Abstract
The bumble bee Bombus terrestris L. is a geographically variable species with a wide distribution in Europe, the near East, northern Africa, Mediterranean islands, the Canary Islands and Madeira. Based on morphological and coat colour pattern differences, the bumble bee populations of the Canary Islands and Madeira are currently treated as separate species, B. canariensis and B. maderensis, respectively. To analyse the phylogeographical associations of these bees with continental B. terrestris, one population each from four islands of the Canaries and one population from Madeira were studied. Genetic variability was assessed at nine microsatellite loci and a fragment of the mitochondrial gene cytochrome b. Genetic differentiation among islands, and between islands and the continent, was extensive. A NJ-tree based on microsatellites strongly supported the distinctness of the Canary Island populations, whereas the Madeira sample was genetically more similar to the continental populations of B. terrestris from Europe. MtDNA sequence data were in good agreement with nuclear markers. They suggest that haplotypes ancestral with respect to B. lucorum occur on the Canary Islands, whereas derived haplotypes were found on the European continent. The Madeira population shares the most common haplotype of continental B. terrestris. Nuclear and mtDNA data both indicate that bumble bees from the Canaries and Madeira do not share a common colonization history.
Similar content being viewed by others
Introduction
The bumble bee Bombus terrestris L. (Hymenoptera: Apidae) is the most intensively studied non-Apis bee. Bombus terrestris s.l. is widely distributed in continental Europe, occurs in North Africa, and has colonized most major Mediterranean and several Atlantic Islands (the Canary Islands and Madeira). Populations from the Canary Islands show a distinct colour pattern in that they lack the yellow bands on the thorax and abdomen that are typical for B. terrestris throughout a large part of the European continent (Fig. 1). Therefore, the populations from the Canary Islands were initially described as a colour morph of B. terrestris but later considered a separate species, B. canariensis (Erlandsson, 1979). Specimens from Madeira, on the other hand, are very similar, despite some minute morphological differences, to continental B. terrestris lusitanicus from the Iberian peninsula (Rasmont et al., 1986) (Fig. 1). Nevertheless, Erlandsson (1979) also described the populations from Madeira as a separate species, B. maderensis. Rasmont (1983) also lists canariensis and maderensis as separate species, although other authors still treat these taxa as subspecies (Hohmann et al., 1993). Here, we will refer to B. terrestris s.l. when referring to these several taxa. In Europe, about 800,000 colonies are currently commercially produced every year for crop pollination in the greenhouse industry (R. De Jonghe, pers. comm.). Because they originate from different stocks, commercially bred bumble bees differ genetically from local populations and the escaping sexuals may introduce foreign alleles. Therefore, the commercial use of B. terrestris s.l. for pollination should be of concern with respect to genetic changes in local populations. However, our understanding of the genetic population structure of Bombus spp. is rather limited (Estoup et al., 1996).
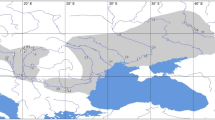
Map of western Europe and northern Africa indicating sampled populations, taxonomic status and coat colour variation of Bombus terrestris, B. canariensis and B. maderensis. The designation of taxa follows Rasmont (1983).
A preliminary genetic analysis of a population of Bombus from Tenerife (Canary Islands) revealed significantly lower levels of genetic variability at microsatellite loci and a distinct autapomorphic mtDNA haplotype (COII gene) compared to continental populations (Estoup et al., 1996). These preliminary genetic data thus suggested that the Canary Island populations may be derived from the continental ancestral stock of B. terrestris, and that these populations went through a bottleneck with reduced effective population size (Estoup et al., 1996). However, as only one population was studied, the robustness of this scenario is low. The aim of this study was to investigate the population genetic structure and colonization history of B. terrestris s.l. populations from the Canary Islands and Madeira in relation to the European continent and Mediterranean basin. In particular, the level of nuclear genetic variability within and among populations from the Canary Islands and Madeira, the genetic differentiation among Canary Island populations, and the relationships between these islands and continental populations of B. terrestris s.l. were investigated. The results support the special status of the Canary Islands populations but suggest that the colonization history of the Canary Islands has been different from that depicted by Estoup et al. (1996). The findings shed new light on the pattern of colonization and present-day population structure of an important temperate-area pollinator.
