Abstract
Allozyme diversity was surveyed at 16 loci across 16 populations of the monospecific, but morphologically diverse, endemic shrub Geleznowia verrucosa Turcz. (Rutaceae). Single-locus diversity measures portrayed G. verrucosa as a genetically depauperate genus (A, 1.4; P, 29.6%), with total genetic diversity (HT, 0.304) partitioned between populations (DST, 0.175), rather than within (HS, 0.129). Some 58% of the total genetic diversity was attributable to interpopulational differences. Although restricted distribution, bottlenecks and/or founder effects and small population size have probably contributed to the low levels of genetic diversity found within this genus, it is evident from this investigation that the genus is not monospecific. The partitioning of genetic diversity, and possible differences in reproductive strategy, suggest that the small- and large-flowered forms represent distinct taxa. Higher levels of divergence between the intermediate-form populations and increased heterozygosity at the Lap locus suggest that this form is of hybrid origin and has arisen through a series of hybrid events between the small and large forms.
Similar content being viewed by others
Introduction
Geographically restricted species are often characterized by lower genetic diversity than more widely distributed species. This diversity is, however, often apportioned in much the same manner as that of more widespread species (Karron, 1987; Hamrick & Godt, 1989). Small population size, reduced gene flow, inbreeding, and founder effects all contribute to the lower genetic diversity of endemic taxa (Hamrick & Godt, 1989).
The Western Australian flora, particularly that of the south-west region, is distinguished by high species diversity and endemism (Hopper, 1979). Many evolutionary patterns occur across the landscape. Refugial species endure in the higher rainfall areas, whereas fragmented relictual species and suites of newly derived taxa exist in the more arid areas (Hopper, 1979). Many species are rare and locally endemic, the product of a long and complex evolution that has generated many distinct genetic systems (Hopper, 1979). Some of the more well-known mechanisms include intraspecific chromosome variation (James, 1965; Coates & James, 1979), polyploidy (Bousfield & James, 1976), dysploidy (James, 1981; Stace, 1995) and complex hybridity (James, 1970).
The levels and distribution of genetic diversity within Western Australian endemics do not always concur with expectations. For example, although some rare and localized species such as Banksia cuneata (Coates & Sokolowski, 1992) and Eucalyptus caesia (Moran & Hopper, 1983) exhibit low levels of genetic diversity, other narrow endemics such as Acacia anomala (Coates, 1988) and Stylidium coroniforme (Coates, 1992) do not.
Genetic diversity has also been found to be apportioned both within (Coates & Hnatiuk, 1990; Byrne & James, 1991) and between (Moran & Hopper, 1983; Sampson et al., 1988) populations. This genetic variability probably reflects the complex evolutionary history of the area, where recently evolved taxa coincide with more ancient relictual taxa.
Geleznowia verrucosa Turcz. (yellow bells) is a little-known woody shrub, endemic to the sand plains of Western Australia. It occurs as small disjunct populations from Cape Range (21°50′S, 116°12′E) in the north of the state, to Dowerin (31°12′S, 117°02′E) in the eastern wheat belt (Hnatiuk, 1990; Keighery & Gibson, 1993). Although considered monospecific, G. verrucosa is morphologically diverse, with populations ranging from a taller (1–2 m) form with abundant, large flowers and leaves, to populations comprising shorter plants (0.5–1 m) with small, often solitary, flowers and leaves. Between these forms are a number of variable populations exhibiting intermediate characters. Although the different forms rarely co-occur, when they do, morphological integrity is generally maintained.
This research was initiated to investigate patterns of genetic diversity within and between the disjunct and morphologically variable populations of G. verrucosa, and to determine whether the different forms are genetically distinct and represent different taxa.
Materials and methods
Sampling strategy
Sixteen populations from within the known distribution of G. verrucosa and representing the three morphological forms (small, intermediate and large) were selected for electrophoretic study (Fig. 1). Where possible, 20 plants at each population were mapped, tagged and the total number of plants counted/estimated. To compare genetic variation within the unusually large Hutt River population, two sites of 20 plants ≈500 m apart were selected. At Pleshkes, where the small and large forms co-occurred, 10 plants from each form were selected, as well as 20 plants along a transect across the integration zone between them. Inflorescences were collected from tagged plants during 1995, placed in sealed plastic bags and transported to the laboratory at 4°C, where buds of a similar age were frozen at −72°C. Allozyme electrophoresis was carried out on anther material from these buds.
