Abstract
Mismatch repair (MMR) deficiency in tumours from patients with the hereditary nonpolyposis colorectal cancer (HNPCC) syndrome is mainly caused by mutations in the MLH1, MSH2, and MSH6 genes. A major challenge in the clinical management of patients with suspected HNPCC is the frequent occurrence of missense mutations in MSH6. These can be considered neither deleterious nor clinically innocent a priori. To assess their significance we studied five novel MSH6 missense mutations in six patients derived from a series of consecutive endometrial and colorectal cancer patients selected for study after their tumours were determined to be microsatellite unstable. We tested each mutated protein for heterodimerisation with MSH2 and for in vitro MMR capability. Four mutations (R128L, P623L, K728T, G881K+S) showed no impairment of these functions while the fifth (E1193K) displayed marked impairment of both functions. These results, taken together with our previous similar findings concerning six other missense mutations in MSH6, allow us to conclude that many or most missense changes in MSH6 likely are clinically innocent, whereas some missense changes such as E1193K, which lead to impaired MMR, are likely to be clinically significant, but have low penetrance.
Similar content being viewed by others
Main
The cancer susceptibility in hereditary nonpolyposis colorectal cancer (HNPCC) syndrome is associated with defective DNA mismatch repair (MMR). An inherited defect in one MMR allele and an acquired defect in the other allele lead to loss of MMR function in the respective polypeptide that accelerates tumour progression. Accordingly, lack of the mutated protein and instability at short tandem repeat sequences such as microsatellites in the genome are the main molecular characteristics of the HNPCC tumours.
More than 450 different MMR gene mutations and 100 intragenic polymorphisms, affecting mostly the MMR genes, MLH1, MSH2, and MSH6, are listed in the international HNPCC mutation database (http://www.nfdht.nl). The clinical phenotype varies between families. Contrary to most mutations affecting the MLH1 and MSH2 genes, a significant proportion of MSH6 mutations occur in HNPCC families with less typical clinical features. Carriers of MSH6 mutations often display late age at onset, carcinomas of the endometrium, and low or no microsatellite instability (MSI) in tumours (Miyaki et al, 1997; Kolodner et al, 1999; Wijnen et al, 1999; Wu et al, 1999; Parc et al, 2000; Wagner et al, 2001; Berends et al, 2002; Peterlongo et al, 2003; Cederquist et al, 2004). Moreover, one-third of MSH6 mutations are nontruncating, and so do not necessarily destroy the encoded protein. These mutations may be associated with a positive immunohistochemical (IHC) staining for the MSH6 protein, which makes them even more difficult to interpret. In the mutation database, 12 MSH6 amino-acid substitutions are listed as pathogenic mutations, while 16 nucleotide changes in the MSH6 coding sequence are listed as polymorphisms. In both groups some changes are found to segregate in putative HNPCC families, while others occur in single endometrial cancer (EC) or colorectal cancer (CRC) patients, which impedes segregation analysis and makes their interpretation impossible.
To address the question, which, if any, MSH6 missense mutations cause susceptibility to HNPCC, we previously studied the functionality of six MSH6 variants (S144I, G566R, P1087T, P1087R, R1095H, and L1354Q) (Kariola et al, 2002, 2003). Here, we expanded the study by analysing five novel MSH6 missense mutations (R128L, P623L, K728T, G881K+S, and E1193K). Distinct from the previous cases, these mutations were derived from a population-based study of EC and CRC patients whose tumours showed MSI, the main hallmark of HNPCC. Our experimental results in conjunction with the clinical characteristics of the patients provide evidence that most MSH6 missense mutations found in a population-based case series of patients with MSI positive tumours are unlikely to be deleterious.
Materials and methods
Patient specimens
MSH6 missense mutations were derived from a project comprising altogether 2000 colorectal and EC cases newly diagnosed in the main hospitals of the Columbus metropolitan area in central Ohio (population 1.5 million). In total, 19% of colorectal and 22% of endometrial tumours showed instability with at least one of five microsatellite markers (Boland et al, 1998). Blood DNA was studied for mutations in all patients with microsatellite unstable tumours. The six patients chosen for this study were the ones, in which a novel MSH6 missense mutation in the coding region was detected, no other mutations detected by DNA sequencing occurred in MSH6, MLH1, or MSH2, and the clinical significance of the change could not be evaluated based on available evidence. All studies were approved by the Institutional Review Boards of the Ohio State University and the University of Helsinki.
