Abstract
An alcohol dehydrogenase allele AdhAAS44 isolated from an Australian natural population of Drosophila melanogaster is associated with low activity in adult flies. A 5.0 kb insertion in the first intron between the distal and proximal promoters has been partially sequenced and shown to be homologous to the retrotransposon Tirant. The insertion is the major change in the transcriptional unit of AdhAAS44, but there is also a single nucleotide change in the adult Adh enhancer. The main phenotypic effect associated with the Tirant insertion is a reduction in alcohol dehydrogenase activity and transcript level in adult flies. There is a very much smaller reduction in 3rd instar larvae. The data indicate that Tirant affects transcription from the distal promoter. The possible mechanisms for this differential effect on transcription are discussed.
Similar content being viewed by others
Introduction
The variety and genomic distribution of transposable elements (TEs) is well documented in Drosophila melanogaster. Over 40 families of TEs have been described (Finnegan, 1989), making up about 10 per cent of the genomic DNA. TEs are important structural components of D. melanogaster heterochromatin (Pimpinelli et al., 1995) and numerous studies have demonstrated that natural populations are highly polymorphic for TE insertions (Charlesworth & Langley, 1991). Studies of restriction fragment length polymorphism (RFLP) in the region of specific gene loci, for example Adh (Aquadro et al., 1986; Jiang & Gibson, 1992a), Amy (Langley et al., 1988), and Gpdh (Symonds & Gibson, 1992) have revealed extensive sequence length variation, most of which is caused by insertions or deletions of less than 200 bp, but some result from known TEs.
Assessing the phenotypic effects of TE insertions in natural populations is of evolutionary significance from at least two aspects. The site of insertion of a TE will determine whether there is a phenotypic effect, either on the equilibrium level of a protein or, in exceptional circumstances, by producing a new protein (Wilanowski et al., 1995). Secondly, the potential for further transposition of a TE from the region of the locus is relevant to the future stability of the gene. For these reasons it is important to identify TE insertions and to assess their phenotypic effects.
Jiang & Gibson (1992a) assessed restriction endonuclease variation in the 12 kb region surrounding the Adh locus in 194 chromosomes sampled from seven Australian and six Chinese populations of D. melanogaster. They found 10 different insertions (of 0.28, 0.35, 0.4, 0.48, 0.7, 1.0, 1.5, 3.0, 4.5 and 5.0 kb) which were subsequently cloned into λEMBL3 and probed with DNA sequences from each of seven known TEs. Insertion 0.28 kb had homology with B104B (Scherer et al., 1982) and insertion 0.4 kb had homology with F101 (Dawid et al., 1981). Two of the insertions — the 1.5 kb inserted 2 kb from the 5§ end of the coding region and the 5.0 kb inserted in intron 1 — were associated with abnormal (low) ADH activity in adult flies (Jiang & Gibson, 1992b).
The first intron in the Adh gene in D. melanogaster separates two promoters which produce developmentally regulated transcripts, with the proximal transcript present in larvae until the mid to late third instar (Savakis et al., 1986). At this time transcription switches to the distal transcript until the beginning of the pupa stage, although the level of transcription is much lower than it had been for the proximal transcript (Corbin & Maniatis, 1989). In adult flies the distal transcript is the major form. Transcription of Adh is regulated by sequences 5§ to each promoter and by two major enhancers upstream from the distal promoter (Fig. 1).
The structure of the Adh gene in Drosophila melanogaster, the positions of its major regulatory elements and the location of the insertion. (a). The Adh exons are shown as black boxes with the two transcription start sites indicated by solid arrows. The Adh adult enhancer (AAE) is shown as a hatched box and the larval enhancer (ALE) as a dotted box. The binding site for P2 protein is shown as an oval. Nucleotide numbering follows Kreitman, 1983. The probes used for Northern analyses are indicated. (b). The site of the Tirant insertion is indicated in the first intron of AdhAAS44. The 4-bp duplication target site is underlined. The two TATA boxes are shown. (c) Inverse PCR strategy used to amplify part of the insert in AdhAAS44. The Adh gene, the site of the insertion and the positions of a number of restriction enzyme sites are shown. The structure of the ligated molecule, with the SspI site used for linearization is indicated. The positions of the two PCR primers are indicated with the insertion sequences represented as a thick black line.
