Abstract
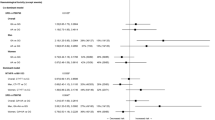
The primary end point of the study was the analysis of associations between polymorphisms with putative influence on 5-fluorouracil/irinotecan activity and progression-free survival (PFS) of patients with advanced colorectal cancer treated with first-line FOLFIRI chemotherapy. Peripheral blood samples from 146 prospectively enrolled patients were used for genotyping polymorphisms in thymidylate synthase (TS), methylenetetrahydrofolate reductase (MTHFR), excision repair cross-complementation group-1 (ERCC 1) xeroderma pigmentosum group-D (XPD), X-ray cross-complementing-1 (XRCC 1), X-ray cross-complementing-3 (XRCC 3) and uridine diphosphate-glucuronosyltransferases-A1 (UGT1 A1). TS 3′-UTR 6+/6+ and XRCC3-241 C/C genotypes were associated with adverse PFS. Hazard ratio for PFS achieved 2.89 (95% confidence interval=1.56–5.80; P=0.002) in 30 patients (20%) with both risk genotypes. Risk for Grade III–IV neutropenia was significantly associated with UGT1A1*28 7/7 genotype. These promising findings deserve further investigations and their validation in independent prospective studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Kelly H, Goldberg RM . Systemic therapy for metastatic colorectal cancer: current options, current evidence. J Clin Oncol 2005; 23: 4553–4560.
Colucci G, Gebbia V, Paoletti G, Giuliani F, Caruso M, Gebbia N et al. Phase III randomized trial of FOLFIRI versus FOLFOX4 in the treatment of advanced colorectal cancer: a multicenter study of the Gruppo Oncologico Dell'Italia Meridionale. J Clin Oncol 2005; 23: 4866–4875.
Marsh S, McLeod HL . Cancer pharmacogenetics. Br J Cancer 2004; 90: 8–11.
Marsh S, McLeod HL . Pharmacogenetics of irinotecan toxicity. Pharmacogenomics 2004; 5: 835–843.
Popat S, Matakidou A, Houlston RS . Thymidylate synthase expression and prognosis in colorectal cancer: a systematic review and meta-analysis. J Clin Oncol 2004; 22: 529–536.
Horie N, Aiba H, Oguro K, Hojo H, Takeishi K . Functional analysis and DNA polymorphism of the tandemly repeated sequences in the 5′-terminal regulatory region of the human gene for thymidylate synthase. Cell Struct Funct 1995; 20: 191–197.
Kawakami K, Watanabe G . Identification and functional analysis of single nucleotide polymorphism in the tandem repeat sequence of the thymidylate synthase gene. Cancer Res 2003; 63: 6004–6007.
Mandola MV, Stoehlmacher J, Muller-Weeks S, Cesarone G, Yu MC, Lenz HJ et al. A novel single nucleotide polymorphism within the 5′ tandem repeat polymorphism of the thymidylate synthase gene abolishes USF-1 binding and alters transcriptional activity. Cancer Res 2003; 63: 2898–2904.
Kawakami K, Omura K, Kanehira E, Watanabe Y . Polymorphic tandem repeats in the thymidylate synthase gene is associated with its protein expression in human gastrointestinal cancers. Anticancer Res 1999; 19: 3249–3252.
Morganti M, Ciantelli M, Giglioni B, Putignano AL, Nobili S, Papi L et al. Relationships between promoter polymorphisms in the thymidylate synthase gene and mRNA levels in colorectal cancers. Eur J Cancer 2005; 41: 2176–2183.
Kawakami K, Watanabe G . The association of thymidylate synthase mRNA expression with its three gene polymorphisms in colorectal cancer. Proc Am Assoc Cancer Res 2004; 45: 484 (abstr 2104).
Ulrich CM, Bigler J, Velicer CM, Greene EA, Farin FM, Potter JD . Searching expressed sequence Tag databases: discovery and confirmation of a common polymorphism in the thymidylate synthase gene. Cancer Epidemiol Biomarkers Prev 2000; 9: 1381–1385.
