Abstract
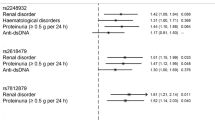
Renal disease occurs in 40–75% of systemic lupus erythematosus (SLE) patients and significantly contributes to morbidity and mortality. We used two pedigree stratification strategies to explore the impact of the ACR renal criterion for SLE classification upon genetic linkage with SLE. In both we used SLE as the phenotype. First, we evaluated genome scan data from >300 microsatellite markers in the 75 pedigrees that had at least one SLE affected with the SLE renal criterion. A maximum-likehood parametric model approach produced a maximum screening LOD score of 3.16 at 10q22.3 in the European-American (EA) pedigrees. The African-American (AA) pedigrees obtained a maximum screening LOD score of 2.58 at 11p15.6. A multipoint sib-pair regression analysis produced P = 0.0000008 in EA at 10q22.3 (SLEN1) and P = 0.000001 in AA at 2q34-35 (SLEN2). A second stratification strategy explored the renal criterion in 35 pedigrees with two or more SLE patients with renal disease and produced a LOD score of 3.34 at 11p15.6 in AA (SLEN3). Sib-pair analysis in these 35 pedigrees revealed P = 0.00003 at 4q13.1 in EA, P = 0.00003 at 11p13 and 0.00007 at 3q23 in AA. Thus, multiple genetic linkages are related to the renal criterion in SLE. Of the significant genetic linkages with SLE described herein, those at 10q22.3 in the EA pedigrees (SLEN1) and at 2q34-35 in the AA pedigrees (SLEN2) have not been previously described.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
García CO, Molina JF, Gutiérrez-Ureña S et al. Autoantibody profile in African-American patients with lupus nephritis Lupus 1996 5: 602–605
Hirsch R, White P . Systemic lupus in children Curr Pediatr 1991 1: 85–88
Walker WG, Solez K . Renal involvement in disorders of connective tissues In: Early LE, Gottschalk (eds) Diseases of Kidneys Little Brown: Boston 1979 2: pp 1259–1288
Bakir AA, Levy PS, Dunea G . The prognosis of lupus nephritis in African-Americans: a retrospective analysis Am J Kidney Dis 1994 24: 159–171
Clark WF . Treatment of lupus nephritis: immunosuppression, general therapy, dialysis and transplantation Clin Invest Med 1994 17: 588–601
Reichlin M, Harley JB . Detection by ELISA of antibodies to small RNA protein particles in systemic lupus erythematosus patients whose sera lack precipitins Trans Assoc Am Physicians 1986 99: 161–171
Molina JF, Molina J, Garcia C, Gharavi AE, Wilson WA, Espinoza LR . Ethnic differences in the clinical expression of systemic lupus erythematosus Lupus 1997 6: 63–67
Arnett FC, Hamilton RG, Roebber M, Harley JB, Reichlin M . Increased frequencies of Sm and nRNP autoantibodies in American blacks compared to whites with systemic lupus erythematosus J Rheumatol 1988 15: 1773–1776
Suzuki N, Harada T, Mizushima Y, Sakane T . Possible pathogenic role of cationic anti-DNA autoantibodies in the development of nephritis in patients with systemic lupus erythematosus J Immunol 1993 151: 1128–1136
Asero R, Banfi G, Radelli L et al. Relationship between antibodies to dsDNA and to soluble cellular antigens and histologically defined glomerulonephritis in patients with SLE Autoimmunity 1990 7: 13–21
Tan EM, Schur PH, Carr RI, Kunkel HG . Deoxyribonucleic acid (DNA) and antibodies to DNA in the serum of patients with systemic lupus erythematosus J Clin Invest 1966 45: 1732–1740
Stetler DA, Cavallo T . Anti-RNA polymerase I antibodies: potential role in the induction and progression of murine lupus nephritis J Immunol 1987 138: 2119–2123
Cameron JS . Lupus nephritis J Am Soc Nephrol 1999 10: 413–424
Lander E, Kruglyak L . Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results Nat Genet 1995 11: 241–247
Nath SK, Kelly JA, Namjou B et al. Evidence for a susceptibility gene (SLEV1) on chromosome 17p13 for vitiligo related systemic lupus erythematosus families Am J Hum Genet 2001 69: 1401–1406
Namjou B, Nath SK, Kilpatrick J et al. Genome scan stratified by the presence of anti-double stranded DNA (dsDNA) autoantibody in pedigrees multiplex for systemic lupus erythematosus (SLE) establishes linkages at 19p13.2 (SLED1) and 18q21.1 (SLED2) Genes Immun 2002 3
