Abstract
Autosomal dominant cone dystrophy (CORD5) (MIM 600977) is a rare disease predominantly affecting cone photoreceptors. Here we refine the CORD5 locus previously mapped to 17p13 from 27 to 14.3 cM and identified a missense mutation, Q626H in the phosphatidylinositol transfer (PIT) membrane-associated protein (PITPNM3) (MIM 608921) in two Swedish families. PITPNM3, known as a human homologue of the Drosophila retinal degeneration B (rdgB), lacks the N-terminal PIT domain needed for transport of phospholipids, renewal of photoreceptors membrane and providing the electroretinogram (ERG) response to light. In our study, the mutation causing CORD5 is located in the C-terminal region interacting with a member of nonreceptor protein tyrosine kinases, PYK2. Our finding on the first mutation in the human homologue of Drosophila rdgB indicates novel pathways and a potential important role of the PITPNM3 in mammalian phototransduction.
Similar content being viewed by others
Introduction
Progressive cone or cone-rod dystrophies (CORDs) are characterized by a defective cone function demonstrated by abnormalities in cone-mediated electroretinogram (ERG) components. The presenting symptoms are defective colour vision, impaired central visual acuity and sensitivity to light.1, 2 The inheritance patterns for CORDs are represented by autosomal-dominant, autosomal-recessive and X-linked forms.3 The classification of CORDs is complicated due to variable dysfunction of the rods. The preservation of rod function can differ both between and within families and is dependent on the disease causing mutation within a gene.2, 4
A variant of autosomal-dominant cone dystrophy has been mapped to 17p12–p13 in a Swedish family5 (CORD5, MIM 600977). Two other reports, one from the United Kingdom and one from the United States also showed linkage of dominant cone dystrophy to chromosome 17p.6, 7 The disorder in the British family, designated as CORD6 (MIM 601777) was found to be caused by mutations in the GUCY2D gene.8 Mutations within the GUCY2D gene also cause Leber congenital amaurosis and autosomal recessive retinitis pigmentosa.9 In the USA family, the disease mapped to 17p12–p13 was later reported to be caused by the same mutation as the in British CORD6 family, and therefore it was concluded that CORD5 and CORD6 is the same disease.6, 10 However, in CORD5 patients of Swedish origin, there were no indications of mutations in the GUCY2D gene analysed by single strand conformational polymorphism (SSCP).2
Besides GUCY2D, another gene, AIPL1 (MIM 604392) mapped to 17p13 is associated with autosomal-dominant CORD.11 Moreover, autosomal-dominant central areolar choroidal dystrophy (CACD) characterized by progressive profound loss of vision in patients during middle age has been mapped to 17p13 in an Irish family. To date, however, the disease causing gene has not been identified.12, 13 Other variants of autosomal-dominant CORDs are caused by mutations in RIM1 (6q14) (MIM 606629), peripherin/RDS (MIM 179605) (6p21.2), GUCA1A (6p21.1) (MIM 600364) and CRX (19q13) (MIM 602225) genes.14, 15, 16, 17
The CORD5 locus in families of Swedish origin was initially mapped to a 27 cM region on chromosome 17p12–p13; however, the gene causing the disease was not identified.5 Thus, the aim of the present study was identification of the genetic defect in CORD5 patients of Swedish origin.
Materials and methods
Patients and clinical examination
To identify a gene causing CORD5 we analysed two multigeneration Swedish families (151 and 152) originating from the same geographical area in Northern Sweden. Pedigrees of both families are presented in Figure 3. Family 151, reported previously,5 is a five-generation family with a total of 48 individuals of which 18 are affected. Family 152 is a seven-generation family of 32 individuals with 15 affected and two individuals with unknown disease status. Informed consent was obtained from all individuals participating in the study and the research following to the tenets of Declaration of Helsinki was approved by the Ethics Committee of University of Umeå. All individuals classified as unaffected were symptomless, and had normal visual acuity and normal colour vision. Patients with a history of defective colour vision, low visual acuity and signs of macular degeneration were assigned as affected. Affected members in family 151 have been subject for a thorough evaluation. In family 152 the corresponding clinical examinations in progress, however have not yet been completed. A full ophthalmologic examination included assessment of visual acuity (VA) and evaluation of the anterior segment and the fundus. Visual fields were tested in a Goldmann perimeter using standard objects. In selected cases, a more detailed investigation was performed. The course of dark adaptation was determined using a Goldmann-Weekers adaptometer. Colour vision was tested with pseudoisochromatic plates (Ishihara and Dvorine). Electrooculography (EOG), and full-field, single flash and flicker ERGs were recorded using UTAS-E 2000 and UTAS-E 3000-mf LKC Technologies Inc.