Mandola MV, Stoehlmacher J, Zhang W, Groshen S, Yu MC, Iqbal S et al. A 6 bp polymorphism in the thymidylate synthase gene causes message instability and is associated with decreased intratumoral TS mRNA levels. Pharmacogenetics 2004; 14: 319–327.
Sohn KJ, Croxford R, Yates Z, Lucock M, Kim YI . Effect of the methylenetetrahydrofolate reductase C677 T polymorphism on chemosensitivity of colon and breast cancer cells to 5-fluorouracil and methotrexate. J Natl Cancer Inst 2004; 96: 134–144.
Etienne MC, Ilc K, Formento JL, Laurent-Puig P, Formento P, Cheradame S et al. Thymidylate synthase and methylenetetrahydrofolate reductase gene polymorphisms: relationships with 5-fluorouracil sensitivity. Br J Cancer 2004; 90: 526–534.
Kawakami K, Ruszkiewicz A, Bennett G, Moore J, Watanabe G, Iacopetta B . The folate pool in colorectal cancers is associated with DNA hypermethylation and with a polymorphism in methylenetetrahydrofolate reductase. Clin Cancer Res 2003; 9: 5860–5865.
Paz MF, Avila S, Fraga MF, Pollan M, Capella G, Peinado MA et al. Germ-line variants in methyl-group metabolism genes and susceptibility to DNA methylation in normal tissues and human primary tumors. Cancer Res 2002; 62: 4519–4524.
Iacopetta B . Methyl-group metabolism and the response of colorectal cancer to 5-Fluorouracil. Crit Rev Oncogenesis 2006; 12: 1–12.
Pommier Y . Topoisomerase I inhibitors: camptothecins and beyond. Nat Rev Cancer 2006; 6: 789–802.
Ando Y, Saka H, Ando M, Sawa T, Muro K, Ueoka H et al. Polymorphisms of UDP-glucuronosyltransferase gene and irinotecan toxicity: a pharmacogenetic analysis. Cancer Res 2000; 60: 6921–6926.
Rouits E, Boisdron-Celle M, Dumont A, Guerin O, Morel A, Gamelin E . Relevance of different UGT1A1 toxicity: a molecular and clinical study of 75 patients. Clin Cancer Res 2004; 10: 5151–5159.
Marcuello E, Altes A, Menoyo A, Del Rio E, Gomez-Pardo M, Baiget M . UGT1A1 gene variations and irinotecan treatment in patients with metastatic colorectal cancer. Br J Cancer 2004; 91: 678–682.
Toffoli G, Cecchin E, Corona G, Russo A, Buonadonna A, D'Andrea M et al. The role of UGT1A1*28 polymorphism in the pharmacodynamics and pharmacokinetics of irinotecan in patients with metastatic colorectal cancer. J Clin Oncol 2006; 24: 3061–3068.
Kawato Y, Aonuma M, Hirota Y, Kuga H, Sato K . Intracellular roles of SN-38, a metabolite of the camptothecin derivative CPT-11, in the antitumor effect of CPT-11. Cancer Res 1991; 51: 4187–4191.
Wu J, Yin MB, Hapke G, Toth K, Rustum YM . Induction of biphasic DNA double strand breaks and activation of multiple repair protein complexes by DNA topoisomerase I drug 7-ethyl-10-hydroxy-camptothecin. Mol Pharmacol 2002; 61: 742–748.
Vallbohmer D, Iqbal S, Yang DY, Rhodes KE, Zhang W, Gordon M et al. Molecular determinants of irinotecan efficacy. Int J Cancer 2006; 119: 2435–2442.
de Boer JG . Polymorphisms in DNA repair and environmental interactions. Mutat Res 2002; 509: 201–210.
Louvet C, de Gramont A, Tournigand C, Artru P, Maindrault-Goebel F, Krulik M . Correlation between progression free survival and response rate in patients with metastatic colorectal carcinoma. Cancer 2001; 91: 2033–2038.
Therasse P, Arbuck SG, Eisenhauer EA, Wanders J, Kaplan RS, Rubinstein L et al. New guidelines to evaluate the response to treatment in solid tumors. European Organization for Research and Treatment of Cancer, National Cancer Institute of the United States, National Cancer Institute of Canada. J Natl Cancer Inst 2000; 92: 205–216.
