Abstract
The Sahel that extends from the Atlantic Ocean to the Ethiopian highland is a historical reservoir of Africa's cultures and grandest populations and a known arena of ancient and recent migrations. We are interested in the issue whether such migrations were also carriers of genetic traits and whether this introgression could be associated with population genetic markers. Based on analysis of Y-chromosome haplogroups, we present evidence that the sickle gene, one of the major protective polymorphisms known in malaria, has in fact found its way only recently to the gene pool of the populations in eastern Sahel. We discuss the possible dynamics of the process and give estimates of the age of the introduction of the S allele into eastern Sahel.
Similar content being viewed by others
Introduction
The origin of malaria-resistant sickle cell genetic variants from either single or multiple events was controversial until recently.1 The debate was partly settled in favor of multiple origins; owing to βS haplotypes in linkage disequilibrium with the sickle mutation.2, 3 Complementary evidence for parallel origins of sickle cell is discernable in ethnicity, some notable examples being the concurrency of gene population spread attributed to expansions of Iron Age Bantu speakers both south and eastwards in Africa emerging from the area of present-day Cameroon, and subsequently across the Atlantic by events associated with the ‘Middle Passage’.
Here we report Y-chromosome co-introgression with the S gene in the eastern part of Africa's Sahel, where it is known that the gene existed previously in low frequencies4, 5 in contrast to being more prevalent among west African migrants.
Materials and methods
A total of 237 unrelated males (some of whom are known sicklers) were typed, for the sickle-cell mutations using amplification refractory mutation system PCR and the results confirmed by protein electrophoresis. The same sample was typed for the major Y chromosome haplogroup-defining mutations. The populations studied included Hausa, Masalit, Borgu as well as other groups and populations inhabiting the area across Africa's Sahel into Chad and Sudan. Populations were classified according to their linguistic affiliation, which comprised the main linguistic families of Africa (Afro-Asiatic, Niger Kordofanian and Nilo-Saharan). The population sample was selected based on preliminary knowledge of the distribution of the S gene among various groups in the Sudan.4, 5, 6 The sample was further classified to include the Hausa as a separate group due to the high frequency of a particular distinctive haplogroup within this group (P25) whose closest phylogenetic molecular ancestor, haplogroup R1-M173* marks a back migration to Africa from west Eurasia.7 The Hausa, originally of western Africa, have migrated to the eastern Sahel during the past 300 years, employed mainly in agricultural economies (Figure 1).
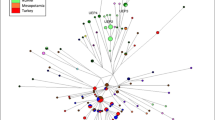
Map of Africa showing the hypothetical movement of Y-chromosome haplogroups across the Sahel. The yellow color of the arrow is for the haplogroups in non-sicklers and the white arrow is for sicklers. The various colors represent approximate estimations of the frequency of the S gene in the continent with red representing the highest frequency followed by the dark and light blue for lower frequencies.
We employed an extension of the coalescence to estimate the possible age and date of introduction of the S gene into the eastern Sahel, using the following formula provided an equal assortment and no gender preferential introduction of the gene:

where X(t) is the frequency of the Y haplogroup among bearers of S mutation in generation t and Y is the frequency of Y haplogroup in the chromosome and a is the decay in association due to assortment.
Results
Table 1 shows the distribution of the S gene in 122 cases and 115 controls with Y chromosome genotyped for the major haplogroups. Two haplogroups, R1b-P25 and E3b-M78, display the highest frequency, and were differentially distributed among the various groups and between sicklers and non-sicklers. The distribution of the observed values were significantly different from the expected (Pearson χ2 44.9 P<0.000). The association of the P25 with the S gene in Hausa, which barely holds (P=0.07) in spite of the relatively large effective population size of the Hausa (Hassan et al, unpublished), might indicate that the S mutation may have been affiliated with a male founder belonging to Y haplogroup R1b-P25. The observed frequency among S-gene carriers in populations other than Hausa makes a strong case for gene flow from the Hausa to local groups in Sudan.