Gaffney PM, Ortmann WA, Selby SA et al. Genome screening in human systemic lupus erythematosus: results from a second Minnesota cohort and combined analyses of 187 sib-pair families Am J Hum Genet 2000 66: 547–556
Lindqvist AK, Steinsson K, Johanneson B et al. A susceptibility locus for human systemic lupus erythematosus (hSLE1) on chromosome 2q J Autoimmun 2000 14: 169–178
Bleesing JJ, Straus SE, Fleisher TA . Autoimmune lymhoproliferative syndrome. A human disorder of abnormal lymphocyte survival Pediatr Clin North Am 2000 47: 1291–1310
Rieux-Laucat F, BlachŁre S, Danielan S et al. Lymphoproliferative syndrome with autoimmunity: a possible genetic basis for dominant expression of the clinical manifestations Blood 1999 94: 2575–2582
Siaiki O, Tanaka T, Kishimoto S . Defective expression of p70/75 interleukin 2 receptor in T cells in patients with systemic lupus erythematosus: a possible defect in the process of increased intracellular calcium leading to p70/75 expression J Rheumatol 1990 17: 1303–1307
Hochberg MC . Updating the American College of Rheumatology Criteria for systemic lupus erythematosus Arthritis Rheum 1998 41: 751
Gray-McGuire C, Moser KL, Gaffney P et al. Genome scan of human systemic lupus erythematosus by regression modeling: evidence of linkage and epistasis at 4p16-15.2 Am J Hum Genet 2000 67: 1460–1469
Olson JM . Relationship estimation by Markov-process models in a sib-pair linkage study Am J Hum Genet 1999 64: 1464–1472
S.A.G.E. Statistical Analysis for genetic epidemiology, release 4.0, Beta 3 Computer package, Department of Epidemiology and Biostatistics, Case Western Reserve University, Cleveland, OH 1999
Cottingham RW Jr, Idury RM, Schaffer AA . Fast sequential genetic linkage computation Am J Hum Genet 1993 53: 252–263
Schaffer AA, Gupta SK, Shriram K, Cottingham RW Jr . Avoiding recomputation in linkage analysis Hum Hered 1994 44: 225–237
Terwilliger JD . A powerful likehood method for the analysis of linkage disequilibrium between trait loci and one more polymorphic marker loci Am J Hum Genet 1995 56: 777–787
Moser KL, Neas BR, Salmon JE et al. Genome scan of human systemic lupus erythematosus: evidence for linkage on chromosome 1q in African-American pedigrees Proc Natl Acad Sci USA 1998 95: 14869–14874
Elston RC, Buxbaum S, Jacobs KB, Olson JM . Haseman and Elston revisited Gen Epi 2000 19: 1–17
Acknowledgements
Special thanks are given to all the families that participated in the study. The study was supported by the National Institutes of Health (AI42460, AR24717, AR45231, AI31584 and AR52221), the Lupus Multiplex Registry and Repository (AR-1-2253) and the US Department of Veterans Affairs.
Author information
Authors and Affiliations
Corresponding author
Additional information
Grant Support: This work was supported by the National Institutes of Health (AI42460, AR24717, AR45231, AI31584 and AR52221) and the US Department of Veterans Affairs.
Rights and permissions
About this article
Cite this article
Quintero-Del-Rio, A., Kelly, J., Kilpatrick, J. et al. The genetics of systemic lupus erythematosus stratified by renal disease: linkage at 10q22.3 (SLEN1), 2q34-35 (SLEN2), and 11p15.6 (SLEN3). Genes Immun 3 (Suppl 1), S57–S62 (2002). https://doi.org/10.1038/sj.gene.6363901
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6363901
Keywords
This article is cited by
-
Prevalence of genetic renal disease in children
Pediatric Nephrology (2013)
-
A novel pathogenetic concept—antiviral immunity in lupus nephritis
Nature Reviews Nephrology (2012)
-
Glomerular diseases: genetic causes and future therapeutics
Nature Reviews Nephrology (2010)
-
Three checkpoints in lupus development: central tolerance in adaptive immunity, peripheral amplification by innate immunity and end-organ inflammation
Genes & Immunity (2009)
-
An investigation of genome-wide associations of hypertension with microsatellite markers in the family blood pressure program (FBPP)
Human Genetics (2007)