Microsatellite instability analysis
Tumours were microdissected as described previously (Chadwick et al, 2001) and DNA extracted from tissue that contained at least 50% tumour cells. Unaffected DNA was obtained either from a blood sample or from microdissected endometrial tissue containing no tumour cells. For the analysis, a panel of five polymorphic microsatellite markers (BAT25, BAT26, D2S123, D5S346, and D17S25 or D18S69) was used.
Mutation detection
Direct exon-by-exon sequencing of MLH1, MSH2, and MSH6 was carried out in an ABI automated sequencer as described (Chadwick et al, 2001). In MSH6, all the exons with the exception of one part of exon 4 were sequenced in a single PCR reaction (Wu et al, 1999).
Population frequency of missense changes
All five missense changes were searched for by SSCP using primers and conditions available from the authors. All SSCP variants were sequenced. The control population consisted of 140 individuals of mixed European ethnicity including 40 grandparents from the Centre d'Etude du Polymorphisme Humain collection obtained from the Coriell Institute, New Jersey.
Immunohistochemical analysis
Paraffin embedded tissue was cut into 4 μ M sections and placed on positively charged slides. The slides were placed in a 60°C oven for 1 h, cooled, and deparaffinised and rehydrated through xylenes and graded ethanol solutions to water. All slides were quenched for 5 min in a 3% hydrogen peroxide solution in methanol to block for endogenous peroxidase. Antigen retrieval was performed by a heat method for all three antibodies, in which the specimens were placed in a citric acid solution (Dako's Target Retrieval Solution, pH 6.1), for 30 min at 94°C using a vegetable steamer. After allowing slides to cool for 15 min at room temperature, slides were placed on a Dako Autostainer immunostaining system, for use with immunhistochemistry. The primary antibody was incubated for 1 h at room temperature. The primary antibodies used were MLH1, clone G168-728 (BD PharMingen) 1/60; MSH2, Ab-2 (Oncogene Research Products) 1/200; and MSH6 (Transduction Laboratories) 1/400. The detection system used for all antibodies was a labelled Streptavidin–Biotin complex. This method is based on the consecutive application of (1) a primary antibody against the antigen to be localised, (2) biotinylated linking antibody, (3) enzyme conjugated streptavidin, and (4) substrate chromogen (DAB). The tissue was protein blocked using Dako's serum free protein block prior to the primary antibody application. Endogenous avidin and biotin were blocked prior to the biotinylated-linking antibody. Slides were then counterstained in Richard Allen hematoxylin, dehydrated through graded ethanol solutions and coverslipped.
MLH1 methylation analysis
Two sections of the MLH1 promoter region were studied for methylation. Methylation-specific PCR was used to assess a region approximately 700 bp upstream of the translation initiation site (Herman et al, 1998), designated H in this report. A combination of bisulphite PCR and cleavage by restriction enzyme digestion was used to study a second region that lies immediately upstream of the translational start site (Deng et al, 1999) and designated D in this report. The methods have been described in detail previously (Nakagawa et al, 2001).
Site-directed mutagenesis and production of recombinant proteins in insect cells
Site-directed mutagenesis was used to introduce the mutations into MSH6 cDNA, which was cloned into the plasmid pFastBac1 (Gibco BRL), as described previously (Kariola et al, 2002, 2003). The primer sequences, PCR product sizes, and cloning sites are listed in Table 2. The mutated cloned fragments were verified by DNA sequencing (AbiPrism 3100 Genetic Analyzer, Applied Biosystems). The recombinant baculoviruses, which were used as expression vectors, were generated using the Bac-to-Bac system according to the manufactureŕs instructions (Gibco BRL). For protein production, Sf9 insect cells were co-transfected with MSH2 and MSH6 recombinant baculoviruses in combinations of wild-type MSH2 (MSH2-WT) with mutated MSH6 (MSH6-R128L, MSH6-P623L, MSH6-K728T, MSH6-G881K+S, or MSH6-E1193K) or with wild-type MSH6 (MSH6-WT). The co-transfection was used since both in vivo studies in mice (de Wind et al, 1999) and in vitro studies in human cells (Marra et al, 1998; Chang et al, 2000) have shown that the MSH6 protein (Msh6 in the mouse) is unstable in the absence of its partner MSH2 (Msh2). After 72 h of culture, the total protein extracts (TE), including the overexpressed MSH6 and MSH2 proteins, were prepared as described previously (Nyström-Lahti et al, 2002).