The dating estimates based on a formula that is an extension of the coalescence gives a recent figure of 1–3 generations for the introduction of the gene and associated haplotypes to eastern Sahel.
Discussion
Patterns of genetic diversity are the result of locus-specific forces (natural selection) and population-level forces (eg demographic growth and range expansion). When multiple independent loci correlate with geography, population-level forces are likely responsible. Conversely, when patterns diverge, natural selection is implicated in modulating observed diversity.
Mitochondrial DNA and Y-chromosome diversity closely correlate with geography and these loci have recently become the markers of choice in defining both provenance and trajectory of population movements and also provide insights into population substructure and group membership. By typing both the haploid Y chromosome and the S gene in the same samples it should be possible to test for common demography as well as detect affinities of particular S variants observed in Sudan to other regions of Africa.
Our results suggest that the sickle cell gene may have been preferentially introduced through males of migrating west African tribes (Figure 1), particularly Hausa-Fulani, and Bagara in the large migrations that began in the eighteenth century and escalated during the nineteenth and early twentieth century. The estimates of a recent figure of 1–3 generations for the introduction of the gene and associated haplotypes to eastern Sahel, is consistent with demography during the past 100 years and with a hypothesis of a recent origin of malaria as a major human infection.6, 8, 9
References
Pagnier J, Mears JG, Dunda-Belkhodja OD et al: Evidence for the multicentric origin of the sickle cell hemoglobin gene in Africa. Proc Natl Acad Sci USA 81, (1984); 1771–1984.
Lanclos KD, Oner C, Dimovski AJ, Gu YC, Huisman TH : Sequence variations in the 5′ flanking and IVS-II regions of the G gamma- and A gamma-globin genes of ßS chromosomes with five different haplotypes. Blood (1991); 77: 2488–2496.
Oner C, Dimovki A, Olivieri N et al: βs haplotypes in various world population's. Hum Genet (1992); 89: 99–104.
Elhassan AM, Godber MG, Kopec AC, Mourant AE, Tills D, Lehmann H : The inherited blood factors of the Beja of the Sudan. Man (1968); 3: 272.
Bayoumi RA, Taha TS : A study of some genetic characteristics of the Fur and Baggara tribes of the Sudan. AM J Phys Anthropol (1985); 67: 363–370.
Mohammed AO, Attalla B, Bashir FMK et al: The relationship of the sickle cell gene to the ethnic and geographic groups populating the sudan. Community Genet (2006); 9: 113–120.
Cruciani F, Santolamazza P, Shen P et al: A back migration from Asia to sub-Saharan Africa is supported by high-resolution analysis of human Y-chromosome haplotypes. Am J Hum Genet (2002); 70: 1197–1214.
Tishkoff SA, Varkonyl R, Cahinhinan N et al: Haplotype diversity and linkage disequilibrium at human G6PD: Recent origin of alleles that confer malaria resistance. Science (2001); 293: 455–462.
Volkman SK, Barry AE, Lyons EJ et al: Recent origin of Plasmodium falciparum from a single progenitor. Science (2001); 293: 482–483.
Acknowledgements
This work is part of the activities of the BioMalPar European Network of Excellence supported by a European grant (LSHP-CT-2004–503578) from the Priority 1 ‘Life Sciences, Genomics and Biotechnology for Health’ in the 6th Framework Programme.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bereir, R., Hassan, H., Salih, N. et al. Co-introgression of Y-chromosome haplogroups and the sickle cell gene across Africa's Sahel. Eur J Hum Genet 15, 1183–1185 (2007). https://doi.org/10.1038/sj.ejhg.5201892
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201892
Keywords
This article is cited by
-
Y-chromosome E haplogroups: their distribution and implication to the origin of Afro-Asiatic languages and pastoralism
European Journal of Human Genetics (2014)
-
Y-chromosome R-M343 African lineages and sickle cell disease reveal structured assimilation in Lebanon
Journal of Human Genetics (2011)
-
A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe
European Journal of Human Genetics (2011)
-
Loss of balancing selection in the βS globin locus
BMC Medical Genetics (2010)