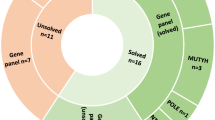
Abstract
Partial and whole gene deletions represent a large proportion (4–33%) of the APC mutations found in polyposis patients, who previously had negative test results. The genotype–phenotype correlations for these APC deletions have not been studied in detail. We aimed to assess the number of germ line APC deletions in Dutch polyposis patients, to describe the clinical phenotype(s), and to review the current literature. We screened 296 index patients with polyposis, who previously had negative test results for APC or MUTYH mutations, for germ line APC gene deletions using Multiplex Ligation-dependent Probe Amplification. APC deletions were identified in 19 polyposis patients; seven had a whole gene deletion, nine had a deletion involving two or more exons, and three had single exon deletions. Most of the deletion families (83%) displayed a classic familial adenomatous polyposis (FAP) phenotype (100–2000 adenomas). We saw no patients with APC deletions and a severe phenotype (ie >2000 polyps); on the contrary, two families carrying a deletion of exons 7–13 and one family with a deletion of exons 1–5 showed a distinctly attenuated FAP phenotype. APC deletions were found in a considerable proportion of polyposis patients previously tested negative for APC or MUTYH (6%, 19/296) and represent 8% of all APC mutations found at our clinics (19/242). Methods to identify such deletions should therefore be included in routine germ line APC mutation analysis. While most total and partial APC deletions lead to a classic FAP phenotype, specific (in-frame) deletions may lead to an attenuated polyposis phenotype.
Similar content being viewed by others
Introduction
Familial adenomatous polyposis, FAP (MIM175100), is an autosomal dominant inherited disease caused by germ line mutations in the APC gene; it affects approximately one in 13 500 people.1 Mutation carriers are predisposed in the majority of cases to develop hundreds or even thousands of polyps and subsequently go on to develop colorectal cancer (CRC).1, 2 In addition, extracolonic features such as fundic gland polyps (FGP), duodenal polyps, congenital hypertrophy of the retinal pigment epithelium (CHRPE) and desmoid tumors are often present, while a more attenuated form of FAP has also been described in which patients develop less than a hundred polyps at a relatively late age.3 Another inheritable form of polyposis is caused by recessive mutations of the MUTYH gene.4 MUTYH-associated polyposis (MAP) is found in 10–20% of polyposis patients.5, 6, 7
The APC gene is a relatively large gene, containing 15 exons and encoding a protein of 2843 amino acids. Exon 15 is by far the largest exon containing over three-quarters of the coding sequence.8 Depending on clinical features and mutation detection techniques applied, germ line mutations in the APC gene are found in 30–80% of patients with polyposis2, 9, 10 and most APC mutations occur in the 5′ half of its coding region. Genotype–phenotype relations have been reported extensively for small nucleotide alterations (point mutations). A classic phenotype (ie 100–2000 polyps) is associated with mutations in codon 168–1250 (exons 4/5–exon 15) or codon 1400–1580 (exon 15). Mutations occurring between codon 1250 and 1464 (exon 15), but particularly at codon 1309;2, 11 lead in most cases to a more severe phenotype (>2000 polyps at a relatively young age). A more attenuated form of polyposis is associated with mutations in three regions: (i) in the 5’ part of the APC gene; (ii) alternative spliced region in exon 9 and (iii) mutations in the extreme 3′ site of the gene.12, 13
Partial and whole gene deletions have been shown to represent a large proportion (4–33%) of APC mutations in polyposis patients who previously had negative test results.14, 15, 16, 17 Large deletions have also been found in other oncogenes such as BRCA1, BRCA2, MLH1, MSH2 and MSH6.17, 18, 19, 20 Recently, APC duplications of exon 4 and 15 have also been described in FAP patients.21, 22
Using the new MLPA (Multiplex Ligation-dependent Probe Amplification) technique,23 we searched for deletions of the whole gene of one or more exons in 296 APC- and MUTYH-negative index patients with polyposis. Here we report on genotype–phenotype correlations in 19 index patients with identified deletions and review the current literature on APC deletions.
