Abstract
Metabolic response to the triglyceride (TG)-lowering drug, fenofibrate, is shaped by interactions between genetic and environmental factors, yet knowledge regarding the genetic determinants of this response is primarily limited to single-gene effects. Since very low-density lipoprotein (VLDL) is the central carrier of fasting TG, identifying factors that affect both total TG and VLDL–TG response to fenofibrate is critical for predicting individual fenofibrate response. As part of the Genetics of Lipid Lowering Drugs and Diet Network (GOLDN) study, 688 individuals from 161 families were genotyped for 91 single-nucleotide polymorphisms (SNPs) in 25 genes known to be involved in lipoprotein metabolism. Using generalized estimating equations to control for family structure, we performed linear modeling to investigate whether single SNPs, single covariates, SNP–SNP interactions, and/or SNP–covariate interactions had a significant association with the change in total fasting TG and fasting VLDL–TG after 3 weeks of fenofibrate treatment. A 10-iteration fourfold cross-validation procedure was used to validate significant associations and quantify their predictive abilities. More than one-third of the significant, cross-validated SNP–SNP interactions predicting each outcome involved just five SNPs, showing that these SNPs are of key importance to fenofibrate response. Multiple variable models constructed using the top-ranked SNP--covariate interactions explained 11.9% more variation in the change in TG and 7.8% more variation in the change in VLDL than baseline TG alone. These results yield insight into the complex biology of fenofibrate response, which can be used to target fenofibrate therapy to individuals who are most likely to benefit from the drug.
Similar content being viewed by others
Introduction
Elevated serum triglyceride (TG) level is an underlying risk factor for coronary heart disease (CHD),1 even after controlling for other known lipoprotein risk factors such as low-density lipoprotein cholesterol (LDL-C) and high-density lipoprotein cholesterol (HDL-C).2, 3 It contributes to CHD directly by triggering vascular endothelial cell dysfunction and inflammation due to increased production of atherogenic TG-rich lipoproteins4 and indirectly through its association with type II diabetes and abdominal obesity, both major risk factors for CHD.5
Reducing high serum TG in individuals through diet or drug therapies has been shown to reduce adverse outcomes (ie, death or myocardial infarction) due to CHD.2 Current TG-lowering therapy includes fibrates, niacin, and omega-3 fatty acids (fish oil).6 Statins, which are often prescribed to lower LDL-C, may also have a mild TG-lowering effect in some patient populations.7 Fibrates (which include fenofibrate, gemfibrozil, and bezafibrate among others) are the primary treatment for high TG and have a very low frequency of adverse events,8 but there is significant interindividual variation in response to these drugs.9 Fibrates bind to transcription factors called PPARs, which then migrate to the nucleus and affect transcription of target genes that are involved in the lipid metabolic pathway.8
Fenofibrate reduces fasting TG level and induces a shift to a less atherogenic TG-rich lipoprotein profile.10 The effects of fenofibrate on the levels of total plasma TG and circulating very low-density lipoprotein (VLDL) particles, the primary carrier of TG during fasting,11 are mediated through its binding to PPAR-α.8 The resulting up- and downregulation of a number of genes involved in lipoprotein metabolism lowers TG by several mechanisms: (1) reducing the availability of free fatty acids for TG formation, (2) increasing the catabolism of VLDL, and (3) reducing neutral lipid (cholesterol ester and TG) exchange between VLDL and HDL.10, 12 Genes that are upregulated in response to PPAR-α include LPL, which is involved in TG lipolysis,8 APOA5, which enhances LPL action and may restrict VLDL secretion from the liver,8, 13 and several genes related to HDL-C metabolism including APOA1, APOA2, and FABP. PPAR-α also downregulates APOC3, an apolipoprotein that inhibits VLDL clearance.8
Although the mechanism of fenofibrate's action is well characterized, significant interindividual variability in response to fenofibrate9 limits the ability to predict individual response to the drug. This variation in fenofibrate response, like other complex phenotypes, is likely due to a combination of genetic and biological risk factors14 and interactions among these factors.15 In this study, the largest to date, we begin to characterize the genetic architecture of fasting TG response to fenofibrate drug therapy through the examination of independent genetic effects, interactions between genetic and demographic or biochemical factors, and interactions between pairs of genetic factors that are associated with changes in total plasma TG (mg/dl) and TG content of its largest constituent, VLDL particles, in response to fenofibrate.
We identified 91 single-nucleotide polymorphisms (SNPs) in 25 genes that have been previously identified as playing a role in lipoprotein metabolism.16, 17 The genes and polymorphisms tested are shown in Table 1, and a more in-depth description of the functional role of the genes is found in Supplementary Table 1. The polymorphisms chosen for this analysis have been previously shown to have a functional effect on lipid metabolism either alone or through interaction with dietary factors.18, 19, 20, 21, 22, 23 Variations in these genes have also been shown to affect response to pharmacological intervention with fibrates and other drugs.24, 25, 26
We examine the effects of these variants on fenofibrate response using a comprehensive, systematic approach that evaluates the role of many genes, many demographic and clinical covariates, and their interactions. Identifying the constellation of genetic and demographic and/or biochemical factors that interact to affect response to fenofibrate will provide insight into the complex mechanism of action of fenofibrate and may improve physicians' ability to identify individuals likely to respond to fenofibrate treatment.
