 753–765.
Hewitt, G. M. (1996). Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc. 59: 247–276.
King, R. A. and Ferris, C. (1998). Chloroplast DNA phylogeography of Alnus glutinosa (L.) Gaertn. Mol Ecol. 7: 1151–1161.
Konnert, M. and Bergmann, F. (1995). The geographical distribution of genetic variation of silver fir (Abies alba, Pinaceae) in relation to its migration history. Pl Syst Evol. 196: 19–30.
Kuittinen, H., Mattila, A. and Savolainen, O. (1997). Genetic variation at marker loci and in quantitative traits in natural populations of Arabidopsis thaliana. Heredity. 79: 144–152.
Lagercrantz, U. and Ryman, N. (1990). Genetic structure of Norway spruce (Picea abies): concordance of morphological and allozymic variation. Evolution. 44: 38–53.
Lesica, P. and Allendorf, F. W. (1995). When are peripheral populations valuable for conservation?. Conserv Biol. 9: 753–760.
Levin, D. A. (1978). Genetic variation in annual Phlox: self-compatible versus self-incompatible species. Evolution. 32: 245–263.
Liao, L. C. and Hsiao, J. Y. (1998). Relationship between population genetic structure and riparian habitat as revealed by RAPD analysis of the rheophyte Acorus gramineus Soland. (Araceae) in Taiwan. Mol Ecol. 7: 1275–1281.
Linhart, Y. B. and Grant, M. C. (1996). Evolutionary significance of local genetic differentiation in plants. Ann Rev Ecol Syst. 27: 237–277.
Mahy, G., Vekemans, X., Jacquemart, A. L. and de Sloover, J. R. (1997). Allozyme diversity and genetic structure in South-Western populations of heather, Calluna vulgaris. New Phytol. 137: 325–334.
Meusel, H., Jäger, E. J. and Weinert, E. (1965). Vergleichende Chorologie der Zentraleuropäischen Flora. Fischer, Jena.
Murphy, R. W., Sites, J. W., Buth, D. G. and Haufler, C. H. (1996). Proteins: isozyme electrophoresis. In: Hillis, D. M., Moritz, C. and Mable, B. K. (eds) Molecular Systematics, pp. 51–120. Sinauer Associates, Sunderland, MA.
Nei, M. (1972). Genetic distance between populations. Am Nat. 106: 283–292.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics. 89: 583–590.
Nei, M. (1987). Molecular Evolutionary Genetics. Columbia University Press, New York.
Nei, M. and Syakundo, M. (1958). The estimation of outcrossing in natural populations. Jap J Genet. 33: 46–51.
Oberdorfer, E. (1983). Pflanzensoziologische Exkursionsflora. Ulmer, Stuttgart.
Raymond, M. and Rousset, F. (1995). Genepop (version 1.2): population genetics software for exact tests and ecumenicism. J Hered. 86: 248–249.
Ritland, K. (1989). Genetic differentiation, diversity, and inbreeding in the mountain monkeyflower (Mimulus caespitosus) of the Washington Cascades. Can J Bot. 67: 2017–2024.
Rohlf, F. J. (1983). Ntsys-pc, version 1.8. Exeter Software, New York.
Selander, R. K., Smith, M. H., Young, S. Y., Johnson, W. E. and Gentry, J. B. (1971). Biochemical polymorphism and systematics in the genus Peromyscus. University Texas Publ. 7103: 49–90.
Soltis, D. E., Haufler, C. H., Darrow, D. C. and Gastony, G. J. (1983). Starch gel electrophoresis of ferns: a compilation of grinding buffers, gel and electrode buffers, and staining schedules. Am Fern J. 73: 9–27.
Swofford, D. L. and Selander, R. B. (1989). Biosys-1. A computer program for the analysis of allelic variation in population genetics and biochemical systematics. Illinois Natural History Survey, Urbana, IL.
Triest, L. (1991). Isozymes in water plants: molecular systematics and biogeography. Opera Botanica Belgica, vol. 4. Nat. Bot. Garden, Meise.
Vogel, A. (1997). Die Verbreitung, Vergesellschaftung und Populationsökologie von Corrigiola litoralis, Illecebrum verticillatum und Herniaria glabra (Illecebraceae). Dissertationes Botanicae. vol. 289. Cramer, Berlin, Stuttgart.
Weir, B. S. and Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. Evolution. 38: 1358–1370.
Wendel, J. F. and Weeden, N. F. (1989). Visualization and interpretation of plant isozymes. In: Soltis, D. E. and Soltis, P. S. (eds) Isozymes in Plant Biology, pp. 5–45. Dioscorides Press, Portland, OR.
Westhoff, V. (1968). Standplaatsen van Corrigiola litoralis L. Gorteria. 4: 137–145.
Wisskirchen, R. (1995). Verbreitung und Ökologie von Flußufer-Pioniergesellschaften (Chenopodion rubri) im mittleren und westlichen Europa. Dissertationes Botanicae, vol. 236. Cramer, Berlin, Stuttgart.
Wright, S. (1965). The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution. 19: 395–420.
Wright, S. (1978). Evolution and the Genetics of Populations, vol. 4, Variability Within and Among Natural Populations. University of Chicago Press, Chicago.
Acknowledgements
I thank Bernd Hänfling for introducing me to allozyme analysis. Valuable comments on drafts of the paper and discussions provided by R. Brandl, M. Brändle, P. Comes, B. Hänfling, T. Schmitt, L. Soldaat and two anonymous referees are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Durka, W. Genetic diversity in peripheral and subcentral populations of Corrigiola litoralis L. (Illecebraceae). Heredity 83, 476–484 (1999). https://doi.org/10.1038/sj.hdy.6886000
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6886000
Keywords
This article is cited by
-
Genetic diversity and reproductive ecology of the sage-leaved rockrose, Cistus salviifolius L., in the Swiss Alps
Plant Ecology (2020)
-
Population genetic structure of the whorl-leaf watermilfoil Myriophyllum verticillatum shaped by topography and geographic distance
Hydrobiologia (2019)
-
Genetic variation of Gentianella campestris ssp. campestris in the Northern Alps: how important are population size and isolation?
Alpine Botany (2019)
-
Interannual dynamics of a rare vegetation on emerged river gravels with special attention to the critically endangered species Corrigiola litoralis L
Folia Geobotanica (2018)
-
Genetic variation and fine-scale population structure in American pikas across a human-modified landscape
Conservation Genetics (2017)