Distribution of Geleznowia verrucosa populations. Symbols: ▴ small, • intermediate, ▪ large, ♦ mixed forms and transect. Population codes: A, Arinya; AW1, Ajana West 1; AW2, Ajana West 2; C1, Coorow 1; C2, Coorow 2; G, Geraldton; HH, Hawks Head; HR, Hutt River (sites 1 and 2); IR, Indarra Reserve; M, Maiseys; MH1, Meanarra Hill 1; MH2, Meanarra Hill 2; N, Nanns; P, Pleshkes (PL, large form; PH, transect; PS, small form); RB, Red Bluff; WB, West Binnu.
Electrophoresis
Anthers were ground with 30 μL of modified extraction buffer (Systma & Schaal, 1985) (50 mg mL–1 PVP, 0.5 mM NAD, 0.3 mM NADP, 1.7 mMEDTA, 1.2 mM ascorbic acid, 0.1% BSA, 10% sucrose w/v and 0.7 mg mL–1 dithiothreitol in 0.1 M Tris–HCl, pH 7), centrifuged for 1.5 min and 12 μL of extract transferred to sample wells from which 1 μL was loaded onto presoaked cellulose acetate plates (Titan III, Helena Laboratories, Beaumont, TX). The plates were transferred to running tanks maintained at 5°C and electrophoresed at 5 mA/gel. Running times varied from 30 to 50 min depending upon the enzyme system under analysis.
Eleven enzyme systems were assayed (Table 1). All systems were run in an 80 mM Tris–EDTA–maleic acid buffer (80 mM Tris, 1 mM Na2EDTA, 1 mM MgCl2, 6.4 mM histidine, 13.3 mM glycine, 3.4 mM glutamic acid, 3.7 mM aspartic acid, pH to 7.6 with maleic acid then pH to 8.1 with sodium hydroxide) except Aat, which was run in Tris–citrate buffer pH 8.1 (100 mM Tris, 10 mM citric acid). Staining procedures followed Richardson et al. (1986), except that 4 mL of stain buffer were used with stain ingredients proportionally increased. Sixteen zones of activity were scored for nine enzymes with each zone assumed to represent a single enzyme locus: Aat-1, Aat-2, Aat-3, Aat-4, Aat-5, Est-1, Gpi-1, Idh-1, Lap-1, Mdh-1, Mdh-2, Mdr-1, Mdr-2, Mdr-3, Pgm-1 and Skd-1. Alleles were designated by enzyme system, locus and allele. For example, Aat-1c is allele c at locus 1 for aspartate aminotransferase.
Genetic analyses
The mean number of alleles per locus (A), proportion of polymorphic loci (P, 0.99 criterion), observed (Ho) and expected heterozygosity (He) were calculated using BIOSYS-1 (Swofford & Selander, 1981). Wright’s (1978) FIS, which is a measure of heterozygote deficiency or excess, was calculated using POPGENE version 1.2 (Yeh et al., 1997). Total genetic diversity and its distribution within and between populations was estimated using Nei’s (1973) gene diversity statistics and calculated using GENESTAT (Whitkus, 1985). Total gene diversity (HT) was partitioned into mean gene diversity within (HS) and between (DST) populations. The proportion of interpopulation differentiation (GST) was determined by GST = DST/HT. Nei’s (1972) genetic distance was used to generate the cluster analysis with standard error bars based on the unweighted pair-group average method using arithmetic means (UPGMA) with Ritland’s (1989) Genetic Distance and Clustering (GD) program.
Results
Genetic variation within and among populations
Allelic frequencies varied among populations (Table 2). Although only three monomorphic loci were observed (Idh-1, Mdh-2 and Mdr-1), in many populations, loci were either homozygous or the alternative alleles were at very low frequencies (e.g. Aat-1c at Nanns, Gpi-1d at Ajana West 2, Gpi-1b at Coorow 1 and Mdr-3b at Hutt River 2).