Materials and methods
Sample
The sample for this study consisted of 688 Caucasian participants from 161 families that participated in the Genetics of Lipid Lowering Drugs and Diet Network (GOLDN) study, which was approved by the Institutional Review Boards at the University of Alabama, the University of Minnesota, the University of Utah, the University of Michigan, and Tufts University. Participants were recruited from two NHLBI Family Heart Study field centers, Minneapolis, MN and Salt Lake City, UT, as described by Lai et al.27 In each case, only families with at least two siblings were recruited, and only participants who did not take lipid-lowering agents (pharmaceuticals or nutraceuticals) for at least 4 weeks prior to the initial visit were included.
Protocol
Participants took part in two clinical visits, at least 3 weeks apart. During both visits, blood was drawn after a 12 h fast and analyzed for TG level, lipoprotein levels, total cholesterol, insulin, and glucose. Age, sex, blood pressure, weight, height, waist-to-hip ratio, and smoking status were ascertained at the initial clinical visit. TG was measured by the glycerol-blanked enzymatic method (Roche Diagnostics Corporation, Indianapolis, IN, USA). VLDL–TG was measured by NMR, a method that has been shown to have a high degree of correlation with the more traditional method of ultracentrifugation separation in the GOLDN population.28 Between the initial and follow-up clinical visits, participants took 160 mg of fenofibrate per day (TriCor®, Abbott Laboratories, Chicago, IL, USA). Fenofibric acid, the active moiety of fenofibrate, was measured by HPLC and the area under the serum fenofibric acid concentration versus time curve was calculated from three measurements over a 6-h period following the last dose of fenofibrate.29 Plasma insulin and glucose levels were measured before and after fenofibrate treatment, and the change in insulin and glucose levels after fenofibrate treatment was calculated.
Genotyping
Ninety-one SNPs from 25 genes known to be involved in lipoprotein metabolism were identified through the genetic association literature and in some cases additional SNPs within a gene were identified using public databases such as dbSNP (http://www.ncbi.nlm.nih.gov/SNP/). SNPs were genotyped using the Applied Biosystems TaqMan SNP genotyping system according to the laboratory protocol (Applied Biosystems, Foster City, CA, USA). Internal controls and repetitive sampling methods were used to ensure genotyping quality. The overall genotyping error and missing genotype call rate were approximately 1%. SNP allele and genotype frequencies and P-values for tests of the Hardy–Weinberg equilibrium are provided in Supplementary Table 2.
Statistics
All analyses were carried out using the R statistical language, v.2.4.30 Correlations between clinical covariates were estimated using Pearson's product moment correlation. Allele and genotype frequencies were calculated using standard gene counting methods. Linkage disequilibrium (LD), as measured by r2, was estimated using an expectation maximization algorithm.31 The Hardy–Weinberg equilibrium was assessed using a χ2 test or Fisher's exact test if a genotype class had less than five individuals.32 Covariates showing a large deviation from a normal distribution in diagnostic plots, such as fasting TG, were log10-transformed. Five individuals were considered outliers (>4 standard deviations from the mean) and were removed from the analysis due to extreme values of ΔTG. If an individual had a missing value for a single SNP or covariate, the individual was excluded from association tests involving that variable.
TG or VLDL–TG response to fenofibrate was defined as the difference between fasting TG and VLDL–TG concentration in mg/dl at the first and second visit, denoted ΔTG and ΔVLDL. In all models, ΔTG and ΔVLDL were adjusted for age, sex, waist-to-hip ratio and smoking status (ever versus never) because these variables have known associations with serum lipoprotein levels33, 34, 35 or because they were statistically significant predictors of TG and lipoprotein variables in this data set. A person was considered as having ever smoked if they had smoked more than 100 cigarettes during their lifetime.
We employed a two-step testing strategy to reduce the number of false-positive results. In step 1, we tested for associations between each of the predictors (SNPs and clinical covariates) with ΔTG and ΔVLDL in the full data set using least-squares linear regression methods.36 We also tested for association between each single SNP and each covariate to discern whether a SNP's effect on the outcome was being mediated through a particular covariate. Results were eligible for step 2 if the predictor was significant at a P-value ≤0.1. To determine if interactions among predictors explained additional variation in either outcome, we tested pairwise interactions among all possible pairs of predictors (ie, SNP–SNP, SNP–covariate, and covariate–covariate interactions) in the full data set. Associations involving interactions were assessed with a partial F-test,37 which compares a full model that includes both the interaction terms and the main effects of the variables comprising the interaction terms to a reduced model that includes only the main effects. Interaction models that had a partial F P-value ≤0.1 were eligible for step 2.