Combined co-immunoprecipitation and Western blot analysis
The combined co-immunoprecipitation and Western blot analysis was performed according to previous reports (Kariola et al, 2002, 2003). The total protein extracts, which were estimated to contain equal quantities of recombinant wild-type MSH2 protein and the immunoprecipitated samples obtained with anti-MSH6 monoclonal antibody (clone 44, BD Transduction Laboratories), were loaded on 7.5% SDS–polyacrylamide gels. After electrophoresis, the proteins were transferred onto nitrocellulose membranes (Hybond-P PVDF, Amercham Pharmacia Biotech), which were blotted with monoclonal antibodies against MSH6 (0.02 μg ml−1, clone 44, BD Transduction Laboratories) and MSH2 (0.3 μg ml−1, Ab-1, Oncogene Research Products).
In vitro MMR assay
The functionality of the recombinant MutSα (MSH2/MSH6) variants was studied in the in vitro MMR assay (Kariola et al, 2002, 2003). In the assay, circular DNA heteroduplexes, which contain a G•T mispair 369 base pairs downstream from a single-strand nick were used as substrates. The functionality of the mutated MSH6 proteins was studied by complementing MMR-deficient nuclear extract (NE) of HCT15 (MSH6−/−) cells with the total protein extract (TE) of Sf9 cells including over expressed human MSH6 and MSH2 proteins. Each protein extract was estimated to contain equal quantities of recombinant wild-type MSH2. NE of the MMR-proficient cell line TK6, and HCT15 complemented with wild-type MutSα TE, were used as positive controls and HCT15 NE as a negative control in the assay. If the repair reaction occurs, it converts G•T heteroduplex to A•T homoduplex, whereafter restriction enzyme BglII can cleave the molecule and the repair efficiency can be measured by the cleavage efficiency. The quantification of the repaired DNA was carried out with Image-Pro Version 4.0 (MediaCybernetics®). Since the heteroduplex molecules are not all repairable they are always added in excess in the assay, and the repair capacity of the proficient wild-type controls is used as a reference level.
Results
Genetic and clinical data in MSH6 mutation carriers
Five novel MSH6 missense mutations (MSH6-R128L, MSH6-P623L, MSH6-K728T, MSH6-G881K+S, MSH6-E1193K) were found in six patients, whose endometrial or colorectal tumour showed high MSI (when at least two out of five markers showed instability). In each case, mutations detectable by DNA sequencing had been excluded in MLH1 and MSH2. Furthermore, the missense changes had not been found in 140 control individuals of comparable ethnicity. The ages at diagnosis were high, with a range of 58–83 years, and none of the mutation carriers had first-degree relatives with EC or CRC. The identified mutations and molecular and clinical data are shown in Table 1.
IHC staining for MLH1, MSH2, and MSH6 revealed that MSH6 protein was lost or nearly lost (5%) in three ECs, derived from the patient carrying the mutation P623L and the two women both carrying the mutation E1193K. One of the latter tumours also lacked MSH2 protein, while MLH1 was expressed. MLH1 protein was absent in three tumours (R128L, K728T, G881K+S), in which MSH2 and MSH6 were expressed. The lack of MLH1 in these tumours was consistent with the finding that the same tumours also showed hypermethylation of the MLH1 promoter region, which often leads to MLH1 gene inactivation and loss of MMR function.