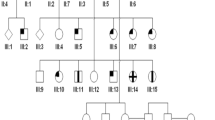
CORD5 haplotypes in families 151 and 152. Filled symbols indicate affected individuals, whereas empty symbols indicate unaffected. Symbols with ‘?’ had unknown disease status. Only haplotypes shared by affected individuals in both families are boxed. Individual IV:10 in family 151 demonstrated a mutation in the sequence of marker D17S720 resulting in allele size 212.
Genotyping, linkage analysis, and mutational screening
DNA from 80 individuals was extracted from peripheral blood lymphocytes according to standard protocols reported by Balciuniene et al.5 Twelve microsatellite markers included in the Rutgers Combined Linkage-Physical Map were used for genotyping. The markers mapped to 17p13 in the proximity to D17S938 were situated approximately 2 cM apart. The GDB Human Genome Database was used for information about microsatellite markers and their primer sequences. PCR mixes were prepared in 7.5 μl reactions using 0.3 U of AmpliTaqGold (Applied Biosystems, Foster City, CA, USA), 0.75 μl of GeneAmpPCRbuffer II, 0.075 mM of dNTP, 0.75 mM of MgCl2, and 0.1 μM of primers. Polymerase chain reaction (PCR) was performed with a temperature profile of 95°C for 10 min, 94°C for 15 s, 55°C for 15 s, 72°C for 30 s for 10 cycles, 89°C for 15 s, 55°C for 15 s and 72°C for 30 s for 20 cycles, and final extension at 72°C for 10 min. Fragment analysis of the samples was performed on a 3730xl DNA analyzer (Applied Biosystems), and the collected raw data were analysed with ABI Prism GeneMapper Software v3.0 (Applied Biosystems).
Two-point linkage analyses were performed using the FASTLINK implementation of the LINKAGE program package.18 CORD5 was analysed as an autosomal-dominant trait, with a disease allele frequency of 0.01. Five age-dependent liability classes with the penetrance set to 0 at <10 years, 0.15 between 10 and 20 years, 0.47 between 21 and 30 years and 0.95 at ages >40 years were used. The liability classes were the same as in the mapping study of CORD5.5 Frequency of phenocopies was set to 0.01. Set up of allele and phenocopies frequencies was the same as for complex traits, which allowed increasing power of linkage analysis.
For bidirectional sequencing of phosphatidylinositol transfer (PIT) membrane-associated protein (PITPNM3) (MIM 608921), coding exons and adjacent intronic sequences were amplified from genomic DNA of two affected patients from each family and one unaffected individual. Primer pairs designed with Primer3 software are listed in Table 1. PCR amplification of the 19 exons was performed in 25 μl reactions with 0.5 U of AmpliTaq Gold (Applied Biosystems), 2.5 μl of PCR buffer II × 10, 1.5 mM of MgCl2, 0.1 mM of dNTP, 0.4 μ M of primers, and 50 ng of genomic DNA. PCR cycling consisted of an initial denaturing step at 94°C for 5 min; then 30 cycles of denaturing at 94°C for 1 min, annealing at temperature dependent on primer sequence (55°C) for 1 min, and extension at 72°C for 1 min; and a final elongation step at 72°C for 7 min. PCR products were purified using MicroSpin columns (Amersham Biosciences, Piscataway, NJ, USA). The sequencing reactions were performed using Big Dye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) in a final reaction of 10 μl. The products of sequencing reactions were run on a 3730xl DNA analyzer (Applied Biosystems). Sequences aligned in the Seqman program of the DNASTAR package were compared with a reference genomic sequence of the PITPNM3 gene available on the Ensembl website (ENSG00000091622).