In step 2, associations (main effects or interactions) with P-values <0.1 in step 1 were evaluated for their predictive ability in independent test samples using a 10-repetition fourfold cross-validation strategy.38, 39 Cross-validation is an internal validation strategy that seeks to reduce false-positive results by eliminating associations that lack predictive ability in independent test sets.40 Because complex traits are likely influenced by a number of moderate genetic effects, P-value correction can be overly conservative and can lead to the removal of important associations. Cross-validation provides a direct assessment of the predictive ability of an association in a new, independent sample of individuals and does not depend strictly (ie, monotonically) on the P-value of an association. Results can be prioritized based on cross-validation, and only those that meet a preset threshold of predictive ability are considered potential ‘true positives.’ Cross-validation methods have been widely used over the last decade in genomic,41, 42 metabolomic,43 proteomic,44, 45 and transcriptomic46, 47, 48 studies as a method for discriminating between true associations and false-positive associations.
For each association, the fourfold cross-validation procedure began by randomly dividing the sample into four equally sized groups without regard to family structure. Three of the four groups were combined into a training data set, and the modeling strategy outlined above (in step 1) was carried out to estimate model coefficients for the training data set. Models were then applied to the fourth group, the testing data set, to make predictions about the value of the outcome variable of each individual in the independent test sample. This process was repeated for each of the four testing sets. Predicted values for all individuals in the test set were then subtracted from their observed values, yielding the total predictive error variability (SSPE), 
. The total variability in the outcome (SST)—the difference between each individual's observed value and the mean value for the outcome—was then calculated, 
. To estimate the proportion of variation in the outcome predicted in the independent test samples the cross-validated R2 (CV R2) was calculated as follows:  .
.
Cross-validation provides a more accurate measure of the predictive ability of the genetic models than the R2 values from linear modeling and will be negative when the model's predictive ability is poor. Because random variations in the sampling of the four mutually exclusive test groups can potentially impact the estimates of CV R2, this fourfold cross-validation procedure was repeated 10 times for each association and the CV R2 values were averaged.38 Univariate associations and interaction effects were considered cross-validated if the average percentage of variation predicted in independent test samples was greater than 0.5.
Each association that was validated in independent test samples in step 2 was then analyzed to determine whether the family structure in the sample influenced the observed association in the full data set. Generalized estimating equations were used to adjust the standard errors of the coefficients in both univariate and pairwise interaction models.49 We considered cross-validated associations with a P-value >0.1 on a global Wald test to be significantly influenced by family structure and therefore did not report them as cross-validated associations.
To visualize the complex genetic architecture, we applied a novel data visualization scheme, the KGraph, described by Kelly et al.50 The KGraph was developed for the visualization of multilocus genetic association results in the context of the underlying relationships among the predictors. We used the KGraph to simultaneously display significant univariate associations and pairwise interactions with the outcomes of interest, ΔTG and ΔVLDL, as well as the relationships among the predictors including SNP–SNP frequency correlations (ie, LD), SNP–covariate associations, and covariate–covariate correlations.
Understanding that total TG and VLDL–TG response to fenofibrate treatment is a complex system unlikely to be adequately explained by a single polymorphism or interaction, we constructed several prediction models using the four top-ranked significant cross-validated SNP–SNP and SNP–covariate interactions (ie, those that had the greatest predictive utility for the outcomes, ΔVLDL and ΔTG). As a point of reference, we compared these multivariable prediction models to a model including only baseline TG.
Results
Descriptive statistics of the clinical covariates and outcomes are shown in Table 2. The mean age of the participants was 47.8 years, 51.9% of participants were female, and 17.6% of participants were classified as an ever-smoker (smoked ≥100 cigarettes lifetime). Participants had a mean BMI of 28.5 kg/m2, waist-to-hip ratio of 0.9, and fasting total cholesterol level of 189 mg/dl. The mean fasting TG level was 132 mg/dl prior to fenofibrate treatment and 88 mg/dl after fenofibrate treatment. The mean ΔTG and ΔVLDL levels were −46.3 and −46.4 mg/dl respectively. Allele and genotype frequencies, rs numbers from dbSNP (build 127), SNP type (synonymous, non-synonymous, intron, and so on), and Hardy–Weinberg equilibrium test results are reported in Supplementary Table 2.
As described in the Methods section, association results were subjected to a two-step analysis process intended to reduce the occurrence of false-positive results and to identify associations with predictive ability in independent test samples. Table 3 shows the number of associations meeting the criteria for each of the steps (significant association and cross-validation) and the percentage of tests from the association step that cross-validated. For example, the 91 SNPs were evaluated for their association with ΔVLDL and ΔTG and in each case, 13 SNPs (14%) showed significant association with the outcome but only one SNP (APOA5_S19W) was validated in independent test samples by predicting >0.5% of the variation in each of these traits. In contrast, there were 1183 tests of SNP–covariate interactions for each outcome, and 156 (13%) tests were significant in each case, with 34 (3%) validated interactions predicting ΔVLDL and 25 (2%) validated interactions predicting ΔTG. Similarly, there were 4095 tests of SNP–SNP interactions for each outcome; 54 (1%) interactions were validated as predictors in independent test samples for ΔTG and 54 (1%) interactions were validated as predictors in independent test samples for ΔVLDL. While P-values and CV R2 were moderately correlated in tests with P-value ≤0.01 (R2=0.13), Figure 1 demonstrates that there is also a moderate amount of discrepancy between the two methods.