Materials and methods
Study area and biological material
The Canary Islands and Madeira are located in the Atlantic ocean at a distance of about 1400 km and 900 km, respectively, from the European continent. The Canary Islands consist of seven large islands, five of which (Tenerife, La Gomera, El Hierro, La Palma and Gran Canaria) are inhabited by Bombus canariensis. No bumble bees occur on Fuerteventura and Lanzarote, located between the African continent and the other Canary Islands (Hohmann et al., 1993). The distance between islands inhabited by B. canariensis ranges between 30 (Tenerife-La Gomera) and 210 km (El Hierro-Gran Canaria). The island ages vary considerably. K-Ar datings (Ancochea et al., 1990; Coello et al., 1992) reveal a general decrease in the age of the islands from east to west. The data suggest that Tenerife formed about 11.6 Ma BP, Gran Canaria 14–16 Ma BP, La Gomera about 10 Ma BP, and El Hierro about 1 Ma BP. None of the Canary Islands seems ever to have been connected to the African continent. The colonization of the Canary Islands by bumble bees must therefore have occurred over open water surfaces. Distances among islands, and between islands and the African continent, were smaller during the ice ages, because the sea level was lower (Hohmann et al., 1993). Madeira, which lies about 630 km off the western coast of Morocco, is of Tertiary volcanic origin and has an estimated age of 60–70 million years.
Bumble bee queens were sampled in April 1995 on Tenerife, La Gomera, El Hierro and Gran Canaria. The sample from Madeira consisted of workers which were collected as described in Estoup et al. (1996). The specimens were stored at ambient temperature in pure ethanol. The geographical origins of sampled populations are indicated in Fig. 1. To test for differences in genetic variability and amount of genetic differentiation between continental and insular populations, some of the microsatellite data from Estoup et al. (1996) were added. These came from the following populations: Avellino (Italy), Banyuls (France), Gif-sur-Yvette (France), Lublin (Poland), Madrid (Spain), Nea-Moudania (Greece), Pleven (Bulgaria) and Tours (France) from the European continent, and the Mediterranean island populations from Cyprus (Greece), Samos (Greece), Elba (Italy), Corsica (France) and Sardinia (Italy). A population of B. lucorum from Bern (Switzerland) was used as an outgroup in our analyses (Estoup et al., 1996).
DNA isolation, PCR and sequencing
DNA extractions were performed from the head of the specimens as described in Garnery et al. (1991) after soaking in sterile water for 10 min to remove any excess of ethanol. The genotypes were assessed at nine microsatellite loci: B10, B11, B96, B100, B116, B118, B124, B126 and B132. All primer sequences are given in Estoup et al. (1995, 1996). Multiplex PCRs were run for combinations of the loci B10-B11-B96, B124-B126 and B100-B118. PCR reactions were performed in a final volume of 10 μL. The reaction mixtures contained 5–10 ng of total DNA, 400 nM of each primer, 75 μM of each nucleotide dGTP, dCTP, dTTP, 6 μM dATP, 1.36×104 Bq alpha-33P-dATP, 1.0 or 1.2 mM MgCl2, 20 μg/mL Bovine Serum Albumin (BSA), 1× reaction buffer and 0.4 units of Taq Polymerase. PCR conditions were as in Estoup et al. (1996). 0.4 vol. of Formamide/EDTA/XC/BPB gel-loading buffer (Sambrook et al., 1989) was added to the PCR products prior to heat denaturation. Samples were run on 6% denaturing polyacrylamide sequencing gels for 3 h at constant voltage (2500 V) and temperature (45°C). Fixed and dried gels were exposed for 24 h.
