Abstract
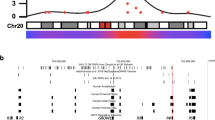
Identification of genes controlling complex diseases has proven to be difficult; however, animal models may pave the way to determine how low penetrant genes interact to promote disease development. We have dissected the Cia5/Eae3 susceptibility locus on mouse chromosome 3 previously identified to control disease in experimental models of multiple sclerosis and rheumatoid arthritis. Congenic strains showed significant but small effects on severity of both diseases. To improve the penetrance, we have now used a new strategy that defines the genetic interactions. The QTL interacted with another locus on chromosome 15 and a partial advanced intercross breeding of the two congenic strains for eight generations accumulated enough statistical power to identify interactions with several loci on chromosome 15. Thereby, three separate loci within the original QTL could be identified; Cia5 affected the onset of arthritis by an additive interaction with Cia31 on chromosome 15, whereas the Cia21 and Cia22 affected severity during the chronic phase of the disease through an epistatic interaction with Cia32 on chromosome 15. The definition of genetic interactions was a prerequisite to dissect the Cia5 QTL and we suggest the partial advanced intercross strategy to be helpful also for dissecting other QTL controlling complex phenotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Nadeau JH . Modifier genes in mice and humans. Nat Rev Genet 2001; 2: 165–174.
Nguyen C, Limaye N, Wakeland EK . Susceptibility genes in the pathogenesis of murine lupus. Arthritis Res 2002; 4 (Suppl 3): S255–S263.
Morel L, Blenman KR, Croker BP, Wakeland EK . The major murine systemic lupus erythematosus susceptibility locus, Sle1, is a cluster of functionally related genes. Proc Natl Acad Sci USA 2001; 98: 1787–1792.
Podolin P . Localization of two insulin-dependent diabetes (Idd) genes to the Idd10 region on mouse chromosome 3. Mamm Genome 1998 1998; 9: 283–286.
Serreze DV, Bridgett M, Chapman HD et al. Subcongenic analysis of the Idd13 locus in NOD/Lt mice: evidence for several susceptibility genes including a possible diabetogenic role for beta 2-microglobulin. J Immunol 1998; 160: 1472–1478.
Vingsbo-Lundberg C, Nordquist N, Olofsson P et al. Genetic control of arthritis onset, severity and chronicity in a model for rheumatoid arthritis in rats. Nat Genet 1998; 20: 401–404.
Morel L, Croker BP, Blenman KR et al. Genetic reconstitution of systemic lupus erythematosus immunopathology with polycongenic murine strains. Proc Natl Acad Sci USA 2000; 97: 6670–6675.
Lyons PA, Armitage N, Lord CJ et al. Mapping by genetic interaction: high-resolution congenic mapping of the type 1 diabetes loci Idd10 and Idd18 in the NOD mouse. Diabetes 2001; 50: 2633–2637.
Cheverud JM, Routman EJ . Epistasis and its contribution to genetic variance components. Genetics 1995; 139: 1455–1461.
Falconer DS, Mackay TFC . Introduction to Quantitative Genetics 1996.
Holmdahl R . Genetics of susceptibility to chronic experimental encephalomyelitis and arthritis. Curr Opin Immunol 1998; 10: 710–717.
Lindqvist AK, Bockermann R, Johansson AC et al. Mouse models for rheumatoid arthritis. Trends Genet 2002; 18: S7–S13.
Sundvall M, Jirholt J, Yang HT et al. Identification of murine loci associated with susceptibility to chronic experimental autoimmune encephalomyelitis. Nat Genet 1995; 10: 313–317.
Jirholt J, Cook A, Emahazion T et al. Genetic linkage analysis of collagen-induced arthritis in the mouse. Eur J Immunol 1998; 28: 3321–3328.
Svensson L, Nandakumar KS, Johansson A, Jansson L, Holmdahl R . IL-4-deficient mice develop less acute but more chronic relapsing collagen-induced arthritis. Eur J Immunol 2002; 32: 2944–2953.
Nandakumar KS, Svensson L, Holmdahl R . Collagen type II-specific monoclonal antibody-induced arthritis in mice: description of the disease and the influence of age, sex, and genes. Am J Pathol 2003; 163: 1827–1837.