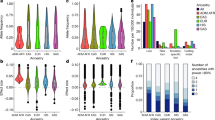
Partial F-test P-values versus CV R2 for SNP–covariate and epistatic (SNP–SNP) interactions predicting ΔTG and ΔVLDL. Percent variation explained in independent test samples (CV R2) versus −log10 partial F-test P-value for SNP–covariate and epistatic (SNP–SNP) interactions that were significant in ΔTG or ΔVLDL (partial F-test P-value ≤0.1). R2 for the correlation between −log P-value and CV R2 is 0.13. Lines are shown at −log P=0.05 and CV R2=0.5%.
Figure 2 is a visual representation of the complex genetic and clinical covariate associations underlying the ΔTG and ΔVLDL outcomes. Using both color and spatial relationships, the KGraph presents both associations with outcomes of interest and the correlation structure of the predictors that underlie those associations. SNPs without at least one significant and cross-validated association either univariately or interacting with another SNP or covariate in either outcome were excluded from display on the KGraph.
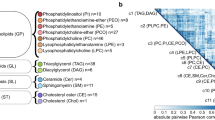
KGraph: fasting total TG and VLDL–TG and response to fenofibrate. Using 8 regions, the KGraph shows the relationships between the SNPs, clinical covariates, and outcomes by displaying the results from tests of correlation, linkage disequilibrium, association and cross-validation. The key at the bottom of the graphic shows the test criterion for each region and the colors associated with the test result. The region number key in the lower left corner shows the location of each region, and indicates whether the results in the region were assessed using cross-validation (shaded regions). Cross-validation of a specific test result is indicated by a black bar. Results for ΔTG are displayed in the top/right half of the cell and results for ΔVLDL are displayed in the bottom/left half. Region 1 displays the association between the SNPs and the clinical covariates, region 2 displays the correlation between the clinical covariates, and region 3 displays the linkage disequilibrium between the SNPs. Region 4 displays covariate association with the outcomes (ΔTG and ΔVLDL), region 5 displays SNP association with the outcomes, region 6 displays covariate–covariate interactions predicting the outcomes, region 7 displays SNP–covariate interactions predicting the outcomes, and region 8 displays SNP–SNP (epistatic) interactions predicting the outcomes.
Region 1 of Figure 2, shown in green, displays the association between the SNPs and clinical covariates. Though this step is often overlooked in association studies, understanding the relationship between SNPs and the covariates can be informative in discerning how the SNPs' effect on the outcome is mediated. In this region, there are a number of cross-validated associations between SNPs in the apolipoprotein gene cluster, which contains APOA1, APOA4, APOA5, and APOC3, and fasting baseline TG or cholesterol. Region 2, shown in gray, illustrates the correlations between the covariates. The majority of the covariates are not significantly correlated (∣r∣<0.3), with only two pairs (glucose before and after fenofibrate treatment and insulin before and after fenofibrate treatment) highly correlated (∣r∣>0.6). LD between SNPs is shown in red in Region 3. Most of the LD observed occurs between SNPs within the same gene, primarily in APOC3, and among other SNPs in the apolipoprotein gene cluster.
The remaining regions are colored in blue, and they represent associations with ΔTG and ΔVLDL. In these regions, the upper right section of the cells represents results for ΔTG and the lower left section represents results for ΔVLDL. Cross-validated associations are indicated by a black bar. Region 4, which displays the univariate association between clinical covariates and the outcomes, shows that baseline TG, cholesterol, and both insulin and glucose before fenofibrate treatment have cross-validated associations with both ΔTG and ΔVLDL, while fenofibric acid area under the curve, glucose level after fenofibrate, and waist-to-hip ratio have cross-validated associations with ΔVLDL but not ΔTG. Region 5, which illustrates univariate association between the SNPs and the outcomes, reveals that only one SNP in APOA5 (APOA5_S19W) has a cross-validated association with both ΔTG and ΔVLDL.
Region 6 displays the covariate–covariate interactions significantly associated with ΔTG and/or ΔVLDL. The majority of the cross-validated interactions were observed between fasting baseline TG and measures of glucose and insulin level before and after fenofibrate and between fenofibric acid area under the curve and insulin level before and after fenofibrate.
Region 7 displays the interactions between the SNPs and covariates that were associated with ΔTG and/or ΔVLDL. A complete listing of all significant and cross-validated SNP–covariate interactions and the corresponding relationships between genotypes and phenotypes are provided in Supplementary Tables 3, 5, and 6. Five of the clinical covariates (sex, baseline TG, fenofibric acid area under the curve, baseline glucose, and change in glucose after fenofibrate) accounted for a large portion of SNP–covariate interactions: 15 (60%) of the validated interactions predicted ΔTG and 16 (47%) of the validated interactions predicted ΔVLDL. The predictive abilities of these interactions ranged from 0.50 to 1.90% (mean=0.89%) for ΔTG and 0.51–1.47% (mean=0.92%) for ΔVLDL. The ΔR2 × 100 values (adjusted R2 from the model including interactions minus adjusted R2 from the main effects only model) for the same interactions, which range from 0.58 to 2.29% (mean=1.35%) for ΔTG and 0.67–2.40% (mean=1.39%) for ΔVLDL, are generally overly optimistic. Baseline glucose and the area under the fenofibric acid concentration curve, a measure of an individual's plasma level of the active form of fenofibrate over a 6 h time period, were most sensitive to genetic context. They each interacted with five SNPs in their effect on ΔVLDL and also interacted with three SNPs in their effect on ΔTG. Sex interacted with three SNPs in its effects on ΔTG and on ΔVLDL, and baseline TG (log) and change in glucose after fenofibrate had three interactions with SNPs in their effects on ΔTG.