Single-locus diversity measures were also low (Table 3). The mean number of alleles per locus (A) was 1.4 (range 1.1–1.8), and the mean percentage of polymorphic loci (P) was 29.6% (range 6.3% to 62.5%.). Observed heterozygosity (Ho, 0.055) was considerably lower than expected (He, 0.097). Populations exhibited marked differences in single-locus diversity measures. Heterozygous individuals were absent from the Ajana West 2, Hawks Head and Coorow 2 populations, and very low levels of heterozygosity characterized those at Ajana West 1 (0.006), Arinya (0.011), Coorow 1 (0.017), Maiseys (0.016) and Hutt River 2 (0.022), where individuals with similar genotypes dominated. In contrast, populations at Meanarra Hill, Red Bluff and West Binnu exhibited higher levels of diversity and approached panmixia. These results are also reflected in the FIS values which varied in a similar manner across populations (range −0.025 to 0.387).
Genetic variation within and among forms
Forms also exhibited marked differences in single-locus diversity measures (Table 3). The percentage of polymorphic loci was lower in the small (26.1%) and intermediate (23.2%) forms than in the large form (36.3%). The large form also approached panmixia, whereas observed heterozygosity was considerably lower than that expected in the other two forms. This was again reflected by the fixation indices for each form.
Some loci showed fixed allelic or highly significant allele frequency differences among the morphological forms. Alleles Aat-2a and Aat-4a were restricted to the large form, except for the small form at Pleshkes, where both forms co-occurred (Table 2). The allele Aat-3a was similarly restricted to the large form, except for small-form populations at Indarra Reserve and Pleshkes.
The distribution of alleles at Pleshkes where the small and large forms co-occurred strongly suggests a zone of hybridization. The small form at this site exhibited unique alleles Aat-1b and Aat-3c as well as other alleles restricted to the large form (Aat-2a, Aat-4a, Aat-5b and Aat-5c), indicating that introgression between the forms has been, and probably still is, occurring (Broadhurst, unpubl. obs.).
The mean total genetic diversity (HT) was 0.304, with diversity distributed among populations (DST, 0.175) rather than within them (HS, 0.129) (Table 4). The proportion of genetic diversity among populations (GST) was very high at 0.576, indicating that 58% of the total diversity was attributable to interpopulation differentiation.
Gene diversity statistics generated according to morphological forms showed that the mean total genetic diversity (HT) of the small form was lower (0.192) than that of the two other forms, with more of the diversity apportioned within populations (HS, 0.122) rather than among them (DST, 0.070) (Table 4). Interpopulational differentiation accounted for 36% of this variation. Although genetic diversity measures for the large form were somewhat higher (HT, 0.254), this form also partitioned genetic diversity within populations (HS, 0.164) rather than among them (DST, 0.090), with similar interpopulational differentiation (GST, 35%). Total gene diversity in the intermediate form (HT, 0.226) was similar to the large form but was apportioned among populations (DST, 0.121) rather than within them (HS, 0.105). These intermediate-form populations, with the highest interpopulational differentiation (GST, 54%), contributed significantly to the high overall interpopulation differentiation encountered within the complex, and to the differentiation between the small and large forms.
Genetic divergence among populations
Average genetic distance (D) within the small and large forms was low, 0.071 and 0.070, respectively, but higher within the intermediate form (0.112). Distances between the intermediate and large forms were considerably higher (0.301) than those between the small and large forms (0.223) and the small and intermediate forms (0.122).
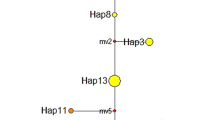
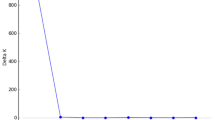
Relationships among populations based on estimates of genetic distance (D) are summarized in Fig. 2. Two distinct clusters were observed. The initial dichotomy separated the large-form populations at Pleshkes, Meanarra Hill, Red Bluff and Geraldton as a distinct group, within which the small form at Pleshkes was a discrete entity. The transect at Pleshkes (Pleshkes Hybrid) was also included in this group. The second group consisted of both the small and intermediate forms. All small-form populations, with the exception of Pleshkes Small, were clustered together with the intermediate form at Hutt River.
Cluster analysis of Geleznowia verrucosa populations based on Nei’s genetic distance (D) and UPGMA algorithm. Population clusters are statistically significant if the shaded bar is less than half the branch length (Ritland, 1989). Population and symbol codes in Fig. 1.
