Abstract
Colonizing events may expose organisms to physical and ecological environments found nowhere else in their range. Novel selection pressures can then influence subsequent rapid evolutionary changes. Here, I investigate the genetics of one such rapid change in the sexual signal of Polynesian field crickets, Teleogryllus oceanicus, that recently colonized the Hawaiian Islands. In Hawaii, T. oceanicus encounter a deadly parasitoid fly found nowhere else in their range. In <20 generations, a wing mutation, flatwing, that eliminates the crickets' song, an important sexual signal, but protects them from the fly, spread to >90% of males on the island of Kauai. I show, using crosses between flatwing males and females from a population that has never contained flatwings, that the song-suppressing mutation is due to a change in a single sex-linked locus. Contemporary evolution of secondary sexual characteristics has only rarely been identified as the result of single-gene changes and never before as a single sex-linked locus, but sex-linked inheritance is thought to facilitate the rapid evolution of these types of traits. Because divergence of sexual signals can influence reproductive isolation, understanding how colonization events and subsequent selection affect signals, and the genetic mechanisms of such change, can shed light on processes likely to play a role in speciation.
Similar content being viewed by others
Introduction
The colonization of a new environment places organisms in circumstances that are favorable to rapid evolutionary change (Reznick and Ghalambor, 2001). Founding populations generally consist of only a few individuals and are subject to genetic drift, encounter novel selection pressures and may experience rapid population growth in their new environments (MacArthur and Wilson, 1967; Carson and Templeton, 1984). Sexual signals, in particular, are amenable to rapid evolutionary change in such situations (Yeh, 2004; Svensson and Gosden, 2007) because founders possess a ‘first sexual signal’ that may be well or poorly fit to its new environment. Differences in the physical and ecological environment, such as, novel predators and parasites, competing signalers, and transmission properties can impose selection on the founders' sexual signal (Boake, 2002).
The conspicuousness of sexual signals to predators and parasites has been recognized by many (Darwin, 1871; Verrell, 1991; Endler, 1992), and the selective pressure of these eavesdropping natural enemies generally opposes that of the intended signal receivers (usually females). Natural enemies are often attracted to the very same elements of a signal that are preferred by females and because of this sexual signals represent a compromise between natural and sexual selection (Zuk and Kolluru, 1998). Perhaps the best-known example of this comes from túngara frogs where both females and predatory fringe-lipped bats prefer males who include ‘chucks’ in their mating calls, yet males regularly give whining calls without chucks in areas of high bat predation (Ryan et al., 1982). It is possible, then, that varying selection pressures across a species' range (in portions of the range with and without a given predator, for instance) can lead to divergence of sexual signals.
Such differentiation of mating signals can contribute to the reproductive isolation of organisms, potentially leading to speciation (West-Eberhard, 1983; Servedio and Noor, 2003). Thus, understanding the influence of colonization and subsequent selection on signal evolution can shed light on the processes that contribute to reproductive isolation. When sexual signals do change, we want to understand not only the forces responsible (for example, adaptive or non-adaptive), but also the underlying genetic architecture. This is important because understanding how changes in sexual signals are controlled genetically will identify types of genetic change involved in reproductive isolation.
Relatively few examples of rapid evolutionary change in sexual signals have been well documented, and even fewer after a natural colonization event (but refer Yeh, 2004). In fact, a recent review identified only 11 studies in which investigators identified rapid evolution of sexual signals in natural populations (Svensson and Gosden, 2007). Of these, the majority are changes in continuously varying quantitative traits, for instance, male coloration, horn size or tail length (Endler, 1980; Coltman et al., 2003; Möller and Szep, 2005, respectively). Only 3 of the 11 studies are of discrete polymorphisms. They are the color polymorphism in female blue-tailed damselflies (Svensson et al., 2005), color morphs in side-blotched lizards (Sinervo and Lively, 1996; Sinervo et al., 2000) and the normal-wing/flatwing polymorphism in Polynesian field crickets (Zuk et al., 2006). Color polymorphisms in both side-blotched lizards and blue-tailed damselflies have been identified as autosomal single-gene traits, the latter with three alleles and a hierarchy of dominance (Sinervo and Lively, 1996; Sinervo et al., 2000; Svensson et al., 2005). Here, I investigate the mode of inheritance of the third example, the normal-wing/flatwing polymorphism of Hawaiian Teleogryllus oceanicus. I use crosses between flatwing males and females from a population that has never contained flatwings to determine the mode of inheritance of the song-suppressing mutation, flatwing.