Overall, four SNPs are responsible for a large portion of the SNP–covariate interactions that predict ΔTG (APOA4_M35, APOC3_3U386, ABCA1_I27943, and LIPC_T224T). Together, these SNPs account for eight (32%) of the validated SNP–covariate interactions in their effect on ΔTG, with predictive ability ranging from 0.52 to 1.29% (mean=0.82%) and ΔR2 × 100 values ranging from 0.73 to 1.33% (mean=1.02%). With respect to predicting ΔVLDL, four SNPs are responsible for a substantial proportion of the SNP–covariate interactions (ABCG8_C54Y, LIPC_i67180, FABP1_m2353, and FABP1_T94A). These SNPs account for 10 (29%) of the validated SNP–covariate interactions for ΔVLDL, with predictive abilities ranging from 0.56 to 1.47% (mean=0.90%) and ΔR2 × 100 values ranging from 0.87 to 2.40% (mean=1.41%). For both outcomes, the remaining SNPs were involved in at most one validated interaction with a covariate (see Supplementary Table 3 for details).
Region 8 displays the epistatic interactions among the SNPs that have a significant and validated predictive ability for ΔTG and ΔVLDL. As with SNP–covariate interactions, a large proportion of epistatic interactions are accounted for by a small number of SNPs. The SNPs that have the most interactions that were validated as predictors of ΔTG are APOA5_S19W with six interactions, PDZK1_i4201 with five interactions, and APOA1_M2630 and FABP1_T94A with four interactions each. Together, these SNPs account for 16 (30%) of the validated epistatic interactions that predict ΔTG, have predictive abilities ranging from 0.53 to 2.19% (mean=0.85%), and have ΔR2 × 100 values ranging from 0.71 to 2.80 (mean=1.58%). The SNPs with the most interactions that were validated as predictors of ΔVLDL are APOA5_S19W with six interactions, APOB_E4181K with five interactions, and APOA1_M3012, APOB_T2515T, and APOC3_3U386 with four interactions each. These SNPs account for 18 (33%) of the validated epistatic interactions that predict ΔVLDL, have predictive abilities ranging from 0.51 to 2.42% (mean=0.81%), and have ΔR2 × 100 values ranging from 0.52 to 2.56% (mean=1.18%). A complete listing of all significant and cross-validated epistatic interactions is given in Supplementary Table 6.
Results for the multivariable modeling using the top-ranked SNP–SNP and SNP–covariate interactions are shown in Table 4. Only two of the SNPs used in the models had cross-validated associations with any of the covariates (PPARG_P12AEB with fenofibric acid area under the curve and PPARG_C1A i27289 with insulin after fenofibrate), which indicates that the associations between these top-ranked SNPs and the outcome is not primarily a result of their relationship with the covariates. A model that includes the four top-ranked epistatic (SNP–SNP) interactions minimally increases the percentage of variation explained beyond that explained by baseline TG alone (0.4% for ΔTG and 0.4% for ΔVLDL). However, a model that includes the four top-ranked SNP–covariate interactions significantly increases the percentage of variation explained beyond baseline TG (11.9% for ΔTG and 7.8% for ΔVLDL).
Discussion
Understanding the genetic architecture that underlies a person's TG response to fenofibrate will help identify those who will most likely benefit from fenofibrate treatment and shed light on the complexities of the network that controls the metabolic response to drugs such as fenofibrate. By looking beyond simple, single SNP effects and examining interactions between SNPs and clinical covariates and pairs of SNPs, this study takes an important first step toward the mapping of this genetic architecture.
The key factor in an analysis such as this is to attempt to appropriately model the system's complexity to the extent possible. Fasting TG response to fenofibrate treatment is influenced by environmental changes (at both the cellular and organismal levels) and large-scale gene expression networks that are controlled in part by genetic variations. A study of this size is unlikely to be adequately powered to reconstruct this network in its entirety, but can be used to begin constructing a better map of the gene–covariate interactions and associations that play a role in interindividual variation.
As the number of SNPs considered in an association study increases, so does the problem of separating true results from spurious results caused by random chance.51 Attempts to combat this problem often focus on P-values, with the rationale that by choosing the models least likely to be found by random chance alone, the probability of including a false positive will be greatly reduced.52 Unfortunately, when considering a complex trait, the effect sizes of individual SNPs and pairwise interactions are generally fairly small. This limits the significance level that these models are able to attain in typical epidemiologic studies, which means that true positives may be removed during P-value adjustment. Additionally, the very low P-values that are required for an association to remain significant after adjustment (eg, by Bonferroni correction or false discovery rate adjustment) are likely to be obtained by SNPs with low minor allele frequencies that capture groups on the extreme of the phenotypic tail (sometimes simply by chance alone).