Results and discussion
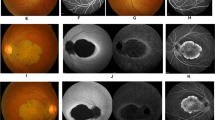
Two Swedish families (151 and 152), originating from the same geographical area in Northern Sweden were included in this study. The majority of patients in family 151 (pedigree shown in Figure 3) had subnormal visual acuity and light sensitivity from childhood. Signs of macular degeneration were observed early as well. There was a progressive decrease of VA leading to legal blindness in early adulthood. Electrophysiological testing showed a progressive loss of photoreceptor function restricted to the cones. The macular regions presented a normal appearance at early stages of the disease. In the young patient (V:2, at 15 years of age, Figure 1b) a bull's-eye maculopathy could be seen. In more advanced cases, findings varied from pigment mottling to pronounced central choroidal atrophy (IV:23 and III:24 at ages 33 and 61 years respectively, Figure 1c and d). A pronounced macular involvement was associated with central scotomas at visual field testing. There was no peripheral restriction of the visual field and patients did not have any history of nyctalopia. Photopic ERGs representing cone function were greatly diminished or absent, whereas scotopic ERGs showed normal amplitudes (Figure 1a–d). The EOG was normal. Dark adaptation curves showed absence of a normal cone segment, but the final rod dark adaptation threshold was not elevated.
Fundus photographs and ERG recordings. A healthy 35-year-old individual (a) and CORD5 patients from family 151. (b) A 15-year-old boy (V:2). (c) His 33-year-old mother (IV:23). (d) His 61-year-old grandmother (III:24). The photographs show a progression of maculopathy with increasing age from bull's-eye maculopathy to an extended macular atrophy. Scotopic rod responses diminish with increasing age, but are within the normal range. Photopic cone ERGs are greatly diminished or absent in all patients. The amplitudes decrease with increasing age and in the oldest patient there are no registerable amplitudes. The implicit times are prolonged with increasing age as well. The same is valid for the registrations of the 30-Hz flicker ERGs.
A milder phenotype was observed among clinically examined individuals from family 152 (pedigree shown in Figure 3). A female (V:15), with a visually affected mother (IV:4), had first complaint of impaired vision at age about 45 years. Examination showed defective colour vision but a normal full-field cone response at electrophysiological testing. Her daughter (VI:11) presented with low visual acuity, defective colour vision and light sensitivity at the age of 30, and the ERG was abnormal in accordance with cone dystrophy. A more detailed phenotype description presented in these families will be published elsewhere.
For fine mapping of the CORD5 locus 12 microsatellite markers in close proximity of the D17S938 on 17p13 were chosen (Figure 2a, Table 2). Significant logarithmic odds (LOD) scores at θ=0 were revealed for markers D17S945 to D17S1828 with a maximum of 12.67 at the marker D17S938 (Table 2). The reconstructed haplotypes in both families (Figure 3) confirmed segregation of CORD5 with markers D17S678, D17S938, D17S1881, D17S720 and D17S1844. In family 151, the interval containing the disease gene was limited by one recombination event for D17S945 in three affected individuals (III:29, IV:27 and V:4). In family 152, recombination events at D17S1828 and D17S1854 were detected (V:17, VI:14 and VI:10). Thus, CORD5 locus was narrowed down from 26.9 to 14.3 cM including flanking markers. In the refined region at least three genes of interest were present: AIPL1, an aryl-hydrocarbon interacting protein-like 1; GUCY2D, guanylate cyclase 2D, both known to be mutated in cone rod dystrophy and Leber congenital amaurosis; and a membrane associated phosphatidylinositol transfer protein 3, PITPNM3, a human homologue of the Drosophila retinal degeneration B (rdgB). AIPL1 and GUCY2D were excluded as a cause of CORD5 by SSCP2 and sequence analysis (data not shown).