Patients and methods
Between 1995 and 2005, a total of 599 index polyposis patients were referred for FAP testing to the Molecular Genetics Laboratory, Center for Human and Clinical Genetics (Leiden, The Netherlands). Informed consent was obtained for DNA testing according to protocols approved by LUMC Ethics Review Board.
The 599 index patients were all screened for germ line mutations in the APC gene by DGGE, PTT and sequence analysis. In 223/599 patients, we were able to identify a pathogenic APC mutation (37%). Mutations in the APC gene involve frameshift mutations (n=124, 55%), nonsense mutations (n=70, 31%) or splice mutations (n=26, 12%), and three large cytogenetically detected deletions (cytogenetic analysis is only done when developmental disorders are present).
In 296 polyposis index patients with available DNA, in whom no APC mutations were detected and MUTYH was ruled out in most of the cases (225/296), we went on to perform MLPA to look for APC deletions.
For MLPA, the P043 kit was used (http://www.mrc-holland.com). This kit contains 20 probes covering the APC region, three probes for the promoter regions, 14 for exons 1–14 and three for exon 15. The MLPA analysis was performed according to the MRC Holland protocol23 with some adaptation: all reagents in the kit were used at 1/4 of the advised volume and hybridization time was cut from 16 to 2.5 h. To exclude false-positive results due to variants underlying the probe, single exon deletions containing only one MLPA probe (exons 11 and 14) were confirmed by a long-range PCR (see Figure 1) with the following primers:
Two single-exon deletions (exons 11 and 14) confirmed by long-range PCR. An agarose gel shows the PCR products of two families with an exons 11 and 14 deletion, respectively (left), compared with controls (centre and right). In the patients, a smaller second fragment is also visible besides the normal product. WT, wild type; del, deletion.

The valued size of the deleted exons 11 and 14 was approximately 0.7 and 2.6 kb, respectively. A false-positive deletion was unlikely in the case of the single exon 15 deletion, since three MLPA probes were used for this exon. Clinical and pathological data were obtained from patients’ records to confirm the FAP diagnosis. The polyposis patients and their families were grouped according to the following phenotype definitions:
-
1
Severe: >2000 polyps.
-
2
Classic: 100–2000 polyps before the age of 30–40.
-
3
Attenuated: 10–99 adenomas diagnosed at a more advanced age (over 30–40 years). If other family members had a classic polyposis phenotype, in most cases the family was considered as classic FAP.
We describe the clinical details of Dutch index cases and their family members. Furthermore, we combined our results for index cases with whole and partial APC gene deletions with those of previous studies. For this, we searched the PubMed and EMBASE databases up to December 2006 using the terms ‘Adenomatous Polyposis Coli’, ‘Genes, APC’, ‘Gene Deletion’, ‘Gene Dosage’, ‘Gene Disruption, ‘Gene Loss’ and ‘Gene Dosage’, with relevant subheadings. We also searched reference lists in articles identified by this strategy and selected additional articles that we considered relevant, including only those describing large germ line APC deletions in patients that could not be detected by cytogenetic techniques. We calculated Pearson's R to analyze trends and considered P<0.05 statistically significant. All our results were analyzed with SPSS 11.01 (SPSS, Chicago, IL, USA).
Results
APC deletions were detected in 19/296 index polyposis patients. Their clinical details and their family members are shown in Table 1. Deletions spanned the whole APC gene in seven index cases,1, 2, 3, 4, 5, 6, 7 and clinical information was available for six of these; the remaining cases were classified as classic, but not severe. Cases with deletions of exons 9–15 (cases 13–15), exons 14–15 (case 18) and exon 15 (case 19) were also classified as classic. In four cases (9, 10, 16 and 17) with deletions of exons 4–5, 6–15, 11 and 14, no clear-cut phenotypic classification was possible. The index patients 9 and 10 developed less than 100 polyps, but they were young (27 and 20 years old, respectively) and were therefore classified as classic FAP. In cases 16 and 17, the index patients developed between 50 and 100 polyps at the ages of 36 and 44 years, respectively, but since a majority of family members showed a classic phenotype, both families were classified as classic. Two cases (11 and 12) with exons 7–13 deletion, and one case8 with an exons 1–5 deletion had an apparently attenuated phenotype.