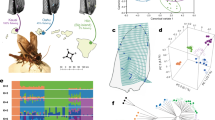
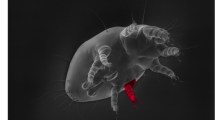
The Polynesian field cricket, T. oceanicus, was recently introduced to Hawaii (Kevan, 1990), where it encounters a deadly parasitoid fly, Ormia ochracea, that is found nowhere else in its range (Lehmann, 2003). This novel natural enemy is attracted to the crickets' mating signal, the calling song (Cade, 1975; Zuk et al., 1993), and is known to have influenced the crickets' song structure and mating behavior in Hawaii (Zuk et al., 1993, 1998, 2001; Lewkiewicz and Zuk, 2004). Gravid female flies deposit larvae on and around calling males, some of which will burrow into the body cavity of the cricket and consume fat and muscle tissue for the next 7–10 days, before eventually emerging and killing the cricket host in the process (Zuk et al., 1993; Adamo et al., 1995). Recently, strong selective pressure by the fly favored a silent cricket morph called ‘flatwing’ that lacks the wing structures used to produce song (Figure 1; Zuk et al., 2006). The advantage of this seemingly maladaptive trait is that flatwing males are ‘invisible’ to the flies, which solely use song to locate their hosts (Cade, 1975; Zuk et al., 2006). To secure matings, instead of calling themselves, flatwing males pursue a satellite strategy, honing in on the song of the remaining calling males and attempting to mate with females attracted to them (Zuk et al., 2006).
Wing morphology of T. oceanicus. Forewings from a (a) flatwing male, (b) normal-wing male and (c) female. Flatwing males lack the file (1) and scraper as well as the resonating structures of the wing (harp (2) and mirror (3)). Flatwing morphology is not, however, identical to that of females. Instead, flatwing males possess a vestigial file that is reduced in size and relocated on the wing but visible by scanning electron microscopy (Zuk et al., 2006). Male crickets normally produce song via stridulation, whereby a file on one forewing strikes the scraper of the other (Bennet-Clark, 1989) and a pulse of sound is produced with each wing-closing motion (Walker, 1962).
The rise of flatwing morphology from negligible in the late 1990s to 91% of the population in 2004 took only 16–20 generations (Zuk et al., 2006). Laboratory populations bred from eggs collected on Kauai have been maintained in the lab in a constant environment and have shown flatwings for 12 generations (suggesting that the trait is genetically controlled and not the result of the rearing environment). No intermediate forms have been observed in the field or laboratory stocks (personal observation). Two possible modes of inheritance exist: either the difference between flatwing and normal-wing morphology is controlled by one or a few genes inherited in Mendelian fashion or the trait is polygenic with a threshold of expression giving rise to discrete morphs (threshold trait; Roff, 1996). Given the rapid spread of the flatwing mutation and the lack of intermediate types, I hypothesized that the polymorphism is controlled by a single locus with two alleles. The initial rise in allelic frequency is fastest when an allele is dominant, since a mutant recessive allele would have to arise twice in the population before ever being expressed (Hedrick, 2000). I therefore also hypothesized that the allele for flatwing morphology is dominant. The locus could be autosomal or sex-linked.
Because Zuk et al. (2006) were fortuitous in observing the rapid change in singing ability in the Kauai population, I can link the evolution of a sexual signal following colonization with the genetic architecture of the trait and the speed of the evolutionary change. My results are discussed in the contexts of the genetic architecture of changes in sexual signals, the genetics of reproductive isolation and invasion success.