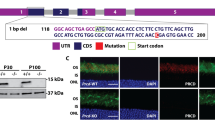
Fine mapping of the CORD5 disease, genomic structure of the PITPNM3 gene, detection of the Q626H mutation and segregation analysis in family 151. (a) Schematic representation of the genomic region in proximity to the D17S938 marker. D17S678, D17S938, D17S1881, D17S720 and D17S1844 segregated with CORD5. PITPNM3 consists of 20 exons, where exon 2 (shown in red) encodes a Ca2+-binding domain, six transmembrane domains (shown as black lines) spread over exons 6, 7, 10, 11, 12 and exons 14–20 encoding PYK2-binding domain (shown in green). The PIT domain characteristic for this family is missing in PITPNM3. Q626H in exon 14 is marked with an asterisk. (b) Sequence analysis demonstrating wild-type sequence and presence of the Q626H mutation in an affected individual. Genomic analysis of PITPNM3 was performed by bidirectional sequencing as described in Materials and methods. (c) Pedigree of family 151 and segregation analysis of CORD5 and the Q626H mutation in the PITPNM3 gene. Filled symbols indicate affected individuals, whereas empty symbols indicate unaffected. Screening for the Q626H mutation was carried out by PCR-RFLP analysis of the PITPNM3 exon 14. (d) Partial sequence alignment of PYK2-binding domain between mammalian PITPNM3. The position 626 where the Q626H mutation occurs is indicated in bold.
PITPNM3 contains 20 exons spanning approximately 101 kb of genomic sequence and encodes a protein belonging to phosphatidyl-inosytol transfer protein (PITP) family. The human rdgB homologue, PITPNM3 also called Nir1 was cloned in 1999. Based on its chromosomal localisation together with linkage data of several retinal disorders to the same locus, the PITPNM3 was proposed to be a candidate gene causing retinal degenerations.19, 20 Taking into account our own linkage data we decided to screen the PITPNM3 for possible mutations. Sequence analysis of 19 exons of PITPNM3 revealed a transversion c.1878G>C (NM_031220) in exon 14 resulting in substitution of a glutamine at position 626 by a histidine (p.Q626H) (Figure 2b). Q626H mutation introduces a restriction site for MaeII endonuclease which allowed screening for the Q626H mutation by PCR-RFLP analysis. A 268 bp long PCR products amplified according to the conditions described in ‘Materials and methods’ were digested with MaeII. Products of digestion, fragments 157 and 111 bp were resolved on 2% agarose gel (SeaKem, ME Agarose) and visualized by GelStar® Nuclear Acid Staining (Cambrex, Karlskoga, Sweden). Consistent segregation of the Q626H in all affected individuals in family 151 (Figure 2c) was shown.
In family 152 three individuals VI:9, VI:12 and VI:16 at age of 9, 18 and 28 years, had unknown disease history at the time of blood collection . These individuals shared the same haplotype as affected individuals (Figure 3) and also had the Q626H mutation. Case VI:16 examined at the age of 29 showed defective colour vision, low visual acuity and abnormal cone response when tested with ERG. Patients VI:9 and VI:12 have not been fully clinically evaluated.
Shared haplotypes and presence of the Q626H mutation in both families indicate that these families have a common descent (Figure 3).
The Q626H mutation was absent on 322 control chromosomes of ethnically matched healthy individuals and in 140 individuals affected with autosomal-dominant or recessive forms of retinitis pigmentosa. Residue 626 in PITPNM3 is located in a protein tyrosine kinase PYK2-binding domain,19 which is evolutionary conserved in mammals (Figure 2d). To predict significance of Q626H substitution we used web-based analysis programs. According to ‘SNPper – aminoacid variation’ (http://snpper.chip.org/bio/show-amino) a change in the chemical structure due to replacement of an uncharged polar glutamine with a positive polar histidine is indicated as major. Furthermore, glutamine and histidine have different alpha-helix propensity and this difference can have effect on secondary structure of PITPNM3. Conformational change can in its turn influence binding to PYK2. We also used SIFT (Sorting Intolerant from Tolerant (http://blocks.fhcrc.org/sift-bin) as a measure of sequence conservation. The Q626H change was predicted to be deleterious and affect protein function (probability 0.03) using SWISS-PROT database, however searching through TREMBLE and NCBI made this change tolerant due to presence of histidine in zebrafish and mouse Pitpnm2. Finally, PolyPhen (Polymorphism Phenotyping) predicting possible impact of an amino acid substitution on the structure and function of a human protein (http://coot.embl.de/PolyPhen/) failed to reveal significance of Q626H substitution. Despite of the latest predictions, presence of the Q626H mutation in all CORD5 individuals of Swedish origin, it's absence on healthy chromosomes and the fact that PITPNM3 is a Drosophila rdgB homologue causing light-induced retinal degeneration and severely impaired ERG21 indicate a potential pathogenic role of the Q626H substitution.