Five (1, 2, 5, 9 and 14) of 18 index patients (28%) were apparently de novo cases, because parents and siblings had no polyps or CRC; in one case, no family history was available. In five index cases (7, 9, 10, 14 and 16) and seven family members, extra colonic features such as duodenal adenomas, FGP and/or epidermoid cysts were diagnosed. Other FAP-related manifestations in family members included a deletion carrier with hepatoblastoma (case 4) and a deletion carrier with duodenal carcinoma (case 10). One index patient (case 19) developed breast cancer twice at age 40 and 47 years, histologically defined as an adenocarcinoma (no further details could be retrieved) and a ductal adenocarcinoma, respectively, and a small-cell lung cancer at age 51. None of the families showed developmental disorders, which might be an indication of a deletion extending beyond the APC gene, although in case 13, two family members with an exons 9–15 deletion had a learning disability and one also had epilepsy.
Deletion of exons 7–13 is an in-frame deletion, because this region contains an exact threefold of nucleotides (corresponding with the codons), deletions of exons 4–5, 11 and 14 are out of frame. The in- or out-of-frame status in deletions including the promoter region and/or exon 15 cannot be predicted because it is not known to what extend these deletions reach.
Discussion
We identified 19 large APC deletions in a MPLA screening of 296 polyposis patients in whom no APC and MUTYH mutations were detected: seven whole gene deletions, nine deletions of two or more exons and three single exon deletions. Most cases had classic polyposis, none could be classified as severe, and three families had AFAP. Remarkably, the 7–13 deletion found in two AFAP families is an in-frame deletion, and we therefore studied the literature to determine possible genotype–phenotype correlations in APC deletion carriers.
Up to August 2006, a total of 89 submicroscopic APC gene deletions, including this series of patients, were reported using MLPA, PCR and real-time PCR, quantitative PCR, polymorphic markers, and cDNA analysis.14, 15, 16, 17, 22, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33 The reported deletions are summarized in Figure 2 and clinical details are given in Table 2. Not in all index patients’ clinical data were available, in 39 whole and 33 partial APC gene deletion cases, data were available for the presence or absence of CRC and extracolonic features. The age of diagnosis was reported in 27 cases with whole gene deletions.
Exon deletions reported in our series and by previous studies.14, 15, 16, 17, 22, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33 The deletions found in this study are given between brackets. Deletions due to proven splice site mutations were not included. The exact breakpoint (in introns) of most deletions was not described exactly; the exonic deletions are shown here.
Whole gene deletions
Including our cases, a total of 47 whole gene deletions have been reported in the literature. In most cases, the deletion included the promoter region. The mean age at diagnosis (as reported in 27 cases) was 32 years, ranging from 14 to 65 years, which is comparable to that reported by Friedl et al2 in classic FAP patients with APC mutations other than deletions (mean age: 30 years). Most cases with whole gene deletions can be classified as classic rather than severe, since the number of polyps in such carriers varies between ‘multiple’ and 2000 or profuse/numerous. Only one case with a whole gene deletion displayed a clearly attenuated phenotype,32 ie, a proband with less then 70 polyps and CRC at an age of 65 years old. In six further whole gene deletion cases, the number of polyps was described as ‘multiple’, which is generally reserved for a polyp count of less than 100, although patients were classified by the authors as ‘typical’ and ‘FAP’, probably because of their young age at presentation.14, 17
Partial deletions
Including our cases, a total of 42 partial APC gene deletions have been reported in the literature. The most frequent deletions were exons 14 (n=6), 14–15 (n=4), 15 (n=4), 8–15 (n=4) and 9–15 (n=4). Most cases with partial APC deletions were classified as classic FAP.