Pullarkat ST, Stoehlmacher J, Ghaderi V, Xiong YP, Ingles SA, Sherrod A et al. Thymidylate synthase gene polymorphism determines response and toxicity of 5-FU chemotherapy. Pharmacogenom J 2001; 1: 65–70.
Stoehlmacher J, Park DJ, Zhang W, Yang D, Groshen S, Zahedy S et al. A multivariate analysis of genomic polymorphisms: prediction of clinical outcome to 5-FU/oxaliplatin combination chemotherapy in refractory colorectal cancer. Br J Cancer 2004; 91: 344–354.
Dotor E, Cuatrecases M, Martinez-Iniesta M, Navarro M, Vilardell F, Guino E et al. Tumor thymidylate synthase 1494del6 genotype as a prognostic factor in colorectal cancer patients receiving fluorouracil-based adjuvant treatment. J Clin Oncol 2006; 24: 1603–1611.
Jakobsen A, Nielsen JN, Gyldenkerne N, Lindeberg J . Thymidylate synthase and methylenetetrahydrofolate reductase gene polymorphism in normal tissue as predictors of fluorouracil sensitivity. J Clin Oncol 2005; 23: 1365–1369.
Marcuello E, Altes A, del Rio E, Cesar A, Menoyo A, Baiget M . Single nucleotide polymorphism in the 5′ tandem repeat sequences of thymidylate synthase gene predicts for response to fluorouracil-based chemotherapy in advanced colorectal cancer patients. Int J Cancer 2004; 112: 733–737.
Cohen V, Panet-Raymond V, Sabbaghian N, Morin I, Batist G, Rozen R . Methylenetetrahydrofolate reductase polymorphism in advanced colorectal cancer: a novel genomic predictor of clinical response to fluoropyrimidine-based chemotherapy. Clin Cancer Res 2003; 9: 1611–1615.
Etienne MC, Formento JL, Chazal M, Francoual M, Magne N, Formento P et al. Methylenetetrahydrofolate reductase gene polymorphisms and response to fluorouracil-based treatment in advanced colorectal cancer patients. Pharmacogenetics 2004; 14: 785–792.
Marcuello E, Altes A, Menoyo A, Baiget M . Methylenetetrahydrofolate reductase gene polymorphisms: genomic predictors of clinical response to fluoropyrimidine-based chemotherapy? Cancer Chemother Pharmacol 2006; 57: 835–840.
Kealey C, Brown KS, Woodside JV, Young I, Murray L, Boreham CA et al. A common insertion/deletion polymorphism of the thymidylate synthase (TYMS) gene is a determinant of red blood cell folate and homocysteine concentrations. Hum Genet 2005; 1160: 347–353.
Pullmann Jr R, Abdelmohsen K, Lal A, Martindale JL, Ladner RD, Gorospe M . Differential stability of thymidylate synthase 3′-untranslated region polymorphic variants regulated by AUF1. J Biol Chem 2006; 281: 23456–23463.
Innocenti F, Undevia SD, Iyer L, Chen PX, Das S, Kocherginsky M et al. Genetic variants in the UDP-glucuronosyltransferase 1A1 gene predict the risk of severe neutropenia of irinotecan. J Clin Oncol 2004; 22: 1382–1388.
McLeod HL, Parodi L, Sargent DJ, Marsh S, Green E, Abreu P et al. UGT1A1*28, toxicity and outcome in advanced colorectal cancer: results from Trial N9741. Proc Am Soc Clin Oncol 2006; 24: 151 (abstr. 3520).
Schulz C, Schalhorn A, Schwabe W, Haeusler P, Zwingers T, Heinemann V . Impact of gene promoter polymorphism of the UGT1A1-gene on the occurrance of irinotecan-induced side effects and drug effiacy. Proc Am Soc Clin Oncol 2005; 23: 263 (abstr. 3570).
Seymour MT, Braun MS, Richman SD, Daly C, Thompson LC, Meade A et al. Association of molecular markers with toxicity outcomes in a randomized trial of chemotherapy for advanced colorectal cancer (FOCUS Trial Investigators). Proc Am Soc Clin Oncol 2006; 24: 84 (abstr. 2022).