The 5.0 kb insertion in the AdhAAS44 allele separates the two promoters and the enhancer regions from the proximal promoter. For this reason it was important to assess the effects of this insertion on ADH activity. We have identified the 5.0 kb insertion in AdhAAS44 and shown that it has differential effects on ADH activity in larvae and adult flies.
Materials and methods
Drosophila melanogaster stocks
The AdhAAS44 allele associated with low ADH activity was identified in a survey of restriction site variation made on flies collected from All Saints (Vic.) (Jiang & Gibson, 1992a). The ADH produced by this allele is electrophoretically ‘fast’. The chromosome bearing the allele is homozygous lethal and we have not been able to recombine the lethality from the Adh locus. AdhAAS44 is maintained heterozygous with Df(2L)64j, a deficiency that spans the Adh locus (Lindsley & Grell, 1968).
Heterozygotes between AdhnLA248 and AdhAAS44 were produced for experimental purposes. AdhnLA248 is an X-ray induced Adh null allele, which produces an Adh transcript 250 bp longer than normal (Chia et al., 1985b). The chromosome bearing AdhnLA248 also carries the markers cn and bw (Lindsley & Grell, 1968) and when homozygous gives white-eyed flies. Heterozygous AAS44/nLA248 larvae were harvested from mass cultures set up on riboflavin-supplemented media (0.3 g/L). Under these culture conditions heterozygotes ++/cn bw can be distinguished from cn bw/cn bw homozygotes by the colour of the Malpighian tubules. Larvae were collected at 72–80 h and 88–96 h. Adult AAS44/nLA248 flies were separated from the progeny of single-pair matings between AAS44/ nLA248 males and homozygous nlA248 virgin females by virtue of their wild-type eye colour.
A normal activity Adh allele, AC5, was used as a control; AC5 encodes electrophoretically fast enzyme.
Extraction of total DNA
Genomic DNA was extracted from whole flies according to the method described by Chia et al., 1985a
Inverse PCR
The lambda clone containing AdhAAS44 (Jiang & Gibson, 1992a) lost titre, so to identify the 5.0 kb insertion in AdhAAS44, the 5§ part of the insertion was amplified using inverse PCR (Triglia et al., 1988). The AdhAAS44 genomic DNA was digested with Sau3AI, as there were likely to be several Sau3AI sites along the 5 kb insertion fragment. There was a Sau3AI site in the first intron of Adh (Fig. 1). An SspI restriction site between the primers JG83 and JG13 was used to linearize the circularized DNA. The primer pair JG83 (position 104–184, 5′-GCCCTTTTGCTACTTACTT-3′) and JG13 (position 288–307, 5§-CCTGTTCCGCATATTCCTCT-3§), which are located between the Sau3AI site and the insertion site, were used for PCR amplification (Fig. 1).
PCR amplification and sequencing
The rest of the AdhAAS44 gene was amplified with appropriate primers described by Gibson et al. (1992). The PCR amplification protocol and the purification of amplified DNA for sequencing has been described by Reed & Gibson (1993).
ADH enzyme activity assays
Flies used for ADH assays were raised from single-pair cultures on a high-protein medium (10 g agar, 5 g yeast, 15 g sucrose, 50 g malt, 30 mL karo and 10 g hi-pro per L) at 25°C. Male progeny were collected and aged for 7–10 days at 20°C before assaying. Flies were frozen in liquid nitrogen and stored at −90°C for up to 7 days. Assay extracts were prepared and ADH activities were measured as previously described (Gibson & Wilks, 1988).
RNA extraction and quantitative Northern analysis
The methods used for total RNA extraction from adult flies for Northern analyses have previously been described (Freeth et al., 1988). Total RNA was also extracted from 72 to 80 h and 88–96 h larvae using the same procedure. The filters loaded with the adult RNA were hybridized with a nick-translated labelled pSAF-2 probe (Goldberg, 1980) and the filters loaded with the larval RNA were hybridized with a nick-translated labelled 604 bp BamHI/HindIII fragment (+651 to+1256) cloned into M13mp9 (Fig. 1, Lockett & Ashburner, 1989). Filters were also probed with a 386 bp SalI/HpaI fragment (−63 to +326), which is specific for the distal transcript (Fig. 1).
The hybridization intensities of the transcript bands were measured on the PhosphorImager instrument (Molecular Dynamics). The signal intensity of each band observed was measured separately and the relative intensity for each sample was calculated.