We found three index cases with an attenuated phenotype (8, 11 and 12), two of which had a previously unreported exons 7–13 deletion and one a 1–5 deletion. One obviously attenuated FAP case was reported by Su et al30 whose patient had an exon 15 deletion, 70–90 polyps at age of 42 years, and no other family members diagnosed with polyps before the age of 30.
Another deletion of exons 1–5 was reported by Aretz et al14 in a patient with multiple colorectal polyps after the age of 31. Family members of this patient had a classic FAP phenotype, and the family was therefore classified as classic according to our definition. However, there were other index patients reported with deletions in exons 1–5 who did not show an attenuated phenotype: case (9) in our study had an exons 4–5 deletion but showed a classic FAP phenotype. A family reported by Aretz et al14 with a single exon 1 deletion, including the promoter region, was also classified as classic. Renkonen et al33 described a classic FAP family with an exon 4 deletion with an index patient who had 300 polyps at the of age 69 years.
The in- or out–of-frame status of the APC deletion could be relevant for the phenotype of APC deletion patients, as this is a well-known phenomenon in Duchenne and Becker dystrophies.34 If the deletion is in-frame, a shorter but possibly functional APC product can be produced. The exons 7–13 deletion is an in-frame deletion, because this region contains an exact threefold of nucleotides (corresponding with the codons), possibly thereby explaining the milder phenotype. The in- or out-of-frame status in deletions including the promoter region and/or exon 15 could not be predicted. Without these regions, whether or not in- or out-of-frame, it is very unlikely that a functional APC product will be made. The promoter region is needed for starting transcription and exon 15 contains seven 20 amino acid repeats and three 15 amino acid repeats that are essential for regulating the β-catenin turnover and localization.35, 36 In our two AFAP families with a 7–13 deletion, the deletion was in frame. All the other partial deletions (without the promoter region or exon 15) reported here and in the literature were out frame.
The diversity of the reported deletions does not indicate the presence of ‘hot spot’ breakpoints. We also found that the size of the exon 14 deletion in our study was larger than that reported by Aretz et al14 (2.6 versus 2.0 kb), indicating at least one different breakpoint. More breakpoints should be expected in large introns, but we did not find any relation between the length of the intron (ranging from 681 to 14 111 nucleotides (http://www.ensembl.org/, Gene ID ENSG00000134982) and the number of breakpoints (P=0.4 for trend, Pearson's R). For this analysis, we included only studies that had looked for deletions in all the exons separately using MLPA.14, 17, 24, 28, 29, 33
Colorectal Cancer
Almost half of our patients developed colon cancer (44%, 8/18), one patient with a whole gene deletion had two synchronous carcinomas, and in one patient no clinical data were available. Five of the nine carcinomas were located in the distal part of the colon (55%).
Combining our results with those from previous studies (see Table 2), we found no significant difference between partial and whole gene APC deletions for the occurrence of CRC. In 39 whole gene deletion cases with detailed clinical information, nine CRCs were found (23%) with a mean age of 40 years (range: 31–65). In 33 cases with one or more exon deletions – for whom detailed clinical information was reported – 8 (24%) developed colon cancers with a mean age of 29 years (range: 29–45). The actual risk for developing CRC is most probably higher than this 23–24%, since a large number of cases will probably have undergone colectomy at a younger age (ie <30 years) to prevent the development of CRCs.