We approached the problem from a different direction. The utility of the associations being investigated lies in their ability to make a prediction about the outcome of interest, so it is natural to select the best models based on predictive ability. By using a cross-validation procedure that successively leaves one quarter of the data out and constructs genetic models in the remaining three quarters of the data, we can estimate the performance of the model on a new, independent sample of individuals from the same population.
Interestingly, the correlation between P-value and CV R2 is moderate at best (Figure 1), and many associations with only marginally significant P-values have a non-negative CV R2. With the movement toward genome-wide association studies that measure hundreds of thousands of SNPs, the problems in correcting for multiple testing will be magnified. Cross-validation is an attractive option to genetic epidemiologists both because it offers a direct assessment of an association's predictive utility and because of the relative simplicity and stability of the cross-validation estimate.38 In addition, replication in an independently collected sample has become the standard tool to validate genetic association results. However, no comparable samples exist for specialized intervention trials such as this one. In the absence of replication samples, cross-validation gives us the ability to validate our results while simultaneously assessing their predictive capacity. Without a replicate sample, however, we cannot estimate whether the predictive ability of the multiple variable models established in this data set would be the same in other data sets.
The 91 SNPs investigated in this study span genes known to be involved in many facets of lipid metabolism. Interestingly, 78 of them (86% of the SNPs) have at least one significant cross-validated association with a clinical covariate or have at least one epistatic or SNP–covariate interaction that predicted either outcome. It is sobering but not completely unexpected that individual polymorphisms and interactions make relatively small contributions to explaining the variation in ΔTG and ΔVLDL. However, the percentage of trait variation explained by SNPs in this study is similar to that observed in studies of other complex phenotypes. For example, a recent genome-wide association study of diabetes and associated traits showed that the single SNPs most strongly associated with TG level explained between 0.5 and 1.2% of the residual variance of TG level after adjustment for traditional risk factors.53 A candidate gene study of blood pressure phenotypes found that the CV R2 of the most predictive SNPs for systolic blood pressure ranged between 1.9 and 6.2%.54
Combining these individual interactions into a multiple variable model did increase the amount of variation explained, even if only moderately in the case of epistatic interactions. We do note that, in general, non-cross-validated genetic models all overestimate the contribution of each predictor or interaction, so it is likely that the percentage of variation explained using cross-validation, while small, is a more realistic estimate than R2 values obtained without cross-validation.
The large number of SNPs involved in significant cross-validated interactions illustrates the complexity of the network underlying these traits, comprising single gene effects and interactions among multiple variations in many genes as well as their interaction with clinical factors. Also, while there is substantial overlap between the clinical factors and interactions that predicted ΔTG and ΔVLDL, there is also evidence of associations that are unique to each outcome, which shows that there are genetic and environmental interactions that affect the TG response to fenofibrate during fasting that lie outside of the metabolism of VLDL particles.
As genotyping has become increasingly less expensive, the scale of genetic association studies has risen dramatically,55 but most studies, even genome-wide studies, concentrate on single, univariate associations. There is no denying that some single polymorphisms can significantly influence physiology; many diseases are the result of a single genetic error with serious phenotypic consequences. Complex, non-Mendelian traits, however, are likely influenced less by single polymorphisms than by interacting effects, particularly epistatic interactions.56 Our results indicate that a wealth of pairwise interactions can be found even in this fairly limited subset of polymorphisms. Methods, such as the multivariable modeling approach that we describe, must now be developed that will allow the integration of these results into a predictive framework, so that the understanding of the genetic architecture of TG response to fenofibrate treatment can be refined and harnessed.
References
Haymore BR, Parks JR, Oliver TG, Glister BC : Hypertriglyceridemia. Hosp Physician 2005; 41: 17–24.
Cullen P : Evidence that triglycerides are an independent coronary heart disease risk factor. Am J Cardiol 2000; 86: 943–949.
Hopkins PN, Wu LL, Hunt SC, Brinton EA : Plasma triglycerides and type III hyperlipidemia are independently associated with premature familial coronary artery disease. J Am Coll Cardiol 2005; 45: 1003–1012.
Ooi TC, Ooi DS : The atherogenic significance of an elevated plasma triglyceride level. Crit Rev Clin Lab Sci 1998; 35: 489–516.
Yoshino G, Hirano T, Kazumi T : Dyslipidemia in diabetes mellitus. Diabetes Res Clin Pract 1996; 33: 1–14.
McKenney JM, Sica D : Role of prescription omega-3 fatty acids in the treatment of hypertriglyceridemia. Pharmacotherapy 2007; 27: 715–728.
Stein EA, Lane M, Laskarzewski P : Comparison of statins in hypertriglyceridemia. Am J Cardiol 1998; 81: 66B–69B.
Chapman MJ : Fibrates in 2003: therapeutic action in atherogenic dyslipidaemia and future perspectives. Atherosclerosis 2003; 171: 1–13.
Keating GM, Ormrod D : Micronised fenofibrate: an updated review of its clinical efficacy in the management of dyslipidaemia. Drugs 2002; 62: 1909–1944.