For the study of mitochondrial DNA sequence variation, a fragment of the mitochondrial cytochrome b gene was amplified by PCR. PCR and sequencing reaction conditions were carried out as described in Pirounakis (1995). PCR fragments were produced using the mtDNA primers CB1 (coordinates 11400–11425 as in Crozier & Crozier, 1993) and CB2 (coordinates 11859–11884 as in Crozier & Crozier, 1993). In a first step, double-stranded PCR products were generated, an aliquot of which was used as template in a second PCR reaction with only one of both primers to produce a single-stranded PCR product. Ss-PCR products were sequenced using the Sequenase version 2.0 kit (USB). Sequence information was collected from 21 B. terrestris s.l., six B. canariensis, three B. maderensis and from one B. lucorum as the outgroup to root the tree.
Data analysis
Allele frequencies, observed heterozygosities and unbiased estimates of expected heterozygosities (Nei, 1978) were calculated for each population separately, using the BIOSYS-1 package (version 1.7) (Swofford & Selander, 1989). Departures from Hardy-Weinberg equilibrium (HWE) were assessed with the software package GENEPOP (version 1.2) (Raymond & Rousset, 1995). F-statistics were computed with FSTAT (Goudet, 1996) and F-values were tested for their statistical significance using permutation tests (Goudet, 1996). The appropriate permutation units depend on the F-value under consideration. For FST we permuted genotypes among populations, for FIS, alleles were permuted among individuals within populations, and for FIT, alleles were permuted among populations. Confidence intervals for F-values were obtained by jack-knifing over populations. We calculated a global FST over the Canary Islands and Madeira, and exclusively for the Canary Islands, to estimate the effective number of migrants (Nm) between Madeira and the Canary Islands, as well as among the Canary Islands themselves, using the private alleles method according to Barton & Slatkin (1985). This estimator is most appropriate when levels of gene flow are low (Barton & Slatkin, 1985). In addition, pairwise multilocus FST-values and estimates of Nm based on these FST-values, were calculated between pairs of populations of B. canariensis and B. maderensis using FSTAT.
Comparisons of genetic variability between the Canary Islands and Madeira, Mediterranean Islands and the continent were calculated for six microsatellite loci used in this study and also by Estoup et al. (1996): B11, B96, B100, B116, B118 and B132. Because the number of alleles depends on the sample size, the sample size was adjusted to 40 alleles per population, using Ewens' formula (Ewens, 1972). One-tailed Mann–Whitney U-tests were used to test for differences in Hexp. Where multiple significance tests were performed, P-values were adjusted using the sequential Bonferroni procedure to control for Type I error (Rice, 1989).
A Neighbour-Joining (NJ) tree (Saitou & Nei, 1987) relating the island and continental populations, was constructed based on microsatellite data using the chord-distance (Cavalli-Sforza & Edwards, 1967). To assess the stability of the tree nodes, we ran 2000 bootstrap replications (Hedges, 1992). All calculations were carried out using programs provided by J.-M. Cornuet (personal communication).
Analyses of phylogenetic relationships among B. terrestris s.l. mtDNA haplotypes were carried out with B. lucorum as an outgroup taxon. A maximum likelihood (ML) analysis was carried out with the program DNAML in the PHYLIP package (Felsenstein, 1993) to assess phylogenetic relationships among haplotypes. The optimal transition/transversion ratio was determined in repeated runs by maximizing the log likelihood value. To construct ML trees, an extensive search for the best tree was run, using global rearrangement. A maximum parsimony analysis (MP) was performed with the software package PAUP 3.1.1 (Swofford, 1993). Searches for all most parsimonious trees were performed with the exhaustive search algorithm assuming equal weights for transitions and transversions. The presence of phylogenetic signal in the sequence matrices was tested using the g1 statistic test based on the skewness of tree-length distributions (Hillis & Huelsenbeck, 1992). The g1 value was obtained from the exhaustive search with PAUP.
