Abstract
T helper (h) lymphocytes in pathogenic immune response at mucosal effector site play a key role in IgA nephropathy (IgAN). We evaluated the impact of some Th1/Th2/Th3/TR-type, and of monocyte/macrophage cytokines on IgAN susceptibility with a family-based association study including 53 patients, 45 complete trios, 4 incomplete trios and 36 discordant siblings. Cytokine gene polymorphisms with a potential regulatory role on their production were investigated using the family-based association test (FBAT): IFNγ intron-1 CA repeat at position 1349–1373; IL-13 −1055C/T; TGFβ +915G/C; IL-10 5′-proximal and distal microsatellites; TNFα −308G/A, −238G/A. The FBAT multi-allelic analysis showed an association between IFNγ polymorphism and susceptibility to IgAN (P=0.03). The bi-allelic analysis evidenced that the 13-CA repeat allele was preferentially transmitted to the affected individuals (P=0.006; Bonferroni P-value=0.04). The direct sequencing of IFNγ amplicons showed a strict association between the 13-CA repeat allele and the A variant of the +874T/A single nucleotide polymorphism (SNP rs2430561) directly adjacent to the 5′ end of the microsatellite. The in vitro production of IFNγ evaluated in peripheral blood mononuclear cells from 10 genotyped patients demonstrated a correlation between the +874A allele and a lower production of IFNγ (P=0.028 Mann–Whitney test). This SNP affects IFNγ production lying within a binding site for the transcription factor NF-κB. No significant difference was observed in the 15 years renal survival between IgAN patients carrying different IFNγ gene polymorphisms. This first family-based association study demonstrates that the +874A allele, strictly associated with IFNγ 13-CA repeat allele, confers susceptibility to IgAN, without influencing renal survival.
Similar content being viewed by others
Introduction
IgA nephropathy (IgAN) is a disease characterized by an abnormal immune response with an increased synthesis of deglycosylated IgA1, occurring in the blood and then depositing in the glomeruli of the kidney. Mucosal immunity is largely involved in the pathogenesis of the disease, as the exposure of the upper respiratory tract to bacteria or virus is often associated with recurrent episodes of macroscopic hematuria. A key component of the mucosal immune defense against pathogens is represented by CD4+ T lymphocytes that can differentiate into functionally distinct subsets.1, 2, 3 Whereas T-helper1 (Th1) cells secrete the cytokine interferon-γ (IFNγ) and tumor necrosis factor-β (TNF-β), Th2 cells secrete interleukin-4 (IL-4), IL-5, IL-9 and IL-13. Th3 and regulatory CD25+ CD4+ T (TR) cells produce transforming growth factor-β (TGF-β) and IL-10, respectively.4, 5 Tumor necrosis factor-α (TNF-α) produced by monocytes and macrophages is a cytokine with pleiotropic biological activity including the regulation of immune cell proliferation and differentiation.6 Cytokines act in a highly complex network in which they induce or repress their own synthesis as well as that of other cytokines and cytokine receptors. The imbalance in cytokine production drives the immune system towards a cell-mediated (Th1 type) or a humoral response (Th2 type). There is evidence for Th1/Th2 polarization of nephritogenic immune responses in IgAN, even if Th1 or Th2 predominance has not been clearly defined. The predominance of T lymphocyte subsets is the consequence of mutual Th1/Th2 cytokine antagonism. The in vitro capacity to produce different cytokines in response to mitogen stimulation has been shown to vary between individuals. Conservative mutations within coding regions, nucleotide variations within more pronounced regulatory regions (ie promoter sequences) and polymorphisms within cytokine gene introns have all been described as potential mechanisms involved. Different polymorphisms were shown to be implied in the genetic control of cytokines produced by T helper cells and monocyte/macrophages: IFNγ intron-1 microsatellite polymorphism positioned at 1339–1373; IL-13 promoter polymorphism (C – T) at position –1055; TGF-β1 polymorphism at position +915 in the signal sequence; microsatellite loci (IL-10R, IL-10G) described in the IL-10 promoter region; TNF-α gene polymorphisms at nucleotide positions −308 and −238.7, 8, 9, 10, 11, 12 We investigated the effects of these polymorphisms on the IgAN onset and progression using a family-based association study. High priority was given to IFNγ gene intron-1 polymorphism, as a previously published case–control association study demonstrated a significantly higher incidence of IFNγ 13-CA repeat allele in Japanese IgAN patients than in healthy subjects.13 The observed preferential transmission of 13-CA repeat allele to IgAN patients suggests a functional role of either the microsatellite itself, or, more probably, of a functional polymorphism in linkage disequilibrium with this allele. An in vitro increased production of IFNγ correlated with the presence of 12-CA repeat allele, which was in linkage disequilibrium with the +874T/A single nucleotide polymorphism (SNP rs2430561) within a putative NF-κB binding site.14, 15 Our study speculates also on the functional role of this polymorphism.