Tsimihodimos V, Miltiadous G, Daskalopoulou SS, Mikhailidis DP, Elisaf MS : Fenofibrate: metabolic and pleiotropic effects. Curr Vasc Pharmacol 2005; 3: 87–98.
Mason TM : The role of factors that regulate the synthesis and secretion of very-low-density lipoprotein by hepatocytes. Crit Rev Clin Lab Sci 1998; 35: 461–487.
Staels B, Dallongeville J, Auwerx J, Schoonjans K, Leitersdorf E, Fruchart JC : Mechanism of action of fibrates on lipid and lipoprotein metabolism. Circulation 1998; 98: 2088–2093.
Merkel M, Heeren J : Give me A5 for lipoprotein hydrolysis!. J Clin Invest 2005; 115: 2694–2696.
Shah RR : Pharmacogenetics in drug regulation: promise, potential and pitfalls. Philos Trans R Soc Lond B Biol Sci 2005; 360: 1617–1638.
Sing CF, Stengard JH, Kardia SL : Genes, environment, and cardiovascular disease. Arterioscler Throm Vasc Biol 2003; 23: 1190–1196.
Ordovas JM, Corella D : Nutritional genomics. Annu Rev Genomics Hum Genet 2004; 5: 71–118.
Hachem SB, Mooradian AD : Familial dyslipidaemias: an overview of genetics, pathophysiology and management. Drugs 2006; 66: 1949–1969.
Corella D, Ordovas JM : Single nucleotide polymorphisms that influence lipid metabolism: interaction with dietary factors. Annu Rev Nutr 2005; 25: 341–390.
Ordovas JM : HDL genetics: candidate genes, genome wide scans and gene-environment interactions. Cardiovasc Drugs Ther 2002; 16: 273–281.
Ye SQ, Kwiterovich Jr PO : Influence of genetic polymorphisms on responsiveness to dietary fat and cholesterol. Am J Clin Nutr 2000; 72: 1275S–1284S.
Brisson D, Ledoux K, Bosse Y et al: Effect of apolipoprotein E, peroxisome proliferator-activated receptor alpha and lipoprotein lipase gene mutations on the ability of fenofibrate to improve lipid profiles and reach clinical guideline targets among hypertriglyceridemic patients. Pharmacogenetics 2002; 12: 313–320.
Talmud PJ, Hawe E, Robertson K, Miller GJ, Miller NE, Humphries SE : Genetic and environmental determinants of plasma high density lipoprotein cholesterol and apolipoprotein AI concentrations in healthy middle-aged men. Ann Hum Genet 2002; 66: 111–124.
Lai CQ, Parnell LD, Ordovas JM : The APOA1/C3/A4/A5 gene cluster, lipid metabolism and cardiovascular disease risk. Curr Opin Lipidol 2005; 16: 153–166.
Yamada M : Influence of apolipoprotein E polymorphism on bezafibrate treatment response in dyslipidemic patients. J Atheroscler Thromb 1997; 4: 40–44.
Zhang X, Chen ZQ, Wang Z, Mohan W, Tam SP : Protein-DNA interactions at a drug-responsive element of the human apolipoprotein A-I gene. J Biol Chem 1996; 271: 27152–27160.
Nemeth A, Szakmary K, Kramer J et al: Apolipoprotein E and complement C3 polymorphism and their role in the response to gemfibrozil and low fat low cholesterol therapy. Eur J Clin Chem Clin Biochem 1995; 33: 799–804.
Lai CQ, Arnett D, Straka RJ et al: Fenofibrate effect on triglyceride and postprandial response of apolipoprotein A5 variants: the GOLDN study. Arterioscler Thromb Vasc Biol 2007; 27: 1417–1425.
Tsai MY, Georgopoulos A, Otvos JD et al: Comparison of ultracentrifugation and nuclear magnetic resonance spectroscopy in the quantification of triglyceride-rich lipoproteins after an oral fat load. Clin Chem 2004; 50: 1201–1204.
Straka RJ, Burkhardt RT, Fisher JE : Determination of fenofibric acid concentrations by HPLC after anion exchange solid-phase extraction from human serum. Ther Drug Monit 2007; 29: 197–202.
R Core Development Team: R: A language and environment for statistical computing, 2008.
Lynch M, Walsh B : Genetics and Analysis of Quantitative Traits. Massachussettes: Sinauer Associates, Inc., 1998.
Weir B : Genetic Data Analysis II. Massachussettes: Sinauer Associates, Inc., 1996.
Freedman DS, Otvos JD, Jeyarajah EJ et al: Sex and age differences in lipoprotein subclasses measured by nuclear magnetic resonance spectroscopy: the Framingham study. Clin Chem 2004; 50: 1189–1200.
Mansfield E, McPherson R, Koski KG : Diet and waist-to-hip ratio: important predictors of lipoprotein levels in sedentary and active young men with no evidence of cardiovascular disease. J Am Diet Assoc 1999; 99: 1373–1379.
Billimoria JD, Pozner H, Metselaar B, Best FW, James DC : Effect of cigarette smoking on lipids, lipoproteins, blood coagulation, fibrinolysis and cellular components of human blood. Atherosclerosis 1975; 21: 61–76.