Results
All nine loci revealed genetic polymorphisms within or among the Canary Islands and Madeira populations (Table 1). Genetic variation was higher in Madeira than in the Canary Island populations (Table 2). Samples from Tenerife and Gran Canaria revealed a similar mean number of alleles per locus (2.3 and 2.5, respectively) and the same number of polymorphic loci (five each). At La Gomera and El Hierro, only three loci were polymorphic. Locus B124 was fixed for allele 50 on El Hierro, but polymorphic on La Gomera, whereas locus B126 was monomorphic on La Gomera and polymorphic on El Hierro (Table 1). Two loci (B10 and B100) showed significant deviations from HWE (P<0.001 for B10, and P=0.033 for B100), but only the deviation at locus B10 remained significant after sequential Bonferroni adjustment of P for the number of loci tested. Strong deviations from HWE at this particular locus were found in the samples from Tenerife and La Gomera, and are responsible for the positive FIS -values in these samples (Table 2). The only population where an excess of heterozygotes was observed, was Gran Canaria (FIS=−0.06).
To test whether populations from the Canary Islands and Madeira display reduced genetic variability, the mean number of alleles and expected multilocus heterozygosities were estimated from the six loci used in both this study and Estoup et al. (1996) (see Methods for list of localities). Genetic variability of the Canary Island populations (¯Hexp=0.23±0.08) was low compared to the continental populations of B. terrestris (¯Hexp=0.62±0.04), or those of Madeira (¯Hexp=0.54±0.08; one-tailed Mann–Whitney; P<0.008 for both comparisons). Similarly, the mean number of alleles for Canary Island populations (x=2.1±0.4) was lower than that found on Madeira (x=3.6±0.6) or on the continent (x=6.8±0.3; P<0.05 for both comparisons).
The Mediterranean islands investigated by Estoup et al. (1996) provide an interesting comparison. For those, the average number of alleles per locus (one-sided Mann–Whitney; P=0.006), but not heterozygosity (P=0.064), was significantly reduced relative to continental populations. But even with respect to the Mediterranean Islands, the Canary Islands populations harboured less variability (P=0.008 for expected heterozygosity and mean number of alleles). Bonferroni adjustment did not change the significance of these comparisons.
Populations of B. terrestris s.l. from the Canary Islands and Madeira were strongly differentiated from each other, as indicated by a highly significant overall FST-value (average FST=0.411, P<0.001). The corresponding average estimate of gene flow, based on the private alleles method, was thus low (Nm=0.15). Multilocus FST estimates among the four populations from the Canary Islands also revealed strong genetic differentiation (FST=0.281, P<0.001) among islands, with correspondingly low gene flow (Nm=0.18). Multilocus pairwise FST estimates between pairs of islands (Canary Islands and Madeira) revealed a relatively weak differentiation between Tenerife and La Gomera (FST=0.092), as compared to other pairs of island populations (FST≥0.274) (Table 3). As expected, when compared to continental populations, the Canary Islands and Madeira showed highly significant levels of genetic differentiation. Pairwise FST estimates between Gif, a French continental population analysed by Estoup et al. (1996) that we took here as a reference, and Tenerife or Madeira was FST=0.426, and FST= 0.107, respectively (P<0.001 for both comparisons).
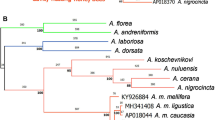
The results of the analysis of variability and differentiation are corroborated by the NJ-tree based on populations (Fig. 2). The Canary Islands represent a clearly distant group from continental B. terrestris, B. maderensis and B. lucorum (Fig. 2). The bootstrap values on loci or individuals reached 100%, confirming the differentiation of these populations. Another well supported cluster is the populations from the Tyrrhenian Islands (Elba, Corsica and Sardinia; Fig. 2), as in Estoup et al. (1996). On the other hand, Madeira surprisingly clusters with the continental and Mediterranean populations.