Discussion
Endemism and G. verrucosa
The levels of genetic diversity in G. verrucosa (P, 29.6%; A, 1.4) were consistent with those observed at the population level in other endemic taxa (P, 26.3%; A, 1.4; Hamrick & Godt, 1989). With few exceptions, G. verrucosa populations are small and disjunct, predisposing them to limited gene flow, genetic drift and fixation (Loveless & Hamrick, 1984; Hamrick & Godt, 1989). Such conditions are highly conducive to creating the population genetic structure and the low levels of genetic diversity observed. The extremely low levels of allozyme polymorphism exhibited by some populations, dominance of others by relatively few genotypes and significant population differentiation suggest that other factors also contribute to genetic diversity within G. verrucosa.
Bottlenecks, both natural and artificial, and/or associated founder effects may play a significant role in the patterns of genetic diversity observed in G. verrucosa. A recent investigation has highlighted this genus to be a disturbance opportunist, with smoke being an important germination factor (Dixon et al., 1995; Roche, 1998). In addition, seed bank trials indicated that little seed remains in the seed soil bank after one year and that ants quickly predate much of the annual production (Roche, 1998). Under these circumstances, seedling recruitment following disturbance is likely to be from limited seed resources primarily produced in the previous season, effectively generating natural bottlenecks.
Several of the G. verrucosa populations may have also experienced artificial bottlenecks. Populations at Hawks Head, Geraldton and Nanns occur in disturbed habitats, those at Ajana West, Coorow, Indarra Reserve and West Binnu populations are located on road verges, and at Pleshkes and Hutt River the populations are periodically disturbed when commercial cut-flower production falls. Many of these populations exhibited low levels of genetic diversity and departure from Hardy–Weinberg equilibrium (Table 3).
Levels of diversity within and among forms of G. verrucosa
The level of genetic divergence between the large form and the small and intermediate forms, the presence of alleles characterizing the various forms, and disparate apportioning of genetic diversity within the three forms, is highly suggestive of systematic separation. Differences in both the levels of genetic diversity and the fixation indices (FIS) further suggest that contrasting reproductive strategies are employed, but insufficient levels of polymorphism among progeny have not allowed useful estimates of outcrossing to be made.
The low levels of genetic diversity and homozygote excesses observed in the small form are consistent with expectations of self-compatible species (Hamrick & Godt, 1989). Other characters such as smaller less attractive flowers, abundant seed and distribution peripheral to that of the large form (Grant, 1975; Solbrig & Rollins, 1977; Cruden & Lyon, 1985; Ritland & Ritland, 1989), are further evidence of a selfing reproductive strategy. In contrast, the large form exhibits levels of genetic diversity concordant with predominantly outcrossed species (Hamrick & Godt, 1989), has larger and more abundant flowers and sets less seed, characteristic of an outcrossing, or at least mixed-mating reproductive strategy (Solbrig & Rollins, 1977; Lyons & Antonovics, 1991). Intermediacy in floral characters and gene diversity indices for the intermediate form suggest that this may be of hybrid origin between the small and large forms.
The apportioning of genetic diversity in the intermediate form strongly supports the notion of hybrid ancestry. The moderate total genetic diversity of the small and large forms resided within populations with relatively low levels of interpopulational differentiation. Such genetic diversity could readily be expected to reside within populations when taxa are compared. In contrast, the higher total genetic diversity of the intermediate form was apportioned between populations with 54% of the total genetic diversity attributable to interpopulational differences. This greater genetic diversity and population differentiation could reflect hybridization events generating a series of diverse hybrid products.
The relationships between populations based on genetic distances further suggest that the intermediate form is of hybrid origin and that several independent hybridization events have occurred (Fig. 2). For example, although the Ajana West populations are only 3 km apart, Ajana West 1 was closely allied with Hawks Head, 5 km west, whereas Ajana West 2 was closely related to Nanns, located 50 km south. The former populations are linked by the allele Aat-4b, exhibited by most of the large-form populations, whereas the latter populations share the Aat-4c allele, common in the small and intermediate forms. The presence of these contrasting alleles despite close proximity in the Ajana West populations may reflect separate hybridization events.