Interactions of the MSH6 variants with wild-type MSH2
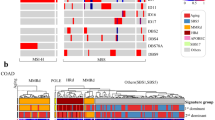
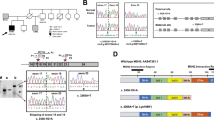
Combined co-immunoprecipitation and Western blot analysis was used to study the effect of MSH6 mutations on the heterodimerisation of MSH6 and MSH2 proteins. Wild-type MutSα (MutSα-WT) was used as a control. As shown in Figure 1A, the total protein extracts from Sf9 cells used for immunoprecipitation assay contained similar amounts of wild-type MSH2 (MutSα-WT, MutSα-R128L, MutSα-P623L, MutSα-K728T, MutSα-G881K+S, and MutSα-E1193K). The immunoprecipitates obtained with anti-MSH6 antibody show that co-expression of four of the MSH6 variants (MSH6-R128L, MSH6-P623L, MSH6-K728T, and MSH6-G881K+S) with wild-type MSH2 yielded stable heterodimers, since the mutated MSH6 co-precipitated similar amount of MSH2 as wild-type MSH6. In contrast, MSH6-E1193K co-precipitated much less MSH2 than MSH6-WT, suggesting interference in their heterodimerisation.
(A) Combined immunoprecipitation and Western blot analysis showing interactions of MSH6 variants with wild-type MSH2. The upper panel shows a Western blot including total protein extracts (TE), the lower panel shows immunoprecipitates (IP) obtained with anti-MSH6 antibody. Following transfer onto the membrane, the proteins were visualised with anti-MSH6 and anti-MSH2 antibodies (see Materials and Methods). The figure shows that the mutated MSH6 proteins R128L, P623L, K728T, and G881K+S are able to form stable MSH2-MSH6 heterodimers and co-precipitate a similar amount of the MSH2 protein as wild-type MSH6, while MSH6-E1193K co-precipitates clearly less MSH2 protein. (B) MMR activity of MMR deficient nuclear extract of HCT15 cells complemented with wild-type MutSα (MutSα-WT) and mutated variants. Mock contains heteroduplex DNA with no protein, TK6 is an MMR-proficient and HCT15 an MMR-deficient (MSH6−/−) control (see Materials and Methods). The fragment lengths on the left indicate the migration of the unrepaired linearised plasmid DNA (3193 bp) and of the two fragments (1833 and 1360 bp) produced following correction of the G•T mispair, which makes the DNA susceptible to cleavage with the restriction endonuclease BglII. The numbers below the panel represent fractions (%) of repaired DNA and display absolute MMR deficiency (0%) of MutSα-E1193K. The values are an average of two independent experiments.
Mismatch repair capacity of MutSα heterodimers
The functionality of the MutSα variants was tested in an in vitro MMR assay (Figure 1B). MutSα-WT, MutSα-R128L, MutSα-P623L, MutSα-K728T, and MutSα-G881K+S were able to complement the MSH6-deficient NE of HCT15 cells and repaired 39, 41, 39, 41, and 41% of G•T heteroduplexes, respectively. However, MutSα-E1193K protein complex failed to complement HCT15 NE and displayed complete MMR deficiency (0%).
Discussion
Genetic counselling of mutation carriers must be based on a reliable assessment of the pathogenicity of the mutation. The present study was undertaken to evaluate the pathogenicity of inherited missense alterations in the MMR gene MSH6. To date, MSH6 gene is the third most common susceptibility gene in the HNPCC syndrome accounting for about 10% of reported mutations found in HNPCC kindreds (http://www.nfdht.nl). Missense mutations, which account for 38% of all MSH6 mutations, are particularly challenging in the clinical management of patients with suspected HNPCC. Even when the amino-acid substitution is pathogenic, the mutation does not necessarily lead to the absence of the protein; thus, it can be detectable by IHC analysis. Furthermore, segregation studies to distinguish pathogenic mutations from polymorphisms are frequently hindered by small family size.