Extracolonic manifestations
We saw no essential difference between partial and whole gene APC deletions for the occurrence of extracolonic manifestations when we combined our results with those of previous studies (a total of 39 patients with whole APC gene deletions and 33 patients with partial exon deletions had available clinical data). Fourteen out of these 39 index cases (36%) had one or more extracolonic features, that is duodenal adenomas (n=6), duodenal cancer (n=1), FGP (n=5), CHRPE (n=3), desmoids (n=4), osteomas (n=5) and epidermoid cysts (n=2). In 33 index cases with one or more exon deletions, nine (27%) developed one or more extracolonic features: duodenal adenomas (n=7), duodenal cancer (n=1), FGP (n=5), stomach polyps (n=1), CHRPE (n=1), desmoids (n=1) and epidermoid cysts (n=2).
Consequences for surveillance
APC deletions show large intrafamilial as well as interfamilial variation in clinical phenotype, suggesting environmental and/or other genetic factors have a modifying influence. Consequently, advice for screening should not be based only on the extent and/or location of the mutation. As the mean age and range of CRC in patients with whole gene deletions is the same as for classic FAP patients, we advise screening according to FAP guidelines,37 ie endoscopic colorectal screening every 2 years starting from age 10 to 12 years, and endoscopic screening of the upper gastrointestinal tract every 1–5 years from age 25 to 30 years, depending on the findings. In those families with two or more members with an apparently attenuated phenotype, screening should start in the late teens.38, 39 If there are more polyps than can be removed endoscopically, we advise performing a colectomy with ileorectal anastomosis, or a proctocolectomy with an ileoanal pouch anastomosis and follow-up screening of the rectum.
Conclusion
Using MLPA, we identified partial and total APC gene deletions in 6% (19/296) of patients who had previously undetected MUTYH and APC mutations; this represents 8% (19/242) of all the APC mutations found in our cohort of 599 polyposis patients. The majority of deletion patients displayed a classic polyposis phenotype, although we also identified three families with APC deletions (16%, 3/19) with a clearly attenuated FAP phenotype. We therefore recommend that MLPA should be used as a standard procedure for identifying APC mutations in polyposis patients with classic FAP as well as those with attenuated FAP.
References
Bisgaard ML, Fenger K, Bulow S, Niebuhr E, Mohr J : Familial adenomatous polyposis (FAP): frequency, penetrance, and mutation rate. Hum Mutat 1994; 3: 121–125.
Friedl W, Caspari R, Sengteller M et al: Can APC mutation analysis contribute to therapeutic decisions in familial adenomatous polyposis? Experience from 680 FAP families. Gut 2001; 48: 515–521.
Knudsen AL, Bisgaard ML, Bulow S : Attenuated familial adenomatous polyposis (AFAP). A review of the literature. Fam Cancer 2003; 2: 43–55.
Al Tassan N, Chmiel NH, Maynard J et al: Inherited variants of MYH associated with somatic G:C–>T:A mutations in colorectal tumors. Nat Genet 2002; 30: 227–232.
Nielsen M, Franken PF, Reinards TH et al: Multiplicity in polyp count and extracolonic manifestations in 40 Dutch patients with MYH associated polyposis coli (MAP). J Med Genet 2005; 42: e54.
Sampson JR, Dolwani S, Jones S et al: Autosomal recessive colorectal adenomatous polyposis due to inherited mutations of MYH. Lancet 2003; 362: 39–41.
Aretz S, Uhlhaas S, Goergens H et al: MUTYH-associated polyposis: 70 of 71 patients with biallelic mutations present with an attenuated or atypical phenotype. Int J Cancer 2006; 119: 807–814.
Kinzler KW, Nilbert MC, Su LK et al: Identification of FAP locus genes from chromosome 5q21. Science 1991; 253: 661–665.
van der Luijt RB, Khan PM, Vasen HF et al: Molecular analysis of the APC gene in 105 Dutch kindreds with familial adenomatous polyposis: 67 germline mutations identified by DGGE, PTT, and southern analysis. Hum Mutat 1997; 9: 7–16.
Powell SM, Petersen GM, Krush AJ et al: Molecular diagnosis of familial adenomatous polyposis. New Engl J Med 1993; 329: 1982–1987.