NJ-tree (unrooted) based on Cavalli-Sforza & Edward's (1967) distance measures, relating continental and insular populations of Bombus terrestris, B. canariensis and B. maderensis. Genetic distances were estimated from six microsatellite loci using a program provided by J.-M. Cornuet (personal communication). Small figures are bootstrap values, computed over 2000 replications.
Cytochrome b sequences revealed intraspecific mtDNA sequence variation in samples of B. terrestris s.l. Six different haplotypes with clear distribution pattern were observed (Table 4). Haplotype A was found in the five continental populations surveyed. Haplotype B was found only once in Nea-Moudania (mainland Greece). Haplotype C was observed in two queens analysed from Great Britain, suggesting that these populations of B. terrestris s.l. are distinct from those of European continent. Two haplotypes could be found on the Mediterranean islands: the continental haplotype A on Mallorca and on the three eastern Mediterranean islands (Samos, Cyprus and Crete), and haplotype D which was exclusively found on the Tyrrhenian islands. The Canary Islands harboured two endemic haplotypes, E and F. All bees from Madeira had the most common continental haplotype A.
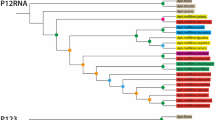
Results of the ML and MP analyses of mtDNA haplotypes were in good agreement. The tree obtained from the ML analysis (TI:TV ratio=2.5; ln likelihood=505.45) was identical with one of the 10 shortest trees (tree length=21, consistency index=0.905) obtained in the MP analysis (Fig. 3a). Other shortest trees differed in the placement of haplotypes A-D or placed the Canary Island haplotypes (E, F) as a sister group to the continental haplotypes (A-D; Fig. 3b). The g1-value (g1= −0.644, P<0.05) indicates a significantly skewed tree-length distribution, suggesting that the dataset contains a phylogenetic signal (Hillis & Huelsenbeck, 1992). All phylogenetic analyses indicated that the most primitive haplotypes are those from the Canary Islands (E and F), followed by haplotype D from the Tyrrhenian Islands. The continental and remaining Mediterranean islands, as well as Madeira, had derived haplotypes (A, B and C).
Two of 10 most parsimonious trees of Bombus terrestris s.l. mtDNA haplotypes: (a) MP identical with the tree obtained in a ML analysis with haplotypes E and F from the Canary Islands ancestral to European haplotypes (A-D); (b) haplotypes E and F form the sister group to the European haplotypes (A-D).
Discussion
The results of this study show that Canary Island populations are characterized by low variability in both allele numbers and heterozygosity, compared to the mainland populations of B. terrestris. At the same time, the mtDNA haplotypes are diverged from those of the continent. These findings are in good agreement with the preliminary study of Estoup et al. (1996). As mentioned by these authors, such low variability could be explained with a founder event by a population derived from the European continental stock. Similarly, it could reflect repeated bottlenecks of a resident population that had immigrated from the continent, coupled with small effective size over extended periods of time (Nei et al., 1975). This would explain the large genetic distance of the Canary Islands from the other populations, as estimated from microsatellite data, because bottlenecks can inflate measures of genetic distance (Chakraborty & Nei, 1977). However, when the mtDNA-sequence data are considered (Fig. 3), the pattern is more suggestive of an ancient, primary colonization of the islands and a long history of the resident population in the archipelago.
Two mtDNA haplotypes, E and F, are endemic to the Canary Islands, whereas haplotypes A, B and C are found on the European continent, most Mediterranean islands and the British Islands, respectively. The Canary Island haplotypes, and to some degree those of the Tyrrhenian Islands, are ancestral with respect to all others if the closely related taxon B. lucorum is taken as an outgroup (Fig. 3). Bombus terrestris and B. lucorum are closely related, as suggested by a phylogenetic analysis of subgenus Bombus s.str., based on mtDNA sequence variation (Widmer et al., unpublished results).