Kleinbaum D, Kupper L, Muller K, Nizam A : Applied Regression Analysis and Other Multivariate Methods. California: Brooks/Cole Publishing Company, 1998.
Rosner BA : Fundamentals of Biostatistics. United States: Thomson Brooks/Cole, 2006.
Molinaro AM, Simon R, Pfeiffer RM : Prediction error estimation: a comparison of resampling methods. Bioinformatics 2005; 21: 3301–3307.
Browne MW : Cross-validation methods. Journal of Mathematical Psychology 2000; 44: 108–132.
Stone M : Cross-validatory choice and assessment of statistical predictions. J R Statist Soc B 1974; 36: 111–147.
Motsinger AA, Ritchie MD : Multifactor dimensionality reduction: an analysis strategy for modelling and detecting gene–gene interactions in human genetics and pharmacogenomics studies. Hum Genomics 2006; 2: 318–328.
Gong R, Liu Z, Li L : Epistatic effect of plasminogen activator inhibitor 1 and beta-fibrinogen genes on risk of glomerular microthrombosis in lupus nephritis: interaction with environmental/clinical factors. Arthritis Rheum 2007; 56: 1608–1617.
Pohjanen E, Thysell E, Jonsson P et al: A multivariate screening strategy for investigating metabolic effects of strenuous physical exercise in human serum. J Proteome Res 2007; 6: 2113–2120.
Agranoff D, Fernandez-Reyes D, Papadopoulos MC et al: Identification of diagnostic markers for tuberculosis by proteomic fingerprinting of serum. Lancet 2006; 368: 1012–1021.
Mertens BJ, De Noo ME, Tollenaar RA, Deelder AM : Mass spectrometry proteomic diagnosis: enacting the double cross-validatory paradigm. J Comput Biol 2006; 13: 1591–1605.
Feng Z, Prentice R, Srivastava S : Research issues and strategies for genomic and proteomic biomarker discovery and validation: a statistical perspective. Pharmacogenomics 2004; 5: 709–719.
Zhu J, Hastie T : Classification of gene microarrays by penalized logistic regression. Biostatistics 2004; 5: 427–443.
Wood IA, Visscher PM, Mengersen KL : Classification based upon gene expression data: bias and precision of error rates. Bioinformatics 2007; 23: 1363–1370.
Liang KY, Zeger SL : Regression analysis for correlated data. Annu Rev Public Health 1993; 14: 43–68.
Kelly RJ, Jacobsen DM, Sun YV, Smith JA, Kardia SL : KGraph: a system for visualizing and evaluating complex genetic associations. Bioinformatics 2007; 23: 249–251.
Thomas DC, Haile RW, Duggan D : Recent developments in genomewide association scans: a workshop summary and review. Am J Hum Genet 2005; 77: 337–345.
Page GP, George V, Go RC, Page PZ, Allison DB : Are we there yet? Deciding when one has demonstrated specific genetic causation in complex diseases and quantitative traits. Am J Hum Genet 2003; 73: 711–719.
Diabetes Genetics Initiative of Broad Institute of Harvard and MIT, Lund University, and Novartis Institutes of BioMedical Research Saxena R, Voight BF, Lyssenko V et al: Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 2007; 316: 1331–1336.
Kardia SL, Sun YV, Hamon SC, Barkely RA, Boerwinkle E, Turner ST : Interactions between the adducin 2 gene and antihypertensive drug therapies in determining blood pressure in people with hypertension. BMC Med Genet 2007; 8: 61.
Hirschhorn JN, Daly MJ : Genome-wide association studies for common diseases and complex traits. Nat Rev Genet 2005; 6: 95–108.
Wagner GP, Laubichler MD, Bagheri-Chaichian H : Genetic measurement of theory of epistatic effects. Genetica 1998; 102–103: 569–580.
Acknowledgements
We thank the families that participated in the GENOA study and Jian Chu, Douglas Jacobsen, Kristin Meyers, and Todd Greene for editorial and computing assistance. This work was supported by grant U 01 HL72524 from the National Heart, Lung, and Blood Institute and training grant T32 HG00040 from the National Human Genome Research Institute (JAS). The authors have no conflicts of interest to disclose relating to the subject of this paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Rights and permissions
About this article
Cite this article
Smith, J., Arnett, D., Kelly, R. et al. The genetic architecture of fasting plasma triglyceride response to fenofibrate treatment. Eur J Hum Genet 16, 603–613 (2008). https://doi.org/10.1038/sj.ejhg.5202003
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5202003
Keywords
This article is cited by
-
Fenofibrate and Dyslipidemia: Still a Place in Therapy?
Drugs (2018)
-
Central obesity in males affected by a dyslipidemia-associated genetic polymorphism on APOA1/C3/A4/A5 gene cluster
Nutrition & Diabetes (2013)
-
The PPAR alpha gene is associated with triglyceride, low-density cholesterol and inflammation marker response to fenofibrate intervention: the GOLDN study
The Pharmacogenomics Journal (2013)
-
Profile of Serum Bile Acids in Noncholestatic Volunteers: Gender-Related Differences in Response to Fenofibrate
Clinical Pharmacology & Therapeutics (2011)