The major argument against the intermediate form being of hybrid origin is the disjunct nature of extant populations. The present distribution patterns indicate that little opportunity for hybridization exists, but such population disjunction may not always have been the case. Geleznowia verrucosa occurs within the transitional-rainfall zone, a species-rich region with a high number of infraspecific variants, cryptic species complexes and natural hybrids (Hopper, 1979, 1992). Plant evolution in this region is considered to have been largely driven by small population size, induced by recurrent late Tertiary–Quaternary events, in a heterogeneous soil mosaic (Hopper, 1992). Range expansion and contraction during these perturbations may have brought about a series of temporally and spatially distributed hybridization events, generating many novel genetic arrays and apportioning genetic diversity between rather than within the resultant hybrid populations. Unique selection within each location would further contribute to population differentiation.
This investigation has revealed the contribution of restricted distributions, population size, disjunction and bottlenecks to genetic diversity within G. verrucosa. More significantly, it has highlighted an enigmatic species complex comprised of two taxa, with strong evidence to suggest the evolution of a third through hybridization. Morphometric and reproductive analyses provide further evidence of systematic separation (Broadhurst, unpubl. obs.). Given the past climatic and geological history of the south-west of Western Australia, hybridization may be a much more significant feature of this flora than previously thought.
References
Bousfield, L. R. and James, S. H. (1976). The behaviour and possible cytoevolutionary significance of B chromosomes in Dampiera linearis (Angiospermae: Goodeniaceae). Chromosoma, 55: 309–323.
Byrne, M. and James, S. H. (1991). Genetic diversity in the cycad Macrozamia riedlei. Heredity, 67: 35–39.
Coates, D. J. (1988). Genetic diversity and population genetic structure in the rare Chittering Grass Wattle, Acacia anomala Court. Aust J Bot, 36: 273–286.
Coates, D. J. (1992). Genetic consequences of a bottleneck and spatial genetic structure in the triggerplant Stylidium coroniforme (Stylidiaceae). Heredity, 69: 512–520.
Coates, D. J. and Hnatiuk, R. J. (1990). Systematic and evolutionary inferences from isozyme studies in the genus Eremaea (Myrtaceae). Aust Syst Bot, 3: 59–74.
Coates, D. J. and James, S. H. (1979). Chromosome variation in Stylidium crossocephalum and the dynamic coadaption of its lethal system. Chromosoma, 72: 357–376.
Coates, D. J. and Sokolowski, R. E. S. (1992). The mating system and patterns of genetic variation in Banksia cuneata A.S. George (Proteaceae). Heredity, 69: 11–20.
Cruden, R. W. and Lyon, D. L. (1985). Patterns of biomass allocation to male and female functions in plants with different mating systems. Oecologia, 66: 299–306.
Dixon, K. W., Roche, S. and Pate, J. S. (1995). The promotive effect of smoke derived from burnt native vegetation on seed germination of Western Australian plants. Oecologia, 101: 185–192.
Grant, V. (1975). Genetics of Flowering Plants. Columbia University Press, New York.
Hamrick, J. L. and Godt, M. J. W. (1989). Allozyme diversity in plant species. In: Brown, A. H. D., Clegg, M. T., Kahler, A. L. and Weir, B. S. (eds) Plant Population Genetics, Breeding, and Genetic Resources, pp. 43–63. Sinauer Associates, Sunderland, MA.
Hnatiuk, R. J. (1990). Census of Australian Vascular Plants. Australian Government Public Service, Canberra.
Hopper, S. D. (1979). Biogeographic aspects of speciation in the southwestern Australian flora. Ann Rev Ecol Syst, 10: 399–422.
Hopper, S. D. (1992). Patterns of plant diversity at the population and species levels in south-west Australian mediterranean ecosystems. In: Hobbs, R. J. (ed.) Biodiversity of Mediterranean Ecosystems in Australia, pp. 27–46. Surrey Beatty & Sons, Chipping North, New South Wales.
James, S. H. (1965). Complex hybridity in Isotoma petraea I. The occurrence of interchange heterozygosity, autogamy and a balanced lethal system. Heredity, 20: 341–353.
James, S. H. (1970). Complex hybridity in Isotoma petraea. Heredity, 25: 53–77.
James, S. H. (1981). Cytoevolutionary patterns, genetic systems and the phytogeography of Australia. In: Keast, A. (ed.) Ecological Biogeography of Australia, pp. 763–782. Dr W. Junk, Publishers, The Hague.
Karron, J. D. (1987). A comparison of levels of genetic polymorphism and self-compatibility in geographically restricted and widespread plant congeners. Evol Ecol, 1: 47–58.