Patients and methods
Family-based association study
A genomic DNA bank was constituted in October 2000 to collect samples from IgAN patients and their relatives living in some European countries (all necessary information is available in the IgAN Consortium website www.igan.net).
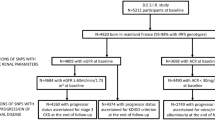
This family-based association study was performed in various pedigree constellations including 183 subjects for whom DNA samples were available (53 IgAN patients, 45 complete trios, 4 incomplete trios and 36 discordant siblings). All the families were living in the Puglia region in Southern Italy. Relatives were checked for urinalysis and examined after informed consent following the protocol of the IgAN Consortium approved by the local Ethics Committee. The IgAN Consortium was supported financially by the European Commission (5th Framework Programme: QLG1-CT-2000-00464). A detailed family history was obtained from all participating relatives. Additional historical information was obtained by family physicians. A database was organized to include personal and clinical data. Exposure to presumptive environmental factors such as type of job, food intake, exposure to solvents or other stimuli was also considered.
Case–control cohort
One hundred and seventy-four IgAN patients were retrospectively enrolled from a large cohort of 437 outpatients, all diagnosed between 1973 and 2000. They agreed to participate in this genetic study from October 2000 to December 2003; this group included the 53 patients analyzed in the family-based study.
Diagnosis of IgAN was based on the presence of IgA as the sole, predominant or co-dominant immunoglobulin in the mesangial area of the glomeruli, excluding systemic disease (Schönlein–Henoch purpura, systemic lupus erythematosus) and liver cirrhosis. The clinical picture was characterized by recurrent episodes of macroscopic hematuria in concomitance with upper respiratory tract infections or persistent microscopic hematuria and/or proteinuria. Measurements of serum creatinine, creatinine clearance, urinalysis, daily proteinuria and blood pressure were obtained from each patient. No patient had received corticosteroid or immunosuppressive therapy in the 3 months before the study. One hundred and sixty-six IgAN patients, with a follow-up period of at least 1 year (median follow-up of 5 years, range 1–30 years), were identified as progressors (n=50) and non-progressors (n=116). End-stage renal disease (ESRD) and/or doubled serum creatinine (sCr) value from the time of renal biopsy were considered as an outcome measure.
One hundred eighty-five subjects were recruited among healthy blood donors. They were matched for age, gender and ethnicity to the IgAN patients. Moreover, they were checked for negative urinalysis and no history of renal disease, diabetes, hypertension or metabolic disorders. Written informed consent for genomic DNA extraction was obtained from each subject enrolled.
Genomic DNA extraction and genotyping of cytokine polymorphisms
Whole blood (5 ml) was collected in EDTA tubes from each subject enrolled in the study. Genomic DNA was isolated by standard method using the Blood and Cells Culture midi DNA kit (Qiagen, Hilden, Germany).
Genotyping of the IFNγ intron-1 microsatellite polymorphism positioned at 1339–1373, C/T polymorphism in the human IL-13 promoter at position –1055, C/T polymorphism in TGF-β codon 25, IL10-G and IL10-R microsatellites, and TNFα –238 and –308 promoter polymorphisms was performed. Loci-specific amplification and gene polymorphisms were analyzed by using primers, conditions and methods previously published and listed in Table 1.8, 10, 16, 17
Genotyping of intron-1 IFNγ microsatellite
Genotyping of the IFNγ CA repeat was performed by PCR amplification and capillary electrophoresis. Genomic DNA (50 μg) was amplified in a reaction volume of 12.5 μl. Forward primer was 6-FAM labeled at the 5′ end and reverse primer was modified with a tail addition (7 bp) to reduce stutter band artifact. The reaction mixture contained 0.5 U AmpliTaq Gold DNA polymerase (Applied Biosystem, Forster City, CA, USA), 0.5 μ M of each specific primer, 2.5 mM MgCl2 and 200 μ M of each dNTP. The initial denaturation (10 min at 95°C) was followed by 40 cycles of 30 s at 95°C, 45 s at 48°C and 30 s at 72°C and a final extension time of 7 min at 72°C. Each amplified product, previously diluted 1:20, was mixed with 0.5 μl of Rox-conjugate Genescan-350 DNA size standard (Applied Biosystem, Forster City, CA, USA) and formamide-EDTA (25 mmol/l). This mixture was loaded onto an ABI Prism 310 capillary DNA sequencer (Applera Biosystem, Forster City, CA, USA). Specific Genotyper 3.7 software was used to size each allele obtained.