Methods
A preliminary study was conducted to ensure that cross-population matings with flatwing males would produce viable offspring and to identify breeding methodology that would maximize survivorship (Methods and Results of this study can be found in an online supplement to Zuk et al., 2006). However, the sample size and crosses performed were insufficient to confirm the mode of inheritance of the flatwing trait. Four replicate F1 crosses between flatwing males and virgin females from a population with no flatwing males were performed, and in the F2 generation only one of these replicates produced enough offspring for statistical evaluation. In that cross, the frequency of flatwing and normal-wing males in the F2 generation approximated a 1:1 ratio, consistent with the inheritance of a single sex-linked locus (χ2=0.032, P=0.8575). Knowing this aided in the design of the current breeding experiment by elucidating the importance of the sex determination system. The sex determination system in crickets is XX/XO with males having only one X chromosome (Hewitt, 1979). Thus, if the song-suppressing mutation is X-linked, males are hemizygous for the flatwing trait. Additionally, and importantly, the flatwing trait is sex-limited (only males have the organs necessary to produce song, females are monomorphic), so we cannot identify the phenotype of females as ‘flat-winged’ or ‘normal-winged’.
Population origins
An archival ‘Kauai’ stock population consisting of only normal-wing males has been maintained in the laboratory since 1992 and was supplemented roughly annually with the offspring of field-caught females until 2001 (the last year in which the population was sampled and no flatwing males found). This stock population has never contained any flatwing males. A flatwing stock was begun in the summer of 2004 with the offspring of field-caught females from the very same field site on Kauai and supplemented in 2005 and 2006. In 2005 and 2006, nearly all male hatchlings emerging from the eggs of field-caught Kauai females had flatwing morphology, so I refer to this as the flatwing population. All stock lab populations consist of at least 100 adults.
Breeding
Breeding experiments were conducted in a growth chamber at 30 °C on a 12:12 light/dark schedule. To produce the F1 generation, virgin females from the pure-breeding normal population (archival Kauai) were removed from mixed-sex groups prior to the final molt and housed in plastic containers (15 × 22 × 13 cm high) in groups of 5–10 with ad libitum access to food (Fluker's cricket chow for juveniles and Purina rabbit chow upon eclosion), water from wet cotton and cardboard egg carton for cover. At least 7 days post-eclosion, each female was placed in an identical plastic container for 24 h with a single flatwing male and allowed to mate. Sexually mature flatwing males used in the crosses were chosen haphazardly from the flatwing lab stocks, and no males or females were re-used. Because the flatwing trait is expressed only in males and we cannot identify females as ‘flatwinged’ or ‘normal-winged’, I did not perform the reciprocal crosses between assumed ‘flatwing’ females and normal-winged males. Mated females remained in these containers with ad libitum food, water and access to wet cotton for oviposition for 72 h after the male had been removed. After this 72 h period, females were removed and each egg pad was placed in a larger (41 × 30 × 20 cm high) plastic box where eggs were raised (approximately 30 individuals per box). In the preliminary study (Zuk et al., 2006, online supplement), this minimized subadult death during molts. Upon hatching, boxes were cleaned at least three times weekly. Each time the boxes were cleaned, I recorded the number of dead offspring. Upon eclosion, I calculated the proportion survivorship based on these observations. Males and females were separated prior to the final molt. When all adults had eclosed, I noted the wing morphology of adult males. I conducted 28 replicate matings between Kauai females and flatwing males to produce F1 offspring, and of these matings, 22 produced eggs and offspring that were successfully raised to adulthood. Replicated tests of goodness of fit (G2 likelihood ratio χ2 tests) were used to determine whether observed wing-type frequencies differed from those expected for the three possible single-gene hypotheses (Table 1). Results for each replicate were obtained simultaneously, so overall tests of goodness of fit (pooled across replicates) were carried out first, with individual replicates analyzed in heterogeneity G tests only if warranted by the data (Sokal and Rohlf, 1995).
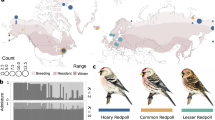
To produce the F2 generation, one virgin female and one virgin male from each of the F1 replicates (siblings) were chosen haphazardly and mated as described above (N=22 replicates). Additionally, each F1 replicate was assigned for use as either a backcross to a normal virgin female pulled from archival ‘Kauai’ lab stocks (N=9) or a backcross to a flatwing male (N=9). Some F1 replicates produced too few offspring to be used in F2 crosses and backcrosses. In these cases, I used the available F1 offspring only to produce F2 offspring. Breeding was conducted as above and offspring were raised until eclosion, after which I documented the ratio of wing types in the male offspring and again used G2 likelihood ratio χ2 tests to determine whether the observed frequencies of each wing type differed from those expected. Table 1 indicates the expected ratios of normal-wing:flatwing male progeny from each of the described crosses if the flatwing trait is due to a single autosomal dominant, autosomal recessive or sex-linked mutation. The overall breeding design is diagrammed in Figure 2.