Keighery, G. and Gibson, N. (1993). Biogeography and composition of the flora of the Cape Range Peninsula, Western Australia. In: Humphreys, W. F. (ed.) The Biogeography of Cape Range Western Australia. Records of the Western Australian Museum Supplement no., 45, pp. 51–85. Western Australian Museum, Perth, Western Australia.
Loveless, M. D. and Hamrick, J. L. (1984). Ecological determinants of genetic structure in plant populations. Ann Rev Ecol Syst, 15: 65–95.
Lyons, E. E. and Antonovics, J. (1991). Breeding system evolution in Leavenworthia: breeding system variation and reproductive success in natural populations of Leavenworthia crassa (Cruciferae). Am J Bot, 78: 270–287.
Moran, G. F. and Hopper, S. D. (1983). Genetic diversity and the insular population structure of the rare granite rock species, Eucalyptus caesia Benth. Aust J Bot, 31: 161–172.
Nei, M. (1972). Genetic distance between populations. Am Nat, 106: 283–292.
Nei, M. (1973). Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA, 70: 3321–3323.
Richardson, B. J., Baverstock, P. R. and Adams, M. (1986). Allozyme Electrophoresis: A Handbook for Animal Systematics and Population Studies, Academic Press, Sydney.
Ritland, K. (1989). Genetic differentiation, diversity and inbreeding in the mountain monkeyflower (Mimulus caespitosus) of the Washington Cascades. Can J Bot, 67: 2017–2024.
Ritland, C. and Ritland, K. (1989). Variation of sex allocation among eight taxa of the Mimulus guttatus species complex (Scrophulariaceae). Am J Bot, 76: 1731–1739.
Roche, S. (1998). Smoke Induced Seed Germination of Selected Australian Native Plants. Ph.D. Thesis, Botany Department, University of Western Australia.
Sampson, J. F., Hopper, S. D. and James, S. H. (1988). Genetic diversity and the conservation of Eucalyptus crucis Maiden. Aust J Bot, 36: 447–460.
Solbrig, O. T. and Rollins, R. C. (1977). The evolution of autogamy in species of the mustard genus Leavenworthia. Evolution, 31: 265–281.
Stace, H. M. (1995). Primitive and advanced character states for chromosome number in Gondwanan angiosperm families of Australia, especially Rutaceae and Proteaceae. In: Brandham, P. E. and Bennett, M. D. (eds) Kew Chromosome Conference IV, pp. 223–232. Royal Botanic Gardens, Kew.
Swofford, D. L. and Selander, R. B. (1981). BIOSYS-1: A FORTRAN program for the comprehensive analysis of electrophoretic data in population genetics and systematics. J Hered, 72: 281–283.
Systma, K. J. and Schaal, B. A. (1985). Genetic variation, differentiation and evolution in a species of complex tropical shrubs based on isozymic data. Evolution, 39: 582–593.
Whitkus, R. (1985). FORTRAN program for computing genetic statistics from allelic frequency data. J Hered, 76: 142–142.
Wright, S. (1978). Evolution and the Genetics of Populations, vol. 4, Variability Within and Among Natural Populations. The University of Chicago Press, Chicago.
Yeh, F. C., Yang, R. -C. and Boyle, T. (1997). POPGENE version 1.21, Microsoft Windows-based freeware for population genetic analysis. University of Alberta, Edmonton.
Acknowledgements
The authors wish to thank Leith Kohler for field assistance and Shauna Roche for additional information. Thanks also to M. Waycott, J. Bussell, M. Byrne and two anonymous reviewers for their constructive comments on the manuscript. This research was undertaken while L.M.B. held an Australian Postgraduate Award at Curtin University of Technology, Perth.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Broadhurst, L., Coates, D. & Tan, B. Genetic diversity in the monospecific Western Australian endemic, Geleznowia verrucosa Turcz. (Rutaceae). Heredity 82, 292–299 (1999). https://doi.org/10.1038/sj.hdy.6884860
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6884860
Keywords
This article is cited by
-
Population genetics of Philotheca sporadica (Rutaceae) to advise an offset translocation program
Conservation Genetics (2015)
-
Genetic entities and hybridisation within the Acacia microbotrya species complex in Western Australia
Tree Genetics & Genomes (2015)
-
Genetic diversity within and divergence between rare and geographically widespread taxa of the Acacia acuminata Benth. (Mimosaceae) complex
Heredity (2002)