We have previously studied the functionality of six MSH6 variants (S144I, G566R, P1087T, P1087R, R1095H, and L1354Q), four of which are listed as pathogenic mutations in the mutation database. The mutations were found in HNPCC kindreds, which did not fulfil the Amsterdam criteria (I or II) and were classified as non-HNPCC or suspected HNPCC families (Vasen et al, 1991, 1999). The same functional assay was previously used to detect impaired MMR caused by the MLH1 missense mutations C77R, I107R, and R659P (Nyström-Lahti et al, 2002). Interestingly, all the six mutated MSH6 proteins were able to repair mismatches (Kariola et al, 2002, 2003), suggesting no MMR defect detectable in our assay. To further address the question of whether MSH6 missense mutations cause MMR defect and susceptibility to HNPCC, the pathogenicity of five novel MSH6 missense variations (R128L, P623L, K728T, G881K+S, and E1193K) was evaluated in the present study. These changes were derived from a series of consecutive EC and CRC patients selected for study after their tumours were determined to be microsatellite unstable. Importantly, in each case, the mutation was searched for in 140 control individuals and not found. Thus, these changes could not be classified a priori as likely polymorphisms. Irrespective of these facts, none of the mutation carriers was shown to have first-degree relatives with EC or CRC or any family history of cancer suggestive of HNPCC.
We studied the interactions of five mutated MSH6 proteins with normal MSH2 and tested the functionality of these heterodimers of MSH6 and MSH2 in the in vitro MMR assay. Four mutations (R128L, P623L, K728t, G881K+S) showed no impairment of these functions while the fifth mutation (E1193K) displayed marked impairment of both functions. The MSH6-E1193K protein co-precipitated much less MSH2 than wild-type MSH6, and the heterodimer MutSα-E1193K could not repair mismatches. The E1193K mutation occurred in two independently ascertained women with EC (ages at onset 59 and 60 years). Both tumours stained negatively or very weakly for MSH6 protein, which is compatible with MMR deficiency associated with MSH6 mutation. The amino acid E1193 is located in a highly conserved region between the ATP and Mg2+ binding sites in MSH6 (Iaccarino et al, 1998, 2000; Dufner et al, 2000) and, accordingly, can be expected to be sensitive to alterations. Irrespective of the definite MMR deficiency associated with the E1193K mutation, the families of the two patients, their siblings, excluding one brother diagnosed with skin cancer at 60, parents and children were unaffected. In immunohistochemical analysis, the three tumours derived from patients carrying mutations R128L, K728T, and G881K+S, expressed MSH6 but lacked MLH1 protein, which may explain the MSI phenotype in these tumours and suggests IHC staining for the MLH1 protein as the first approach in microsatellite unstable tumours. However, the mutation, P623L, which had no detectable phenotype in our functional assay was found in a patient, whose EC showed high MSI, and MSH6 was the only nonexpressed protein in the tumour among the three proteins tested. Thus, we cannot exclude the possibility that pathogenicity of P623L is caused by lower levels of the functional MSH6 protein as we have previously shown to be the case with some MLH1 mutations (Raevaara et al, 2003). Another possibility is that MSH6 inactivation is not at all due to P623L but has been acquired as a somatic event during the development of the tumour.
Our results, taken together with our previous similar findings concerning six other missense mutations in MSH6 and the clinical data of the patients and their families, lead us to conclude that most missense changes in MSH6 cause no or low cancer susceptibility. The results do not exclude the possibility that some of the studied MSH6 mutations, which were functional in the in vitro MMR assay, could still affect biochemical events preceding the MMR function in vivo. Yet, a functional MMR defect, especially in situations where cosegregation of missense mutation and disease phenotype cannot be studied, reliably confirms that cancer susceptibility in a family is linked to a found mutation.