In humans PITPNM3 is shown to be expressed in the brain, spleen and ovary,19 although PITPNM3 expression based on ESTs counting is more broad including adrenal gland, cervix, colon, connective tissue, eye, kidney, lymph, pancreas, lung, larynx, mouth, muscle and testis (http://www.ncbi.nlm.nih.gov/UniGene). Interestingly, the zebrafish homologue, pl-RdgB is mainly expressed in inner segments of cone photoreceptors,22 whereas in rat retina PITPNM3 is expressed through all cell layers including the inner segment and the outer plexiform layer containing photoreceptor terminals but predominant expression was later determined in Muller cells.23
Members of the PITP-family contain a phosphatidylinositol transfer protein domain (PIT), an acidic region/Ca+2 binding domain, six transmembrane domains, the DDHD and a C-terminal domain interacting with PYK2.19, 24 The PITPs participate in multiple processes such as phospholipase C-mediated inositol signaling, ATP-dependent Ca+2-activated secretion, lipid metabolism, trafficking from Golgi-membranes and exocytosis.24 An exclusive importance of a PIT domain needed for phospholipids transport and renewal of photoreceptor membrane was observed in rdgB mutant flies with light-dependent retinal degeneration which was rescued by transfer of the PIT-domain.25 Transgenic expression of zebrafish pl-RdgB lacking a PIT domain in Drosophila rdgB2 null mutant improved photoreceptor survival but did not show any effect on ERG22 suggesting that PIT-domain is necessary for light response and function of other protein domains prevents photoreceptors degeneration.
The signalling pathway for mammalian PITPNM3 is still not completely understood. Genetic studies on Drosophila mutants provided evidence that rdgB functions downstream the protein kinase C, rhodopsin and phospholipase C, since PKC activation results in retinal degeneration in rdgB mutants even in darkness25, 26, 27 and mutations in rhodopsin and phospholipase C prevent light dependent retinal degeneration in rdgB mutants.21
Recently it was shown that the C-terminal region of the PITPNM3 is necessary for the interaction with protein tyrosine kinase PYK2.19 Activation of PYK2 results in phosphorylation of PITPNM3 indicating that it is a substrate for PYK2 although it is not known if this interaction is phosphorylation dependent.19
In conclusion, here we show for the first time that the Q626H mutation in PITPNM3 mediates CORD5. At this point, we can only speculate the potential functional role of this mutation. One obvious expectation might be that the mutation abolishes or modifies the interaction with PYK2 in humans. PITPNM3 might be involved in several more complex pathways affecting photoreceptor survival and light induced response. Further studies on model organisms carrying the Q626H mutation will clarify the mechanisms by which mutations in the PYK2-binding domain result in defective vision. Determination of general importance of PITPNM3 in the development of other CORDs also requires further screening for additional mutations in both familiar and isolated cases. Clinical examination of our patients demonstrated impaired function of cone cells and normal response of rods, which raises a question of disease designation. Historically, the disease was called CORD5, as there was already CORD1–4. At that time it was impossible to predict if rods function would later be affected. Detection of new mutations in PITPNM3 followed by studies of phenotype-genotype correlations might correct the name of the disease.
In summary, this study adds one more gene on 17p, which causes retinal degeneration. We provide evidence that CORD5 in Swedish patients is a distinct clinical entity and describe the first disease causing mutation within the PYK2–binding domain in the PITP family.