Summarized breeding design. Flatwing males and females from the ‘archival’ Kauai population (never contained flatwings) were mated to produce the F1 generation. F1 siblings were mated to produce the F2 generation, and each F1 replicate was used in a backcross with either a normal female or a flatwing male.
Results
I performed four different types of crosses, hereafter referred to as F1, F2, BC1 and BC2. BC1 refers to the backcrosses between an F1 male and a female from the archival Kauai population of all normal-winged males. BC2 refers to the backcrosses between F1 females and flatwing males. I achieved a mean proportion survivorship of over 80% for all crosses and all replicates, with a range of 37.5–100% survivorship. The proportion survivorship in four F1 replicates was below 50% because many hatchlings drowned in excess water that collected in their boxes. The reason for this pooled water is unknown; all boxes used were identical. Complete survivorship data are available as Supplementary Information at Heredity's website.
Tests for goodness of fit to the single autosomal dominant, autosomal recessive and sex-linked hypotheses were performed on the data pooled across replicates for the F1, F2, BC1 and BC2 cross types. Sokal and Rohlf (1995) suggest pooling when the data from all replicates are obtained simultaneously. I corrected for multiple comparisons using a Bonferroni correction (α=0.0125). I could not perform goodness-of-fit tests when the expected ratio of normal-wing/flatwing males was either 0:1 or 1:0 because this places a 0 in the denominator of the statistical equation, making the value of the test statistic undefined. Using the remaining crosses, however, I was able to systematically eliminate all but one possible mode of inheritance.
In the F1 and BC1 crosses, I obtained a perfect or near-perfect match to the ratio of normal-wing:flatwing males expected if the trait is controlled by a single sex-linked or a single autosomal recessive locus (Table 1 and 2). In the F2 and BC2 crosses, the observed ratio of normal-wing:flatwing male offspring did not differ significantly from that expected if the flatwing trait is controlled by a single sex-linked locus (F2 and BC2, respectively: G2=0.3656, d.f.=1, P=0.5454; G2=0.5097, d.f.=1, P=0.4753). The sex-linked and autosomal recessive hypotheses differ only in their expectations for the F2 generation (Table 1). The observed frequency of normal-wing/flatwing males in the F2 generation, however, did not approximate a 3:1 ratio (G2=83.50, d.f.=1, P<0.0001) as would be expected with autosomal recessive inheritance. Additionally, the ratio of normal-wing:flatwing male offspring in the F2 and BC1 crosses did not approximate the 1:3 and 1:1 ratios expected for autosomal dominant inheritance (F2 and BC1, respectively: G2=107.6731, d.f.=1, P<0.0001; G2=235.67, d.f.=1, P<0.0001), eliminating this mode of inheritance as well.
Discussion
Sexual signals are amenable to rapid evolutionary change after colonization events because novel selective agents like predators and parasitoids, transmission properties of the environment, and competing signalers may differ from those in their source populations (Boake, 2002; Yeh, 2004). Divergence of sexual signals is thought to lead to reproductive isolation (West-Eberhard, 1983). Thus, understanding the role of colonization and subsequent selection on sexual signals, and the genetic changes responsible for their divergence, can inform us about types of changes likely to influence speciation. Here, I investigated the genetics of a rapid evolutionary change in the presence of song in Polynesian field crickets. This change appears to be a response to a deadly parasitoid fly (Zuk et al., 2006) found only in the portion of the range where crickets were introduced, the Hawaiian Islands. I found that the song-suppressing mutation, flatwing, which completely eliminates the ability of Kauai males to produce song, an important sexual signal, is due to a change in a single sex-linked locus. Sex determination in field crickets is XX/XO with males having only one X chromosome (Hewitt, 1979), so males are hemizygous for the flatwing trait. Calling morphology is also sex-limited; females never express the trait.