Leonard GD, Fojo T, Bates SE . The role of ABC transporters in clinical practice. Oncologist 2003; 8: 411–424.
Choudhuri S, Klaassen CD . Structure, function, expression, genomic organization, and single nucleotide polymorphisms of human ABCB1 (MDR1), ABCC (MRP), and ABCG2 (BCRP) efflux transporters. Int J Toxicol 2006; 25: 231–259.
Carlini LE, Meropol NJ, Bever J, Andria ML, Hill T, Gold P et al. UGT1A7 and UGT1A9 polymorphisms predict response and toxicity in colorectal cancer patients treated with capecitabine/irinotecan. Clin Cancer Res 2005; 11: 1226–1236.
Nagar S, Blanchard RL . Pharmacogenetics of uridine diphosphoglucuronosyltransferase (UGT) 1A family members and its role in patient response to irinotecan. Drug Metab Rev 2006; 38: 393–409.
Savas S, Kim DY, Ahmad MF, Shariff M, Ozcelik H . Identifying functional genetic variants in DNA repair pathway using protein conservation analysis. Cancer Epidemiol Biomarkers Prev 2004; 13: 801–807.
Kiuru A, Lindholm C, Heilimo I, Ceppi M, Koivistoinen A, Ilus T et al. Influence of DNA repair gene polymorphisms on the yield of chromosomal aberrations. Environ Mol Mutagen 2005; 46: 198–205.
Brenneman MA, Weiss AE, Nickoloff JA, Chen DJ . XRCC3 is required for efficient repair of chromosome breaks by homologous recombination. Mutat Res 2000; 459: 89–97.
Araujo FD, Pierce AJ, Stark JM, Jasim N . Variant XRCC3 implicated in cancer is functional in homology-directed repair of double-strand breaks. Oncogene 2002; 21: 4176–4180.
Arnaudeau C, Lundin C, Helleday T . DNA double-strand breaks associated with replication forks are predominantly repaired by homologous recombination involving an exchange mechanism in mammalian cells. J Mol Biol 2001; 307: 1235–1245.
Savas S, Kim DY, Ahmad MF, Shariff M, Ozcelik H . Identifying functional genetic variants in DNA repair pathway using protein conservation analysis. Cancer Epidemiol Biomarkers Prev 2004; 13: 801–807.
Kweekel DM, Gelderblom H, Guchelaar HJ . Pharmacology of oxaliplatin and the use of pharmacogenomics to individualize therapy. Cancer Treat Rev 2005; 31: 90–105.
Acknowledgements
We thank the Consorzio Interuniversitario per le Biotecnologie (CIB) and Fanoateneo for their financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ruzzo, A., Graziano, F., Loupakis, F. et al. Pharmacogenetic profiling in patients with advanced colorectal cancer treated with first-line FOLFIRI chemotherapy. Pharmacogenomics J 8, 278–288 (2008). https://doi.org/10.1038/sj.tpj.6500463
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.tpj.6500463
Keywords
This article is cited by
-
FOLFIRI-Mediated Toxicity in Human Aortic Smooth Muscle Cells and Possible Amelioration with Curcumin and Quercetin
Cardiovascular Toxicology (2020)
-
Polymorphisms of MTHFR and TYMS predict capecitabine‐induced hand‐foot syndrome in patients with metastatic breast cancer
Cancer Communications (2019)
-
Virtual Clinical Studies to Examine the Probability Distribution of the AUC at Target Tissues Using Physiologically-Based Pharmacokinetic Modeling: Application to Analyses of the Effect of Genetic Polymorphism of Enzymes and Transporters on Irinotecan Induced Side Effects
Pharmaceutical Research (2017)
-
Polymorphisms of MTHFR C677T and A1298C associated with survival in patients with colorectal cancer treated with 5-fluorouracil-based chemotherapy
International Journal of Clinical Oncology (2017)
-
Evaluation of 5-fluorouracil degradation rate and Pharmacogenetic profiling to predict toxicity following adjuvant Capecitabine
European Journal of Clinical Pharmacology (2017)