Results
The insertion in AdhAAS44 is Tirant
Southern analyses (Jiang & Gibson, 1992a) had shown that the 5.0 kb insertion in AdhAAS44 was located in a HpaI (−386)/HindIII (−219) fragment in the first intron between the two Adh promoters (see Fig. 1). This analysis also revealed a 200 bp insertion in a 1.45 kb EcoRI fragment, 1320 bp 5§ to the distal promoter. This region contains the Adh larval enhancer (ALE) (Corbin & Maniatis, 1990), but as we have observed a similar insertion in this position in alleles with normal activity we have not investigated it any further.
The 5.0 kb insertion in AdhAAS44 could not be amplified directly by PCR using primers in the Adh gene. Instead we used inverse PCR, with the protocol outlined in Fig. 1, to amplify part of the insertion. A 150 bp PCR product was subcloned into pBluescript SK(+) and sequenced. The sequence was found to be 97 per cent homologous (EMBL data bank) to the 5§ end of the 5.7 kb Tirant TE, first described as an insert in the extramacrochaetae (emc) locus in D. melanogaster (Garrell & Modolell, 1990) — Tirant was named after a medieval Castilian Knight. The restriction map of the insert (Jiang & Gibson, 1992a) strongly supports the conclusion that the insert is derived from Tirant, although the resolution of the map was insufficient to detect the region deleted compared to the 5.7 kb element. Later, using an Adh primer and one based on the Tirant sequence described by Garrell & Modolell (1990), we amplified by PCR and sequenced the 3§ end of the insertion in AdhAAS4. These data further confirmed that the insertion was Tirant, that it was inserted in Adh at position +406 to+409, i.e. AGCC, and that this target site was duplicated. The target site differs at two nucleotides from the target site (CCCG) previously described (Garrell & Modolell, 1990). Tirant is inserted in the same transcriptional orientation as the Adh gene.
In order to be confident that any phenotypic effects on ADH activity associated with the Tirant insertion were not confounded by nucleotide changes elsewhere resulting in amino acid substitutions, the AdhAAS44 gene was sequenced in the coding regions from the larval leader at+650 to the 3§ flanking region at+1697. In addition, intron 1 was sequenced from +288 to+409 and the region from −640 to −460, which spanned the 142 bp Adh adult enhancer (AAE) (Falb & Maniatis, 1992), was also sequenced (Fig. 1).
The DNA sequence confirmed that AdhAAS44 is a fast electrophoretic allele with C at+1490 (Kreitman, 1983). There were no other changes that would give rise to amino acid substitutions, and other differences found in comparison to the consensus sequence for AdhF had previously been described in normal activity AdhF alleles (Kreitman, 1983; Laurie et al., 1991). There were two nucleotide changes in the AAE sequence (−615 to −473) and one of these (A to T) at position −523 was in the central core region at the binding site for the transcription factor AEF-1. It is possible that this change might affect the function of the AAE.
Effect of Tirant on ADH activity and Adh transcripts
As Tirant was inserted between the two Adh promoters we assayed ADH activity in 3rd instar larvae and adult flies. The second chromosome bearing AdhAAS44 was homozygous lethal so assays were made on larvae and adults heterozygous AdhAAS44/AdhnLA248, which are also heterozygous for cn and bw carried on the nLA248 second chromosome. Larvae of the genotype ++/cn bw, when cultured on media supplemented with riboflavin, are distinguished by the dark yellow colour of their Malpighian tubules, in contrast to the pale yellow colour of the tubules in cn bw/cn bw homozygotes. These assay data (Table 1) show that, compared to the AdhF control allele, AdhAAS44 has about one-third the activity in adult flies but more than 75 per cent of the normal level in larvae. Early third instar larvae (72–78 h) have significantly (P<0.05) lower relative activity than the control, compared to late (90–96 h) third instar larvae.
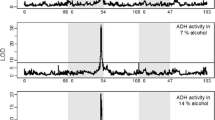
The relative transcript levels between AdhAAS44 and the control (Fig. 2 and Table 2) paralleled the ADH activity differences, although for each developmental stage the difference was less. Thus, in adults, AdhAAS44 produced about 55 per cent of the transcript level of the control compared to 34 per cent of the ADH activity.