Karlsson J, Johannesson M, Lindvall T et al. Genetic interactions in Eae2 control collagen induced arthritis and the CD4+/CD8+ T cell ratio. J Immunol 2005; 174: 533–541.
Johansson AC, Sundler M, Kjellen P et al. Genetic control of collagen-induced arthritis in a cross with NOD and C57BL/10 mice is dependent on gene regions encoding complement factor 5 and FcgammaRIIb and is not associated with loci controlling diabetes. Eur J Immunol 2001; 31: 1847–1856.
Glazier AM, Nadeau JH, Aitman TJ . Finding genes that underlie complex traits. Science 2002; 298: 2345–2349.
Yi N, Xu S, Allison DB . Bayesian model choice and search strategies for mapping interacting quantitative trait loci. Genetics 2003; 165: 867–883.
Peripato AC, De Brito RA, Matioli SR et al. Epistasis affecting litter size in mice. J Evol Biol 2004; 17: 593–602.
R development core team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, 2004.
Mahler M, Leiter EH . Genetic and environmental context determines the course of colitis developing in IL-10-deficient mice. Inflamm Bowel Dis 2002; 8: 347–355.
Podolin PL, Denny P, Lord CJ et al. Congenic mapping of the insulin-dependent diabetes (Idd) gene, Idd10, localizes two genes mediating the Idd10 effect and eliminates the candidate Fcgr1. J Immunol 1997; 159: 1835–1843.
Encinas JA, Wicker LS, Peterson LB et al. QTL influencing autoimmune diabetes and encephalomyelitis map to a 0.15-cM region containing Il2 [letter]. Nat Genet 1999; 21: 158–160.
Dong P, Hood L, McIndoe RA . Detection of a large RIII-derived chromosomal segment on chromosome 10 in the H-2 congenic strain B10.RIII(71NS)/Sn. Genomics 1996; 31: 266–269.
Korstanje R, Paigen B . From QTL to gene: the harvest begins. Nat Genet 2002; 31: 235–236.
Markel P, Shu P, Ebeling C et al. Theoretical and empirical issues for marker-assisted breeding of congenic mouse strains. Nat Genet 1997; 17: 280–284.
Jirholt J, Lindqvist AK, Karlsson J, Andersson A, Holmdahl R . Identification of susceptibility genes for experimental autoimmune encephalomyelitis that overcome the effect of protective alleles at the eae2 locus. Int Immunol 2002; 14: 79–85.
Broman KW, Wu H, Sen S, Churchill GA . R/qtl: QTL mapping in experimental crosses. Bioinformatics 2003; 19: 889–890.
Acknowledgements
We thank Isabell Bohlin and Carlos Palestro for help with the animal care. This work was supported by grants from the Swedish Medical Research Council, the Swedish Foundation for Strategic Research, the Swedish Association against Rheumatism, the Crafoord, Lundberg, the Kock and Österlund Foundations and EU FP5 (QLG1-CT-2001-01407 ‘EUROME’). Dr Patrik Wernhoff was supported by the EU Grant HPMD-2000-00047.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Johannesson, M., Karlsson, J., Wernhoff, P. et al. Identification of epistasis through a partial advanced intercross reveals three arthritis loci within the Cia5 QTL in mice. Genes Immun 6, 175–185 (2005). https://doi.org/10.1038/sj.gene.6364155
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364155
Keywords
This article is cited by
-
Polymorphic estrogen receptor binding site causes Cd2-dependent sex bias in the susceptibility to autoimmune diseases
Nature Communications (2021)
-
Fine-mapping QTLs in advanced intercross lines and other outbred populations
Mammalian Genome (2014)
-
Disease-promoting and -protective genomic loci on mouse chromosomes 3 and 19 control the incidence and severity of autoimmune arthritis
Genes & Immunity (2012)
-
Dissection of a locus on mouse chromosome 5 reveals arthritis promoting and inhibitory genes
Arthritis Research & Therapy (2009)
-
Contribution of genetic studies in rodent models of autoimmune arthritis to understanding and treatment of rheumatoid arthritis
Genes & Immunity (2007)