Nugent KP, Phillips RK, Hodgson SV et al: Phenotypic expression in familial adenomatous polyposis: partial prediction by mutation analysis. Gut 1994; 35: 1622–1623.
Spirio L, Otterud B, Stauffer D et al: Linkage of a variant or attenuated form of adenomatous polyposis coli to the adenomatous polyposis coli (APC) locus. Am J Hum Genet 1992; 51: 92–100.
Soravia C, Berk T, Madlensky L et al: Genotype–phenotype correlations in attenuated adenomatous polyposis coli. Am J Hum Genet 1998; 62: 1290–1301.
Aretz S, Stienen D, Uhlhaas S et al: Large submicroscopic genomic APC deletions are a common cause of typical familial adenomatous polyposis. J Med Genet 2005; 42: 185–192.
De Rosa M, Scarano MI, Panariello L et al: Three submicroscopic deletions at the APC locus and their rapid detection by quantitative-PCR analysis. Eur J Hum Genet 1999; 7: 695–703.
Sieber OM, Lamlum H, Crabtree MD et al: Whole-gene APC deletions cause classical familial adenomatous polyposis, but not attenuated polyposis or ‘multiple’ colorectal adenomas. Proc Natl Acad Sci USA 2002; 99: 2954–2958.
Bunyan DJ, Eccles DM, Sillibourne J et al: Dosage analysis of cancer predisposition genes by multiplex ligation-dependent probe amplification. Br J Cancer 2004; 91: 1155–1159.
Petrij-Bosch A, Peelen T, van Vliet M et al: BRCA1 genomic deletions are major founder mutations in Dutch breast cancer patients. Nat Genet 1997; 17: 341–345.
Grabowski M, Mueller-Koch Y, Grasbon-Frodl E et al: Deletions account for 17% of pathogenic germline alterations in MLH1 and MSH2 in hereditary nonpolyposis colorectal cancer (HNPCC) families. Genet Test 2005; 9: 138–146.
Wijnen J, van der KH, Vasen H et al: MSH2 genomic deletions are a frequent cause of HNPCC. Nat Genet 1998; 20: 326–328.
McCart A, Latchford A, Volikos E, Rowan A, Tomlinson I, Silver A : A novel exon duplication event leading to a truncating germ-line mutation of the APC gene in a familial adenomatous polyposis family. Fam Cancer 2006; 5: 205–208.
Takahashi M, Kikuchi M, Ohkura N et al: Detection of APC gene deletion by double competitive polymerase chain reaction in patients with familial adenomatous polyposis. Int J Oncol 2006; 29: 413–421.
Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, Diepvens F, Pals G : Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 2002; 30: e57.
Michils G, Tejpar S, Thoelen R et al: Large deletions of the APC gene in 15% of mutation-negative patients with classical polyposis (FAP): a Belgian study. Hum Mutat 2005; 25: 125–134.
Cao X, Eu KW, Seow-Choen F, Zao Y, Cheah PY : APC mutation and phenotypic spectrum of Singapore familial adenomatous polyposis patients. Eur J Hum Genet 2000. Vol 8: 42–48.
Flintoff KJ, Sheridan E, Turner G, Chu CE, Taylor GR : Submicroscopic deletions of the APC gene: a frequent cause of familial adenomatous polyposis that may be overlooked by conventional mutation scanning. J Med Genet 2001; 38: 129–132.
Gismondi V, Bafico A, Biticchi R et al: 310 Basepair APC deletion with duplication of breakpoint (439ins15del310) in an Italian polyposis patient. Hum Mutat 1998; (Suppl 1): S220–S222.
Meuller J, Kanter-Smoler G, Nygren AO et al: Identification of genomic deletions of the APC gene in familial adenomatous polyposis by two independent quantitative techniques. Genet Test 2004; 8: 248–256.