Given the close proximity to the African continent, it is most likely that B. canariensis originated from northern African B. terrestris s.l. To what extent the islands of Fuerteventura and Lanzarote, located between the African continent and Gran Canaria, and which today are not inhabited by bumble bees (Hohmann et al., 1993), acted as stepping-stones in the colonization process cannot be answered. Unfortunately, samples from northern Africa (ssp.africanus) were not available for analysis and we tacitly assumed throughout this study that the African populations are not differentiated from continental European populations at the mtDNA and microsatellite level. However, it now becomes clear that molecular data from these bees are urgently needed to understand the divergence of the Canary Island populations in particular, as well as the colonization process in the Mediterranean basin and continental Europe.
The phylogeographical analysis of mtDNA haplotypes (Avise, 1994) supports the close relationship between populations from some Mediterranean islands (Cyprus, Crete, Mallorca, Samos) and the European continent, as they all share haplotype A (Table 4). The Tyrrhenian Islands (Elba, Corsica and Sardinia), on the other hand, form a monophyletic group in the NJ-tree, based on nuclear genetic variation (Fig. 2), which is supported by the exclusive occurrence of haplotype D and high bootstrap values. The colonization history inferred for Canary Island populations based on sequence variation of the cytochrome b gene is not in agreement with the conclusions drawn by Estoup et al. (1996) based on sequence information of the gene COII (247 bp). They found a single substitution between haplotype I from the European continent and the haplotypes from the Tyrrhenian Islands and Tenerife (B. canariensis). The insular haplotypes were autapomorphic, which indicated that island haplotypes could be derived from an ancestral European continental stock (Estoup et al., 1996). In agreement with the view advocated here, an old age of Canary Island populations has also been inferred in other cases, for example, for Drosophila subobscura, where the abundance of endemic nucleomorphs and their great divergence are likely to be the result of the very old age of these Drosophila populations rather than a consequence of drift and founder effects (Afonso et al., 1990). Furthermore, the relationships among the insular populations of B. canariensis are in partial agreement with other animal taxa that have a long history on the archipelago. Phylogenetic studies on darkling beetles of the genus Pimelia (Juan et al., 1995), also found strong evidence for a colonization route from east (Fuerteventura, Tenerife) to west (La Gomera, El Hierro). A slightly different dispersal scenario has been invoked for the lizard genus Gallotia (Thorpe et al., 1994). Molecular data suggest that the western species, G. galloti, arose in Tenerife and dispersed westward in two independent pathways: north to La Palma and south to La Gomera and El Hierro (Thorpe et al., 1994). In contrast to the cases of darkling beetles and lizards, where most islands harbour endemic subspecies or species (Thorpe et al., 1994; Juan et al., 1995), differentiation of bumble bee populations has apparently not led to speciation among the different Canary Island populations, despite significant genetic differentiation. These differences may be the result of different dispersal capabilities which determine inter-island gene flow. Bombus queens have indeed been observed to migrate long distances, sometimes also over open water (Mikkola, 1984). Whether the estimated levels of gene flow among populations of B. canariensis are sufficient to prevent speciation or whether the separation of the insular gene pools is too recent for speciation to occur is not known.
In contrast, the case of Madeira seems to be different. Variability at microsatellite loci in this population was similar to Mediterranean island populations and larger than on the Canary Islands. This observation, and the occurrence of the continental and Mediterranean mtDNA haplotype A on Madeira is indicative of a different evolutionary history. The independent origin of the Madeira and Canary Island populations is also evidenced by the NJ-trees based on microsatellite markers (Fig. 2). Populations and individuals from Madeira are clearly more closely related to B. terrestris from the European continent and Mediterranean islands than to B. canariensis. Interestingly, also in the case of Drosophila subobscura, allozyme, chromosomal and mtDNA polymorphisms suggest that the Madeira populations are more related to those of the continent than to those of the Canary Islands (Larruga et al., 1983; Afonso et al., 1990). However, in contrast to B. terrestris s.l., populations of D. subobscura from Madeira carry several distinct mtDNA haplotypes (Afonso et al., 1990).