We expect the rate of evolution to be slower for complex, or polygenic, traits because of the potential for interactions between genes (Orr, 2000). Perhaps it is not surprising, then, that although cricket song itself is known to be a polygenic trait (Shaw, 1996; Gray and Cade, 1999), the song-suppressing mutation, flatwing, is due to a change in a single locus. Indeed, Zuk et al. (2006) observed the rapid spread of the flatwing mutation from negligible to >90% of Kauai males in <20 generations. The flatwing mutation is not part of the quantitative genetic background of song itself but, instead, a morphological mutation that eliminates males' ability to produce this sexual signal. Previous work in this system has identified changes in song structure, the timing of signaling and associated behaviors, which are consistent with selection to avoid the deadly parasitoid fly (Zuk et al., 1993, 1995, 1998; Lewkiewicz and Zuk, 2004). So, clearly, song itself is also subject to intense selection in this system. The suppression of calling ability in a part of the population (flatwing males) protects the quantitative genetic background of song from the effects of selection, but other factors such as mutational input and drift may have stronger impacts on future evolution than they did before the suppressor arose. Similarly, female choice for components of male song might be expected to evolve not only in response to the suppression of song in flatwing males, but also in response to subsequent changes in the song itself.
Of the 11 examples of contemporary evolutionary change in a sexual signal identified by Svensson and Gosden (2007), 3 are of discrete polymorphisms like the flatwing/normal-wing polymorphism. The other two, rapid changes in the frequencies of color morphs in blue-tailed damselflies and side-blotched lizards, have been identified as changes in single genes, however, not in sex-linked genes (Sinervo and Lively, 1996; Sinervo et al., 2000; Svensson et al., 2005). A rapid rise in the frequency of X-linked mutations might be expected if the mutation is favorable when expressed in the hemizygous sex (Avery, 1984) because such genes are expressed more often in the hemizygous sex (usually males), especially when rare (Avery, 1984). So the mode of inheritance identified for the flatwing mutation is consistent with the rapid spread of the trait that Zuk et al. (2006) observed on Kauai.
Here, I have identified a mutation that is sex-linked and suppresses the expression of sex-limited wing morphology used to produce the sexual signal, song. A large body of literature suggests that the influence of X-linked genes should be greater on sexually selected (often sex-limited) traits than autosomal genes, and in some studies this has been the case (Reinhold, 1998; Hurst and Randerson, 1999; Saifi and Chandra, 1999). When sexual selection favors trait expression in hemizygous males, but not in females, X-linked loci evolve more quickly (Rice, 1984; Charlesworth et al., 1987) and are more likely to persist than autosomal loci (Reinhold, 1999). Among studies searching for large X-effects on sexually selected traits, results are mixed, particularly in QTL analyses, possibly because these studies also identify the epistatic effects of X-linked loci on autosomal loci (Fairbairn and Roff, 2006). Large X-effects have been noted in studies of the inheritance of mating signals and courtship behavior as well as in postzygotic reproductive isolation (Coyne and Orr, 1989; Hollocher and Wu, 1996; Yeh et al., 2006). It remains to be seen whether the sex-limited wing morphology used to produce the song is controlled by genes found on the X-chromosome as well, or whether the flatwing mutation simply interrupts a pathway leading to the normal development of wing structure.
Most empirical studies of the quantitative genetics of reproductive isolation have identified polygenic effects, rather than single-gene effects, as being responsible for sexual incompatibility (Gleason et al., 2005). For instance, wing spot formation and courtship displays in male Drosophila have been identified as polygenic and attributed to the X-chromosome and autosomal loci, respectively (Yeh et al., 2006); cuticular hydrocarbons are polygenic as well (Coyne et al., 1994; Gleason et al., 2005). In swordtail crickets (Laupala; Shaw, 1996) as well as field crickets (Gray and Cade, 1999), male song is a polygenic trait, and it is often the case that closely related species differ only in their songs. Is the flatwing mutation a single-gene change that will contribute to reproductive isolation? Experiments examining the mating success of flatwings with females from populations throughout the crickets' range are underway, and these should reveal whether the loss of this important mating signal is reproductively isolating.