Mihalatos M, Apessos A, Dauwerse H et al: Rare mutations predisposing to familial adenomatous polyposis in Greek FAP patients. BMC Cancer 2005; 5: 40.
Su LK, Kohlmann W, Ward PA, Lynch PM : Different familial adenomatous polyposis phenotypes resulting from deletions of the entire APC exon 15. Hum Genet 2002; 111: 88–95.
Su LK, Steinbach G, Sawyer JC, Hindi M, Ward PA, Lynch PM : Genomic rearrangements of the APC tumor-suppressor gene in familial adenomatous polyposis. Hum Genet 2000; 106: 101–107.
Venesio T, Balsamo A, Rondo-Spaudo M, Varesco L, Risio M, Ranzani GN : APC haploinsufficiency, but not CTNNB1 or CDH1 gene mutations, accounts for a fraction of familial adenomatous polyposis patients without APC truncating mutations. Lab Invest 2003; 83: 1859–1866.
Renkonen ET, Nieminen P, Abdel-Rahman WM et al: Adenomatous polyposis families that screen APC mutation-negative by conventional methods are genetically heterogeneous. J Clin Oncol 2005; 23: 5651–5659.
Monaco AP, Bertelson CJ, Liechti-Gallati S, Moser H, Kunkel LM : An explanation for the phenotypic differences between patients bearing partial deletions of the DMD locus. Genomics 1988; 2: 90–95.
Liu J, Xing Y, Hinds TR, Zheng J, Xu W : The third 20 amino acid repeat is the tightest binding site of APC for beta-catenin. J Mol Biol 2006; 360: 133–144.
Rubinfeld B, Souza B, Albert I et al: Association of the APC gene product with beta-catenin. Science 1993; 262: 1731–1734.
Vasen B : Guidelines for the surveillance and management of familial adenomatous polyposis (FAP): a world wide survey among 41 registries. Colorectal Dis 2003; 1: 214.
Nielsen M, Hes FJ, Nagengast FM et al: Germline mutations in APC and MUTYH are responsible for the majority of families with attenuated familial adenomatous polyposis (AFAP). Clin Genet 2007; 71: 427–433.
Burt RW, Leppert MF, Slattery ML et al: Genetic testing and phenotype in a large kindred with attenuated familial adenomatous polyposis. Gastroenterology 2004; 127: 444–451.
Cao Y, Pieretti M, Marshall J, Khattar NH, Chen B, Kam-Morgan L, Lynch H : Challenge in the differentiation between attenuated familial adenomatous polyposis and hereditary nonpolyposis colorectal cancer: case report with review of the literature. Am J Gastroenterol 2002; 97: 1822–1827.
Acknowledgements
MN received a grant from the Dutch Digestive Diseases Foundation (Grant no. MWO 0355). We thank IA Tomlinson, D Eccles, and S Aretz for providing additional clinical information and Jackie Senior for critically reviewing the manuscript. The author(s) declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nielsen, M., Bik, E., Hes, F. et al. Genotype–phenotype correlations in 19 Dutch cases with APC gene deletions and a literature review. Eur J Hum Genet 15, 1034–1042 (2007). https://doi.org/10.1038/sj.ejhg.5201871
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201871
Keywords
This article is cited by
-
Current status of prophylactic surgical treatment for familial adenomatous polyposis in Japan
Surgery Today (2017)
-
Association between the age and the development of colorectal cancer in patients with familial adenomatous polyposis: a multi-institutional study
Surgery Today (2017)
-
ACG Clinical Guideline: Genetic Testing and Management of Hereditary Gastrointestinal Cancer Syndromes
American Journal of Gastroenterology (2015)
-
APC rearrangements in familial adenomatous polyposis: heterogeneity of deletion lengths and breakpoint sequences underlies similar phenotypes
Familial Cancer (2015)
-
APC Somatic Mosaicism in a Patient with Gardner Syndrome Carrying the E1573X Mutation: Report of a Case
Diseases of the Colon & Rectum (2009)