References
Small KW, Gehrs K : Clinical study of a large family with autosomal dominant progressive cone degeneration. Am J Ophthal 1996; 121: 1–12.
van Ghelue M, Eriksen HL, Ponjavic V et al: Autosomal dominant cone-rod dystrophy due to a missense mutation (R838C) in retinal guanylate cyclase gene (RETGC-1) is associated with considerable variation. Ophthalmol Genet 2000; 21: 197–209.
Michaelides M, Holder GE, Hunt DM, Fitzke FW, Bird AC, Moore AT : A detailed study of the phenotype of an autosomal dominant cone-rod dystrophy (CORD7) associated with mutation in the gene for RIM1. Br J Ophthalmol 2005; 89: 198–206.
Michaelides M, Holder GE, Bradshaw K, Hunt DM, Moore AT : Cone-rod dystrophy, intrafamilial variability, and incomplete penetrance associated with the R172W mutation in the peripherin/RDS gene. Ophthalmol 2005; 112: 1592–1598.
Balciuniene J, Johansson K, Sandgren O, Wachtmeister L, Holmgren G, Forsman K : A gene for autosomal dominant progressive cone dystrophy maps to chromosome 17p12–p13. Genomics 1995; 30: 281–286.
Small KW, Syrquin M, Mullen L, Gehrs K : Mapping of autosomal dominant cone degeneration to chromosome 17p. Am J Ophthalmol 1996; 121: 13–18.
Kelsell RE, Evans K, Gregory CY, Moore AT, Bird AC, Hunt DM : Localization of a gene for dominant cone-rod dystrophy (CORD6) to chromosome 17p. Hum Molec Genet 1997; 6: 597–600.
Kelsell RE, Evans K, Payne AM et al: Mutations in the retinal guanylate cyclase (RETGC-1) gene in dominant cone-rod dystrophy. Hum Molec Genet 1998; 7: 1179–1184.
Perrault I, Rozet JM, Calvas P et al: Retinal-specific guanylate cyclase gene mutations in Leber's congenital amaurosis. Nat Genet 1996; 14: 461–464.
Udar N, Yelchits S, Chalukya M et al: Identification of GUCY2D gene mutations in CORD5 families and evidence of incomplete penetrance. Hum Mutat 2003; 21: 170–171.
Sohocki MM, Perrault I, Leroy BP et al: Prevalence of AIPL1 mutations in inherited retinal degenerative disease. Mol Genet Metab 2000; 70: 142–150.
Lotery AJ, Ennis KT, Silvestri G et al: Localisation of a gene for central areolar choroidal dystrophy to chromosome 17p. Hum Mol Genet 1996; 5: 705–708.
Hughes AE, Lotery AJ, Silvestri G : Fine localisation of the gene for central areolar choroidal dystrophy on chromosome 17p. J Med Genet 1998; 9: 770–772.
Johnson S, Halford S, Morris AG et al: Genomic organisation and alternative splicing of human RIM1, a gene implicated in autosomal dominant cone-rod dystrophy (CORD7). Genomics 2003; 81: 304–314.
Nakazawa M, Kikawa E, Chida Y, Wada Y, Shiono T, Tamai M : Autosomal dominant cone-rod dystrophy associated with mutations in codon 244 (Asn244His) and codon 184 (Tyr184Ser) of the peripherin/RDS gene. Arch Ophthalmol 1996; 114: 72–78.
Downes SM, Holder GE, Fitzke FW et al: Autosomal dominant cone and cone-rod dystrophy with mutations in the guanylate cyclase activator 1A gene-encoding guanylate cyclase activating protein-1. Arch Ophthalmol 2001; 119: 96–105.
Freund CL, Gregory-Evans CY, Furukawa T et al: Cone-rod dystrophy due to mutations in a novel photoreceptor-specific homeobox gene (CRX) essential for maintenance of the photoreceptor. Cell 1997; 91: 543–553.
Weeks DE, Ott J, Lathrop GM : SLINK: a general simulation program for linkage analysis. Am J Hum Genet 1990; 47: A204 (Supplement).