The flatwing/normal-wing polymorphism is unique in that the mutation was observed to completely eliminate, rather than modify, the sexual signal of >90% of T. oceanicus males found on Kauai. The loss of sexual signals, while counterintuitive, is widespread (reviewed by Wiens, 2001). For instance, phylogenetic studies reveal that conspicuous male coloration has been lost repeatedly in birds (Petersen, 1996; Price and Birch, 1996), lizards (Wiens, 1999) and fish (Endler, 1980; Reimchen, 1989). Three types of explanations for the loss of male sexual traits are possible, including environmental factors such as predation risk, random factors such as genetic drift and social factors such as male–male competition and female choice (Wiens, 2001). For a sexually selected trait to be lost, positive female choice must be overcome by random or environmental effects or female preference must be lost, reduced or reversed. A combination is also possible: the preference may be lost, then the trait may be lost because of natural selection or drift.
It seems, then, that behavioral variation or plasticity (particularly in female preferences) might play a role in the loss of sexual signals. As traits that are necessarily accompanied by a set of behaviors (of the signaler and the receiver), one might expect sexual signals and associated behaviors to change in concert. Here, the selective pressure of a deadly natural enemy favored the flatwing mutation because it protects signalers. However, behavior must have facilitated the rapid rise of the song-suppressing mutation. In field crickets, song is used by males in two mating contexts: (1) calling song attracts locomotory females from afar and (2) courtship song is normally required by females before they will mount for mating (Burk, 1983; Libersat et al., 1994; Balakrishnan and Pollack, 1996). If female choosiness was not relaxed or already variable, such that matings with silenced males were acceptable, the flatwing mutation would have been eliminated. This suggests that single-gene changes can have dramatic effects on the evolution of organisms, particularly when working in concert with behavior. Ongoing experiments are investigating whether pre-existing plasticity, in male or female behavior, might have facilitated the spread of the flatwing mutation. This type of behavioral compensation may be an important mechanism for invading populations when rapid evolutionary changes in sexual signaling, or other types of traits, occur soon after colonization (Yeh and Price, 2004).
References
Adamo SA, Robert D, Hoy RR (1995). Effects of a tachinid parasitoid, Ormia ochracea, on the behaviour and reproduction of its male and female field cricket hosts (Gryllus spp.). J Ins Phys 41: 269–277.
Avery PJ (1984). The population genetics of haplo-diploids and X-linked genes. Genet Res 44: 321–342.
Balakrishnan R, Pollack GS (1996). Recognition of courtship song in the field cricket, Teleogryllus oceanicus. Anim Behav 51: 353–366.
Bennet-Clark HC (1989). Songs and the physics of sound production. In: Huber F, Moore TE, Loher W (eds). Cricket Behavior and Neurobiology. Cornell University Press: Ithaca. pp 227–261.
Boake CRB (2002). Sexual signaling and speciation, a microevolutionary perspective. Genetica 116: 205–214.
Burk T (1983). Male aggression and female choice in a field cricket (Teleogryllus oceanicus): the importance of courtship song. In: Gwynne DT, Morris GK (eds). Orthopteran Mating Systems: Sexual Competition in a Diverse Group of Insects. Westview Press: Boulder. pp 97–119.
Cade W (1975). Acoustically orienting parasitoids: fly phonotaxis to cricket song. Science 190: 1312–1313.
Carson HL, Templeton AR (1984). Genetic revolutions in relation to speciation phenomena: the founding of new populations. Annu Rev Ecol Syst 15: 97–131.
Charlesworth B, Coyne JA, Barton NH (1987). The relative rates of evolution of sex chromosomes and autosomes. Am Nat 130: 113–146.
Coltman DW, O'Donoghue P, Jorgenson JT, Hogg JT, Strobeck C, Festa-Bianchet M (2003). Undesirable evolutionary consequences of trophy hunting. Nature 426: 655–658.
Coyne JA, Crittenden AP, Mah K (1994). Genetics of a pheromonal difference contributing to reproductive isolation in Drosophila. Science 265: 1461–1464.
Coyne JA, Orr HA (1989). Two rules of speciation. In: Otte D, Endler JA (eds). Speciation and Its Consequences. Sinauer Associates: Sunderland, MA. pp 180–207.
Darwin C (1871). The Descent of Man and Selection in Relation to Sex. John Murray: London.
Endler JA (1980). Natural selection on color patterns in Poecilia reticulata. Evolution 34: 76–91.
Endler JA (1992). Signals, signal conditions, and the direction of evolution. Am Nat 139: S125–S153.