References
Afonso, J. M., Volz, A., Hernandez, M., Ruttkay, H., Gonzalez, M., Larruga, J. M. et al. (1990). Mitochondrial DNA variation and genetic structure in old-world populations of Drosophila subobscura. Mol Biol Evol, 7: 123–142.
Ancochea, E., Fuster, C. J. M., Ibarrola, E., Cendrero, A., Hernan, F. et al. (1990). Volcanic evolution of the island of Tenerife (Canary Islands) in the light of new K-Ar data. J Vol Geotherm Res, 44: 231–249.
Avise, J. C. (1994). Molecular Markers, Natural History and Evolution. Chapman & Hall, New York.
Barton, N. H. and Slatkin, M. (1985). A quasi-equilibrium theory of the distribution of rare alleles in a subdivided population. Heredity, 56: 409–415.
Cavalli-Sforza, L. L. and Edwards, A. W. F. (1967). Phylogenetic analysis: models and estimation procedures. Am J Hum Genet, 19: 233–257.
Chakraborty, R. and Nei, M. (1977). Bottleneck effects on average heterozygosity and genetic distance with the stepwise mutation model. Evolution, 31: 347–356.
Coello, J., Cantagrel, J. M., Hernan, F., Fuster, J. M., Ibarrola, E., Ancochea, E. et al. (1992). Evolution of the eastern volcanic ridge of the Canary Islands based on new K-Ar data. J Vol Geotherm Res, 53: 251–274.
Crozier, R. H. and Crozier, Y. C. (1993). The mitochondrial genome of the honeybee Apis mellifera: complete sequence and genome organization. Genetics, 133: 97–117.
Erlandsson, S. (1979). Bombus canariensis Perez, 1895 n.stat. & B. maderensis n.sp. from the Macronesian Islands. Entomol Scand, 10: 1427–1431.
Estoup, A., Scholl, A., Pouvreau, A. and Solignac, M. (1995). Monoandry and polyandry in bumble bees (Hymenoptera; Bombinae) as evidenced by highly variable microsatellites. Mol Ecol, 4: 89–93.
Estoup, A., Solignac, M., Cornuet, J.-M., Goudet, J. and Scholl, A. (1996). Genetic differentiation of continental and island populations of Bombus terrestris (Hymenoptera: Apidae) in Europe. Mol Ecol, 5: 19–31.
Ewens, W. J. (1972). The sampling theory of selectively neutral alleles. Theor Pop Biol, 3: 87–112.
Felsenstein, J. (1993). PHYLIP (Phylogeny inference package). Version 3.5c. Department of Genetics, University of Washington, Seattle.
Garnery, L., Vautrin, D., Cornuet, J. M. and Solignac, M. (1991). Phylogenetic relationships in the genus Apis inferred from mitochondrial DNA sequence data. Apidologia, 22: 87–92.
Goudet, J. (1996). FSTAT A computer program to calculate F-statistics. Version 1.2. J Hered, 86: 485–486.
Hedges, S. B. (1992). The number of replications needed for accurate estimation of the bootstrap P value in phylogenetic studies. Mol Biol Evol, 9: 366–369.
Hillis, D. M. and Huelsenbeck, J. P. (1992). Signal, noise, and reliability in molecular phylogenetic analyses. J Hered, 83: 189–195.
Hohmann, H., La Roche, F., Ortega, G. and Barquin, J. (1993). Bienen, Wespen und Ameisen der Kanarischen Inseln (Insecta: Hymenoptera: Aculeata). Uebersee-Museum, Bremen.
Juan, C., Oromi, P. and Hewitt, G. M. (1995). Mitochondrial DNA phylogeny and sequential colonization of Canary Islands by darkling beetles of the genus Pimelia (Tenebrionidae). Proc R Soc B, 261: 173–180.
Larruga, J. M., Cabrera, V. M., Gonzalez, A. M. and Gullon, A. (1983). Molecular and chromosomal polymorphism in continental and insular populations from the south-western range of Drosophila subobscura. Genetica, 60: 191–205.