Change history
16 November 2011
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Berends MJW, Wu Y, Sijmons RH, Mensink RGJ, van der Sluis T, Hordijk-Hos JM, de Vries EGE, Hollema H, Karrenbeld A, Buys CHCM, van der Zee AGJ, Hofstra RMW, Kleibeuker JH (2002) Molecular and clinical characteristics of MSH6 variants: an analysis of 25 index carriers of a germline variant. Am J Hum Genet 70: 26–37
Boland CR, Thibodeau SN, Hamilton SR, Sidransky D, Eshleman JR, Burt RW, Meltzer SJ, Rodriguez-Bigas MA, Fodde R, Ranzani GN, Srivastava S (1998) A National Cancer Institute Workshop on Microsatellite Instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 58: 5248–5257
Cederquist K, Emanuelsson M, Göransson I, Holinski-Feder E, Müller-Koch Y, Golovleva I, Grönberg H (2004) Mutation analysis of the MLH1, MSH2 and MSH6 genes in patients with double primary cancers of the colorectum and the endometrium: a population-based study in northern Sweden. Int J Cancer 109: 370–376
Chadwick RB, Pyatt RE, Niemann TH, Richards SK, Johnson CK, Stevens MW, Meek JE, Hampel H, Prior TW, de la Chapelle A (2001) Hereditary and somatic DNA mismatch repair gene mutations in sporadic endometrial carcinoma. J Med Genet 38: 461–466
Chang DK, Ricciardiello L, Goel A, Chang CL, Boland R (2000) Steady-state regulation of the human DNA mismatch repair system. J Biol Chem 275: 18424–18431
Deng G, Chen A, Hong J, Chae HS, Kim YS (1999) Methylation of CpG in a small region of the hMLH1 promoter invariably correlates with the absence of gene expression. Cancer Res 59: 2029–2033
de Wind N, Dekker M, Claij N, Jansen L, van Klink Y, Radman M, Riggins G, van der Valk M, van't Wout K, te Riele H (1999) HNPCC-like cancer predisposition in mice through simultaneous loss of Msh3 and Msh6 mismatch-repair protein functions. Nat Genet 23: 359–362
Dufner P, Marra G, Räschle M, Jiricny J (2000) Mismatch recognition and DNA-dependent stimulation of the ATPase activity of hMutSα is abolished by a single mutation in the hMSH6 subunit. J Biol Chem 275: 36550–36555
Herman JG, Umar A, Polyak K, Graff JR, Ahuja N, Issa JP, Markowitz S, Willson JK, Hamilton SR, Kinzler KW, Kane MR, Kolodner RD, Vogelstein B, Kunkel TA, Baylin SB (1998) Incidence and functional consequences of hMLH1 promoter hypermethylation in colorectal carcinoma. Proc Natl Acad Sci USA 95: 6870–6875
Iaccarino I, Marra G, Dufner P, Jiricny J (2000) Mutation in the magnesium binding site of hMSH6 disables the hMutSα sliding clamp from translocating along DNA. J Biol Chem 275: 2080–2086
Iaccarino I, Marra G, Palombo F, Jiricny J (1998) hMSH2 and hMSH6 play distinct roles in mismatch binding and contribute differently to the ATPase activity of hMutSα. EMBO J 17: 2677–2686
Kariola R, Otway R, Lönnqvist KE, Raevaara TE, Macrae F, Vos YJ, Kohonen-Corish M, Hofstra RMW, Nyström-Lahti M (2003) Two mismatch repair gene mutations found in a colon cancer patient – which one is pathogenic? Hum Genet 112: 105–109
Kariola R, Raevaara TE, Lönnqvist KE, Nyström-Lahti M (2002) Functional analysis of MSH6 mutations linked to kindreds with putative hereditary non-polyposis colorectal cancer syndrome. Hum Mol Genet 11: 1303–1310
Kolodner RD, Tytell JD, Schmeits JL, Kane MF, Das Gupta R, Weger J, Wahlberg S, Fox EA, Peel D, Ziogas A, Garber JE, Syngal S, Anton-Culver H, Li FP (1999) Germ-line msh6 mutations in colorectal cancer families. Cancer Res 59: 5068–5074
Marra G, Iaccarino I, Lettieri T, Roscilli G, Delmastro P, Jiricny J (1998) Mismatch repair deficiency associated with overexpression of the MSH3 gene. Proc Natl Acad Sci USA 95: 8568–8573
Miyaki M, Konishi M, Tanaka K, Kikuchi-Yanoshita R, Muraoka M, Yasuno M, Igari T, Koike M, Chiba M, Mori T (1997) Germline mutation of MSH6 as the cause of hereditary nonpolyposis colorectal cancer. Nat Genet 17: 271–272