Lev S, Hernandez J, Martinez R, Chen A, Plowman G, Schlessinger J : Identification of a novel family of targets of PYK2 related to Drosophila retinal degeneration B (rdgB) protein. Mol Cell Biol 1999; 19: 2278–2288.
Lev S : Molecular aspects of retinal degenerative diseases. Cell Mol Neurobiol 2001; 21: 575–589.
Harris WA, Stark WS : Hereditary retinal degeneration in Drosophila melanogaster. A mutant defect associated with the phototransduction process. J Gen Physiol 1977; 69: 261–291.
Elagin VA, Elagina RB, Doro CJ, Vihtelic TS, Hyde DR : Cloning and tissue localization of a novel zebrafish RdgB homolog that lacks a phospholipid transfer domain. Vis Neurosci 2000; 17: 303–311.
Tian D, Lev S : Cellular and developmental distribution of human homologues of the Drosophilia rdgB protein in the rat retina. Invest Ophthalmol Vis Sci 2002; 43: 1946–1953.
Lev S : The role of the Nir/rdgB protein family in membrane trafficking and cytoskeleton remodeling. Exp Cell Res 2004; 297: 1–10.
Milligan SC, Alb Jr JG, Elagina RB, Bankaitis VA, Hyde DR : The phosphatidylinositol transfer protein domain of Drosophila retinal degeneration B protein is essential for photoreceptor cell survival and recovery from light stimulation. J Cell Biol 1997; 139: 351–363.
Smith DP, Ranganathan R, Hardy RW, Marx J, Tsuchida T, Zuker CS : Photoreceptor deactivation and retinal degeneration mediated by a photoreceptor-specific protein kinase C. Science 1991; 254: 1478–1484.
Minke B, Rubinstein CT, Sahly I, Bar-Nachum S, Timberg R, Selinger Z : Phorbol ester induces photoreceptor-specific degeneration in a Drosophila mutant. Proc Natl Acad Sci USA 1990; 87: 113–117.
Acknowledgements
We thank all members of CORD5 families and acknowledge E Blennow for collecting blood samples and J Moharer for technical assistance. This study was supported by Grants from Visare Norr and University Hospital of Umeå.
Author information
Authors and Affiliations
Corresponding author
Additional information
Web resources
The accession numbers and URLs for data presented herein are as follows: Information on CORD5–entity can be found on Online Mendelian Inheritance in Man (OMIM) – http://www.ncbi.nlm.nih.gov/OMIM. Primer design was performed using program available on http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi. Genomic sequence of the PITPNM3 gene is available on http://www.ensembl.org (ENSG00000091622).
ClustalW 1.74 multiple sequence alignment program – http://www.ebi.ac.uk/clustalw/.
To sort intolerant from tolerant amino acid substitutions – http://blocks.fhcrc.org and to check possible impact of an amino acid substitution on the structure and function of a human protein was used PolyPhen analys program – http://coot.embl.de/PolyPhen/.
Rights and permissions
About this article
Cite this article
Köhn, L., Kadzhaev, K., Burstedt, M. et al. Mutation in the PYK2-binding domain of PITPNM3 causes autosomal dominant cone dystrophy (CORD5) in two Swedish families. Eur J Hum Genet 15, 664–671 (2007). https://doi.org/10.1038/sj.ejhg.5201817
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201817
Keywords
This article is cited by
-
βA3/A1-crystallin regulates apical polarity and EGFR endocytosis in retinal pigmented epithelial cells
Communications Biology (2021)
-
The clinical spectrum of inherited diseases involved in the synthesis and remodeling of complex lipids. A tentative overview
Journal of Inherited Metabolic Disease (2015)
-
Genome-wide association and linkage analyses localize a progressive retinal atrophy locus in Persian cats
Mammalian Genome (2014)
-
Novel mutations in CRB1 and ABCA4 genes cause Leber congenital amaurosis and Stargardt disease in a Swedish family
European Journal of Human Genetics (2013)
-
Breakpoint characterization of a novel ∼59 kb genomic deletion on 19q13.42 in autosomal-dominant retinitis pigmentosa with incomplete penetrance
European Journal of Human Genetics (2009)