Mikkola, K. (1984). Migration of wasp and bumble bee queens across the Gulf of Finland (Hymenoptera: Vespidae and Apidae). Not Ent, 64: 125–128.
Nei, M. (1978). Estimation of average heterozygosity and genetic distances from a small number of individuals. Genetics, 89: 583–590.
Nei, M., Maruyama, T. and Chakraborty, R. (1975). The bottleneck effect and genetic variability in populations. Evolution, 29: 1–10.
Pirounakis, K. (1995). Genetic Variation Among European Populations of the Bumblebee Bombus pascuorum Using Mitochondrial DNA Sequence Data. Diploma Thesis, ETH Zürich.
Rasmont, P. (1983). Catalogue commenté des bourdons de la région ouest-paléarctique (Hymenoptera, Apoidea, Apidae). Not faunist Gembloux, 7: 1–72.
Rasmont, P., Scholl, A., de Jonghe, R., Obrecht, E. and Adamski, A. (1986). Identité et variabilité des males de bourdons du genre Bombus Latreille sensu stricto en Europe occidentale et centrale (Hymenoptera, Apidae, Bombinae). Rev Suisse Zool, 93: 661–682.
Raymond, M. and Rousset, F. (1995). GENEPOP. Population genetics software for exact tests and ecumenicism. Version 1.2. J Hered, 86: 248–249.
Rice, W. R. (1989). Analyzing tables of statistical tests. Evolution, 43: 223–225.
Saitou, N. and Nei, M. (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol, 4: 406–425.
Sambrook, J., Fritsch, E. F. and Maniatis, T. (1989). Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Swofford, D. L. (1993). Phylogenetic analysis using parsimony (PAUP). Illinois Natural History Survey, Champaign, IL.
Swofford, D. L. and Selander, R. B. (1989). BIOSYS-1. A computer program for the analysis of allelic variation in population genetics and biochemical systematics. Release 1.7. Illinois Natural History Survey, Champaign, IL.
Thorpe, R. S., McGregor, D. P., Cumming, A. M. and Jordan, W. C. (1994). DNA evolution and colonization sequence of island lizards in relation to geological history: mtDNA RFLP, cytochrome B, cytochrome oxidase, 12S rRNA sequence, and nuclear RAPD analysis. Evolution, 48: 230–240.
Acknowledgements
We are indebted to many friends and colleagues who kindly provided technical and theoretical assistance and help: M. Baltisberger, E. Bucheli, J.-M. Cornuet, D. Frey, J. Koella, S. Koulianos, A. Köpf, R. Köpfli, K. Pirounakis, N. Rank and J. Shykoff. This work was part of the Swiss Priority Programme Environment and supported by grant numbers 31–32193.91 and 3100–049040.95 to P. Schmid-Hempel.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Widmer, A., Schmid-Hempel, P., Estoup, A. et al. Population genetic structure and colonization history of Bombus terrestris s.l. (Hymenoptera: Apidae) from the Canary Islands and Madeira. Heredity 81, 563–572 (1998). https://doi.org/10.1046/j.1365-2540.1998.00407.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00407.x
Keywords
This article is cited by
-
Diversity of bumble bees, their molecular analysis and first report of Bombus hypnorum L. (Hymenoptera: Apidae) from Himachal Pradesh
Vegetos (2024)
-
Morphological and genetic data suggest a complex pattern of inter-island colonisation and differentiation for mining bees (Hymenoptera: Anthophila: Andrena) on the Macaronesian Islands
Organisms Diversity & Evolution (2022)
-
Patterns of intraspecific morphological variability in soil mites reflect their dispersal ability
Experimental and Applied Acarology (2021)
-
Contrasting patterns of genetic and morphological diversity in the bumblebee Bombus lucorum (Hymenoptera: Apidae: Bombus) along a European gradient
Journal of Insect Conservation (2019)
-
Population genetics and geometric morphometrics of the Bombus ephippiatus species complex with implications for its use as a commercial pollinator
Conservation Genetics (2017)