Nakagawa H, Chadwick RB, Peltomäki P, Plass C, Nakamura Y, de la Chapelle A (2001) Loss of imprinting of the insulin-like growth factor II gene occurs by biallelic methylation in a core region of H19-associated CTCF-binding sites in colorectal cancer. Proc Natl Acad Sci USA 98: 591–596
Nyström-Lahti M, Perrera C, Räschle M, Panyushkina-Seiler E, Marra G, Curci A, Quaresima B, Costanzo F, D'Urso M, Venuta S, Jiricny J (2002) Functional analysis of MLH1 mutations linked to hereditary nonpolyposis colon cancer. Genes Chromosomes Cancer 33: 160–167
Parc YR, Halling KC, Wang L, Christensen ER, Cunningham JM, French AJ, Burgart LJ, Price-Troska TL, Roche PC, Thibodeau SN (2000) hMSH6 alterations in patients with microsatellite instability-low colorectal cancer. Cancer Res 60: 2225–2231
Peterlongo P, Nafa K, Lerman GS, Glogowski E, Shia J, Ye TZ, Markowitz AJ, Guillem JG, Kolachana P, Boyd JA, Offit K, Ellis NA (2003) MSH6 germline mutations are rare in colorectal cancer families. Int J Cancer 107: 571–579
Raevaara TE, Vaccaro C, Abdel-Rahman WM, Mocetti E, Bala S, Lönnqvist KE, Kariola R, Lynch HL, Peltomäki P, Nyström-Lahti M (2003) Pathogenicity of the hereditary colorectal cancer mutation hMLH1 del616 linked to shortage of the functional protein. Gastroenterology 125: 501–509
Vasen HFA, Mecklin J-P, Khan PM, Lynch HT (1991) The International Collaborative Group on Hereditary Non-Polyposis Colorectal Cancer (ICG-HNPCC). Dis Colon Rectum 34: 424–425
Vasen HFA, Watson P, Mecklin J-P, Lynch HT, the ICG-HNPCC (1999) New clinical criteria for hereditary nonpolyposis colorectal cancer (HNPCC, Lynch syndrome) proposed by the international collaborative group on HNPCC. Gastroenterology 116: 1453–1456
Wagner A, Hendriks Y, Meijers-Heijboer EJ, de Leeuw WJF, Morreau H, Hofstra R, Tops C, Bik E, Bröcker-Vriends AHJT, van der Meer C, Lindhout D, Vasen HFA, Breuning MH, Cornelisse CJ, van Krimpen C, Niermeijer MF, Zwinderman AH, Wijnen J, Fodde R (2001) Atypical HNPCC owing to MSH6 germline mutations: analysis of a large Dutch pedigree. J Med Genet 38: 318–322
Wijnen J, de Leeuw W, Vasen H, van der Klift H, Møller P, Stormorken A, Meijers-Heijboer H, Lindhout D, Menko F, Vossen S, Möslein G, Tops C, Bröcker-Vriends A, Wu Y, Hofstra R, Sijmons R, Cornelisse C, Morreau H, Fodde R (1999) Familial endometrial cancer in female carriers of MSH6 germline mutations. Nat Genet 23: 142–144
Wu Y, Berends MJV, Mensink RGJ, Kempinga C, Sijmons RH, van der Zee AGJ, Hollema H, Kleibeuker JH, Buys CHCM, Hofstra RMW (1999) Association of hereditary nonpolyposis colorectal cancer-related tumors displaying low microsatellite instability with MSH6 germline mutations. Am J Hum Genet 65: 1291–1298
Acknowledgements
This study was supported by grants from the Sigrid Juselius Foundation, the Finnish Cancer Foundation, the European Commission (QLG1-CT-2000-01230), and grants CA67941 and CA-16058 from the National Institutes of Health, USA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Kariola, R., Hampel, H., Frankel, W. et al. MSH6 missense mutations are often associated with no or low cancer susceptibility. Br J Cancer 91, 1287–1292 (2004). https://doi.org/10.1038/sj.bjc.6602129
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.bjc.6602129
Keywords
This article is cited by
-
An MSH6 germline pathogenic variant p.Gly162Ter associated with Lynch syndrome
Human Genome Variation (2022)
-
CoDP: predicting the impact of unclassified genetic variants in MSH6 by the combination of different properties of the protein
Journal of Biomedical Science (2013)
-
Sub-cellular localization analysis of MSH6 missense mutations does not reveal an overt MSH6 nuclear transport impairment
Familial Cancer (2012)
-
A putative Lynch syndrome family carrying MSH2 and MSH6 variants of uncertain significance—functional analysis reveals the pathogenic one
Familial Cancer (2011)
-
MutSβ exceeds MutSα in dinucleotide loop repair
British Journal of Cancer (2010)