Northern analyses for adults and larvae of AdhAAS44/AdhnLA248 and AdhAC5/AdhnLA248 heterozygotes. From left to right, samples were loaded as: AdhAC5 adults, AdhAAS44 adults, AdhAC5 88–96 h larvae, AdhAAS44 88–96 h larvae, AdhAC5 72–80 h larvae and AdhAAS44 72–80 h larvae. The adult filters were probed with the nick-translated labelled pSAF-2 and the larval filters were probed with the nick-translated labelled BSBH6. The upper band represents the nLA248-specific RNA. The lower band represents the normal sized Adh-specific RNA.
Discussion
The sequence data for the 5§ region, together with the restriction map, indicate that the insertion in AdhAAS44 is Tirant (Garrell & Modolell, 1990). This insertion, located in the first intron of Adh is the first example of a TE insertion in the Adh transcriptional unit in alleles sampled from natural populations. Previously Aquadro et al. (1986) found a low-activity Adh allele (R142) with a 5.2 kb copia retrotransposon inserted 243 bp 5§ to the distal transcription start site. Dunn & Laurie (1995), using P element transformation of deletion constructs, showed that excision of copia from the cloned R142 gene caused a threefold increase in ADH activity.
Unlike copia (Mount & Rubin, 1985), the Tirant TE is less well studied, with only two Tirant insertions previously described. The limited data suggest that Tirant has a number of structural features common to Drosophila retrotransposons. For example, the 4 bp target site is duplicated on insertion, the element contains moderately repetitive DNA and has 5 bp inverted terminal repeats (Garrell & Modolell, 1990). Recently, Thomas et al. (1995) have described a spontaneous dominant mutation at the Serrate locus (SerD) caused by a 505 bp Tirant insertion. From comparison of the sequence with that described by Garrell & Modolell (1990), the authors conclude that Tirant has 416 bp LTRs that terminate with 5 bp inverted repeats and is thus a typical retrotransposon. There are no data on the genomic copy number.
The Tirant associated with the Ach mutation at the extramacrochaetae locus is inserted such that it truncates the emc ORF. The functional mutant protein encoded lacks 42 carboxy-terminal amino acids present in the wild-type protein (Garrell & Modolell, 1990). In SerD, Tirant is inserted in the 3§ untranslated region of the gene. The truncation of the Ser RNA within the Tirant LTR eliminates putative RNA degradation signals downstream, which leads to increased stability of Ser RNA, higher levels of Serrate protein and a complex phenotype (Thomas et al., 1995).
The main phenotypic effect associated with AdhAAS44 is an ≈70 per cent reduction in ADH activity in adult flies but less reduction (≈20 per cent) in 3rd instar larvae, and these changes are paralleled by changes in transcript levels. The probe used in the Northern blots identified transcripts both from the distal and proximal promoters. Our attempts to quantify transcripts in larvae from the distal promoter only using a 386 bp distal transcript specific probe (Fig. 1, Lockett & Ashburner, 1989) were not successful, so we cannot differentiate transcripts from the two promoters in the late 3rd instar larvae.
The sequence data indicated that only the Tirant insertion in the first intron and a single nucleotide change in the AAE region at position 523 differed from normal-activity alleles. The change in the AAE might affect the fidelity of the binding site and thus reduce adult activity, but this can only be gauged by further experiments, using P element transformation of appropriate constructs. As the reduction in ADH activity and transcript levels is substantial in adults, it seems more likely that Tirant is the causative factor. The Tirant insertion is 28 bp 5§ to the binding site of the P2 protein (see Fig. 1), which is a transcriptional regulatory factor of the proximal promoter (Heberlein et al., 1985), and this may be the cause of the small effect in larvae. Tirant also distances the larval enhancers (ALE) from the proximal promoter and this could also lead to a reduction in proximal transcript levels. Nevertheless, the major phenotypic effect seems to be brought about by the effect of Tirant on transcription from the distal promoter. It is possible that this effect is mediated by a lowered rate of transcription or by the lower stability of the much larger primary transcript that will be initiated from the distal promoter. There is, however, no known way of deleting the Tirant insertion from AdhAAS44 to test directly for an effect on Adh transcription. As occurs with SerD, it is possible that the poly(A) signal (AATAAA) 100 bp downstream of the 5§ termini of Tirant, might be functional in Adh transcription initiated from the distal promoter, given that Tirant is in the same transcriptional orientation as Adh. However, the Northerns did not reveal such a truncated transcript (of about 500 bp) that would be expected if transcription terminated within Tirant. Finally it should be considered whether sequences within Tirant are binding sites for factors which normally enhance Adh transcription, possibly modifying transcript levels in different tissues, as occurs with the gypsy insertion in the yellow locus (Geyer & Corces, 1992).