Fairbairn DJ, Roff DA (2006). The quantitative genetics of sexual dimorphism: assessing the importance of sex-linkage. Heredity 97: 319–328.
Gleason JM, Jallon J-M, Rouault J-D, Ritchie MG (2005). Quantitative trait loci for cuticular hydrocarbons associated with sexual isolation between Drosophila simulans and D. sechellia. Genetics 171: 1789–1798.
Gray DA, Cade WH (1999). Quantitative genetics of sexual selection in the field cricket Gryllus integer. Evolution 53: 848–854.
Hedrick PW (2000). Genetics of Populations, 2nd edn. Jones and Bartlett: Sudbury, MA.
Hewitt GM (1979). Insecta 1. Orthoptera: Grasshoppers and Crickets, 1st edn. Gebruder Borntraeger: Berlin.
Hollocher H, Wu C-I (1996). The genetics of reproductive isolation in the Drosophila simulans clade: X vs autosomal effects and male vs female effects. Genetics 143: 1243–1255.
Hurst LD, Randerson JP (1999). An exceptional chromosome. Trends Genet 15: 383–385.
Kevan DKM (1990). Introduced grasshoppers and crickets in Micronesia. Bol San Veg 20: 105–123.
Lehmann GUC (2003). Review of biogeography, host range and evolution of acoustic hunting in Ormiini (Insecta, Diptera, Tachinidae), parasitoids of night-calling bushcrickets and crickets (Insecta, Orthoptera, Ensifera). Zool Anz 242: 107–120.
Lewkiewicz DA, Zuk M (2004). Latency to resume calling after disturbance in the field cricket, Teleogryllus oceanicus, corresponds to population-level differences in parasitism risk. Behav Ecol Sociobiol 55: 569–573.
Libersat F, Murray JA, Hoy RR (1994). Frequency as a releaser in the courtship song of two crickets, Gryllus bimaculatus (de Geer) and Teleogryllus oceanicus: a neuroethological analysis. J Comp Physiol A 174: 485–494.
MacArthur RH, Wilson EO (1967). The Theory of Island Biogeography. Princeton University Press: NJ.
Möller AP, Szep T (2005). Rapid evolutionary change in a secondary sexual character linked to climatic change. J Evol Biol 18: 481–495.
Orr H (2000). Adaptation and the cost of complexity. Evolution 54: 13–20.
Petersen AT (1996). Geographic variation in sexual dichromatism in birds. Bull Br Ornithol Club 116: 156–172.
Price T, Birch GL (1996). Repeated evolution of sexual color dimorphism in passerine birds. The Auk 113: 842–848.
Reimchen TE (1989). Loss of nuptial coloration in threespine sticklebacks (Gasterosteus aculeatus). Evolution 43: 450–460.
Reinhold K (1998). Sex linkage among genes controlling sexually selected traits. Behav Ecol Sociobiol 44: 1–7.
Reinhold K (1999). Evolutionary genetics of sex-limited traits under fluctuating selection. J Evol Biol 12: 897–902.
Reznick DN, Ghalambor CK (2001). The population ecology of contemporary adaptations: what empirical studies reveal about the conditions that promote adaptive evolution. Genetica 112–113: 183–198.
Rice WR (1984). Sex chromosomes and the evolution of sexual dimorphism. Evolution 38: 735–742.
Roff DA (1996). The evolution of threshold traits in animals. Q Rev Biol 71: 3–35.
Ryan MJ, Tuttle MD, Rand AS (1982). Bat predation and sexual advertisement in a neotropical anuran. Am Nat 119: 136–139.
Saifi GM, Chandra HS (1999). An apparent excess of sex- and reproduction-related genes on the human X chromosome. Proc R Soc London B 266: 203–209.
Servedio MR, Noor MAF (2003). The role of reinforcement in speciation: theory and data meet. Annu Rev Ecol Syst 34: 339–364.
Shaw KL (1996). Polygenic inheritance of a behavioral phenotype: interspecific genetics of song in the Hawaiian cricket genus Laupala. Evolution 50: 256–266.
Sinervo B, Lively CM (1996). The rock-paper-scissors game and the evolution of alternative male strategies. Nature 380: 240–243.