Whatever the mechanism turns out to be, the Tirant insertion in the Adh locus is associated with a differential effect on ADH activity with a much larger reduction in adult flies. This phenotypic effect on ADH may be associated with reduced fitness (Van Delden, 1982), although there is some evidence that larval ADH levels are more relevant to fitness than are adult levels (Heinstra et al., 1989). The AdhAAS44 allele provides another example of the kind of effects that transposable elements can induce in natural populations and hence contribute to the variation on which selection might act.
References
Aquadro, C. F., Desse, S. F., Bland, M. M., Langley, C. H. and Andlaurie-Ahlberg, C. C. (1986). Molecular population genetics of the alcohol dehydrogenase gene region of Drosophila melanogaster. Genetics, 114: 1165–1190.
Charlesworth, B. and Langley, C. H. (1991). Population genetics of transposable elements in Drosophila. In: Selander, R. K., Clark, A. G. and Whittam, T. S. (eds) Evolution at the Molecular Level, 150–176. Sinauer, Sunderland, MA.
Chia, W., Savakis, C., Karp, R., Pelham, H. and Ashburner, M. (1985a). Mutation of the Adh gene of Drosophila melanogaster containing an internal tandem duplication. J Mol Biol, 186: 679–688.
Chia, W., Karp, R., McGill, S. and Ashburner, M. (1985b). Molecular analysis of the Adh region of the genome of Drosophila melanogaster. J Mol Biol, 186: 689–706.
Corbin, V. and Maniatis, T. (1989). Role of transcriptional interference in the Drosophila melanogaster Adh promoter switch. Nature, 337: 279–282.
Corbin, V. and Maniatis, T. (1990). Identification of cis-regulatory elements required for larval expression of the Drosophila melanogaster alcohol dehydrogenase gene. Genetics, 124: 637–646.
Dawid, I. B., Long, E. O., Dinocera, P. P. and Pardue, M. L. (1981). Ribosomal insertion-like elements in Drosophila melanogaster are interspersed with mobile sequences. Cell, 25: 399–408.
Dunn, R. C. and Laurie, C. C. (1995). Effects of a transposable element insertion on alcohol dehydrogenase expression in Drosophila melanogaster. Genetics, 140: 667–677.
Falb, D. and Maniatis, T. (1992). A conserved regulatory unit implicated in tissue-specific gene expression in Drosophila and man. Genes Devel, 6: 454–465.
Finnegan, D. J. (1989). Eukaryotic transposable elements and genome evolution. Trends Genet, 5: 103–107.
Freeth, A. L., Gibson, J. B. and Wilks, A. V. (1988). Transcription analysis of alcohol dehydrogenase null alleles from natural populations of Drosophila melanogaster. Genome, 30: 25–30.
Garrell, J. and Modolell, J. (1990). The Drosophila extramacrochaetae locus, an antagonist of proneural genes that, like these genes, encodes a helix-loop-helix protein. Cell, 61: 39–48.
Geyer, P. K. and Corces, V. (1992). DNA position-specific repression of transcription by a Drosophila zinc finger protein. Genes Devel, 6: 1865–1873.
Gibson, J. B. and Wilks, A. V. (1988). The alcohol dehydrogenase polymorphism of Drosophila melanogaster in relation to environmental ethanol, ethanol tolerance and alcohol dehydrogenase activity. Heredity, 60: 403–414.
Gibson, J. B., Wilks, A. V. and Agrotis, A. (1992). Molecular relationships between alcohol dehydrogenase null-activity alleles from natural populations of Drosophila melanogaster. Mol Biol Evol, 9: 250–260.
Goldberg, D. A. (1980). Isolation and partial characterisation of the Drosophila alcohol dehydrogenase gene. Proc Natl Acad Sci USA, 77: 5794–5798.
Heberlein, U., England, B. and Tjian, R. (1985). Characterization of Drosophila transcription factors that activate the tandem promoters of the alcohol dehydrogenase gene. Cell, 41: 965–977.
Heinstra, P. W., Geer, B. W., Seykens, D. and Langevin, M. (1989). The metabolism of ethanol-derived acetaldehyde by alcohol dehydrogenase (EC 1.1.1.1) and aldehyde dehydrogenase (EC 1.2.1.3) in Drosophila melanogaster larvae. Biochem J, 259: 791–797.
Jiang, C. and Gibson, J. B. (1992a). The alcohol dehydrogenase polymorphism in natural populations of Drosophila melanogaster: restriction map variation in the region of the Adh locus in populations from two hemispheres. Heredity, 68: 1–14.
Jiang, C. and Gibson, J. B. (1992b). The alcohol dehydrogenase polymorphism in natural populations of Drosophila melanogaster: ADH activity variation, restriction site polymorphism and the Adh cline. Heredity, 68: 337–344.
Kreitman, M. (1983). Nucleotide polymorphism at the alcohol dehydrogenase locus of Drosophila melanogaster. Nature, 304: 412–417.
Langley, C. H., Shrimpton, A. E., Yamazaki, T., Miyashita, N., Matsuo, Y. and Aquadro, C. F. (1988). Naturally occurring variation in the restriction map of the Amy region of Drosophila melanogaster. Genetics, 119: 619–629.
Laurie, C. C., Bridgham, J. T. and Choudhary, M. (1991). Associations between DNA sequence variation and variation in expression of the Adh gene in natural populations of Drosophila melanogaster. Genetics, 129: 489–499.
Lindsley, D. and Grell, E. H. (1968). Genetic Variations of. Drosophila melanogaster. Carnegie Institution of Washington Publication no. 627.
Lockett, T. J. and Ashburner, M. (1989). Temporal and spatial utilization of the alcohol dehydrogenase gene promoters during the development of Drosophila melanogaster. Devl Biol, 134: 430–437.
Mount, S. M. and Rubin, G. M. (1985). Complete nucleotide sequence of the Drosophila transposable element copia: homology between copia and retroviral proteins. Mol Cell Biol, 5: 1630–1638.
Pimpinelli, S., Berloco, M., Fanti, L., Dimitri, P., Bonaccorsi, S., Marchetti, E. et al. (1995). Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc Natl Acad Sci USA, 92: 3804–3808.
Reed, D. S. and Gibson, J. B. (1993). Defective P element insertions affect the expression of sn-glycerol-3-phosphate dehydrogenase alleles in natural populations of Drosophila melanogaster. Proc R Soc B, 251: 39–45.
Savakis, C., Ashburner, M. and Wills, J. H. (1986). The expression of the gene coding for alcohol dehydrogenase during the development of Drosophila melanogaster. Devl Biol, 114: 194–207.
Scherer, G., Tschudi, C., Perera, J., Delius, H. and Pirrotta, V. (1982). B104, a new dispersed repeated gene family in Drosophila melanogaster and its analogies with retroviruses. J Mol Biol, 157: 435–451.
Symonds, J. E. and Gibson, J. B. (1992). Restriction site variation, gene duplication, and the activity of sn-glycerol-3-phosphate dehydrogenase in Drosophila melanogaster. Biochem Genet, 30: 169–188.
Thomas, U., Jonsson, F., Speicher, S. A. and Knust, E. (1995). Phenotypic and molecular characterisation of SerDa dominant allele of the Drosophila gene Serrate. Genetics, 139: 203–213.
Triglia, T., Peterson, M. G. and Kemp, D. J. (1988). A procedure for in vitro amplification of DNA segments that lie outside the boundaries of known sequences. Nucl Acids Res, 16: 8186
van Delden, W. (1982). The alcohol dehydrogenase polymorphism in Drosophila melanogaster Selection at an enzyme locus. Evol Biol, 15: 187–222.
Wilanowski, T. M., Gibson, J. B. and Symonds, J. E. (1995). Retrotransposon insertion induces an isozyme of sn-glycerol-3-phosphate dehydrogenase in Drosophila melanogaster. Proc Natl Acad Sci USA, 92: 12065–12069.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, YH., Gibson, J. A Tirant insertion in the alcohol dehydrogenase locus of Drosophila melanogaster. Heredity 80, 293–299 (1998). https://doi.org/10.1038/sj.hdy.6882830
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6882830