Sinervo B, Svensson E, Comendant T (2000). Density cycles and an offspring quantity and quality game driven by natural selection. Nature 406: 985–988.
Sokal RR, Rohlf FJ (1995). Biometry: the Principles and Practice of Statistics in Biological Research, 3rd edn. WH Freeman: San Francisco.
Svensson EI, Abbott J, Hardling R (2005). Female polymorphism, frequency dependence, and rapid evolutionary dynamics in natural populations. Am Nat 165: 567–576.
Svensson EI, Gosden TP (2007). Contemporary evolution of secondary sexual traits. Func Ecol 21: 422–433.
Verrell PA (1991). Illegitimate exploitation of sexual signaling systems and the origin of species. Ethol Ecol Evol 3: 273–283.
Walker TJ (1962). Factors responsible for intraspecific variation in the calling songs of crickets. Evolution 16: 407–428.
West-Eberhard MJ (1983). Sexual selection, social competition, and speciation. Q Rev Biol 58: 155–183.
Wiens JJ (1999). Phylogenetic evidence for multiple losses of a sexually selected character in phrynosomatid lizards. Proc R Soc London B 266: 1529–1535.
Wiens JJ (2001). Widespread loss of sexually selected traits. Trends Ecol Evol 16: 517–523.
Yeh PJ (2004). Rapid evolution of a sexually selected trait following population establishment in a novel environment. Evolution 58: 166–174.
Yeh PJ, Price TD (2004). Adaptive phenotypic plasticity and the successful colonization of a novel environment. Am Nat 164: 531–542.
Yeh S-D, Liou S-R, True JR (2006). Genetics of divergence in male wing pigmentation and courtship behavior between Drosophila elegans and D. gunungcola. Heredity 96: 383–395.
Zuk M, Kolluru GR (1998). Exploitation of sexual signals by predators and parasitoids. Q Rev Biol 73: 415–438.
Zuk M, Rotenberry JT, Simmons LW (1998). Calling songs of field crickets (Teleogryllus oceanicus) with and without phonotactic parasitoid infection. Evolution 52: 166–171.
Zuk M, Rotenberry JT, Simmons LW (2001). Geographical variation in calling song of the field cricket Teleogryllus oceanicus: the importance of spatial scale. J Evol Biol 14: 731–741.
Zuk M, Rotenberry JT, Tinghitella RM (2006). Silent night: adaptive disappearance of a sexual signal in a parasitized population of field crickets. Biol Lett 2: 521–524.
Zuk M, Simmons LW, Cupp L (1993). Calling characteristics of parasitized and unparasitized populations of the field cricket Teleogryllus oceanicus. Behav Ecol Sociobiol 33: 339–343.
Zuk M, Simmons LW, Rotenberry JT (1995). Acoustically-orienting parasitoids in calling and silent males of the field cricket Teleogryllus oceanicus. Ecol Ent 20: 380–383.
Acknowledgements
I thank Bryoni Yun, Muhammad Schuaib and Jeff Wang for assistance with colony maintenance and breeding. D Roff, C Hayashi and M Zuk provided comments on an earlier version of the manuscript and two anonymous reviewers gave valuable feedback that greatly improved the manuscript. This work was supported by NSF grants to M Zuk and RM Tinghitella.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Heredity website (http://www.nature.com/hdy)
Supplementary information
Rights and permissions
About this article
Cite this article
Tinghitella, R. Rapid evolutionary change in a sexual signal: genetic control of the mutation ‘flatwing’ that renders male field crickets (Teleogryllus oceanicus) mute. Heredity 100, 261–267 (2008). https://doi.org/10.1038/sj.hdy.6801069
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801069
Keywords
This article is cited by
-
Dissecting cricket genomes for the advancement of entomology and entomophagy
Biophysical Reviews (2022)
-
Responses of intended and unintended receivers to a novel sexual signal suggest clandestine communication
Nature Communications (2021)
-
Males and females evolve riskier traits in populations with eavesdropping parasitoids
Behavioral Ecology and Sociobiology (2018)
-
Is it the Song or the Singers? Acoustic and Social Experiences Shape Adult Reproductive Tactics and Condition
Journal of Insect Behavior (2018)
-
Combating the Assumption of Evolutionary Progress: Lessons from the Decay and Loss of Traits
Evolution: Education and Outreach (2012)