DNA sequencing was performed by BigDye terminator chemistry on 15ng of purified PCR product (Montage PCR, Millipore, Bedford, MA, USA) and analyzed on an ABI Prism 3100 capillary DNA sequencer.
In vitro IFNγ production
IFNγ production was measured in the supernatants of the cultured peripheral blood mononuclear cells (PBMC) from five homozygous IgAN patients for the 12-CA repeat/+874T haplotype and five homozygous for the 13-CA repeat/+874A haplotype of IFNγ intron 1. Cultured PBMC were stimulated with concanavalin A (5 μg/ml) for 48 h at 37°C in an atmosphere of 5% CO2. IgAN patients were adequately matched for clinical and laboratory findings, and had no evidence of inflammatory processes at the time of the PBMC isolation. IFNγ levels were measured using an ELISA kit (Bender MedSystems, Austria) according to the manufacturer's instructions.
IFNγ +874T/A variants subcloning
The +874A and +874T IFNγ inserts were obtained by PCR from DNA target of two previously genotyped subjects, respectively, homozygous for the 12-CA repeat/+874T haplotype and homozygous for the 13-CA repeat/+874A haplotype. Each PCR primer used included a different restriction site chosen in the polylinker region of the eligible vector (forward 5′-CTTAACGCTAGCAATATTCAGACATTCACA-3′ containing the NheI restriction site; reverse 5′-TTCATCAGATCTGAGCTTTAAAAGATAGTT-3′ containing the BglII restriction site). Amplicons were digested with NheI and BglII and subcloned into the pGL2-promoter luciferase reporter vector (Promega, Madison, WI, USA). Orientation and identification were confirmed by DNA sequencing.
Transient transfections and luciferase assay
To evaluate the different NF-κB binding activity of IFNγ +874A-pGL2 and IFNγ +874T-pGL2 promoter constructs, we transiently transfected HK2 cells, an immortalized proximal tubular epithelial cell line (ATCC, Manassas, VA, USA). HK2 cells transfected with an expression vector containing the luciferase cDNA under the control of three NF-κB consensus sequences (pLucNF-κB) were used as positive control. Transient transfection was carried out by electroporation using the Gene Pulser II RF module (Biorad, Hercules, CA, USA). HK2 cells (5 × 106) were resuspended in 0.5 ml medium containing 2.5 μg of pCMVβGal and 7.5 μg of IFNγ +874A-pGL2 or IFNγ +874T-pGL2 promoter constructs. Electroporation was carried out at 950 μF and 0.4 kV, and cells were then cultured for 24 h in a six-well plate (8 × 105 cells/well) in DMEM/F12 supplemented with 10% FCS, 100 U/ml penicillin, 100 μg/ml streptomycin and 2 mmol/l L-glutamine. Triplicate wells were incubated with and without human soluble CD40 ligand alone (sCD40L; 0.1 μg/ml) or along with caffeic acid phenetyl ester (CAPE; 10 μg/ml) in 1 ml of serum-free DMEM/F12 for 24 h. Following incubation, the cells were lysed in 100 μl of Luciferase Reporter lyses buffer supplied with the Luciferase Assay System (Promega). The extracts were assayed for luciferase activity using a DIGENE DCR-1 luminometer (Abbott Laboratories, Abbot Park, IL, USA). Luciferase activity was normalized to β-galactosidase.
Statistical analysis
The family-based association test (FBAT) software was used to evidence association between the Th1/Th2/Th3/TR-type cytokine polymorphisms and IgAN. This method is an extension of the original transmission disequilibrium Test (TDT).18 It incorporates a set of statistical procedures to accommodate variable pedigree constellations, dichotomous or quantitative phenotypes, phenotype-unknown subjects, bi- or multi-allelic marker data, multiple loci and various models for the mode of inheritance and gene–environment interactions. The FBAT statistic was calculated under an additive model and for both the multi-allelic (all alleles at a marker were compared simultaneously in one test) and bi-allelic (each allele against all others) mode of testing. The FBAT statistic for the bi-allelic model was calculated using the -o option for unknown disease prevalence in the population.19, 20, 21, 22 The Bonferroni correction, in which the P-value is multiplied for a number of tests equal to the number of alleles, was carried out when each allele was considered separately.
The power of our study design was evaluated by PBAT (analytical power calculation for family-based association test) and it resulted ineffective in detecting association with alleles with a low frequency (P<0.5).23
Cumulative incidence of developing end-point events from the time of renal biopsy was estimated by the Kaplan–Meier method for censored data and compared by log-rank test for stratifying variables. Continuous variables were compared by t-test or Mann–Whitney test, as applicable. Dichotomous and polytomous data were compared by the χ2 test using Statview 5.0 Software (1992–1998 SAS Institute Inc.).
Results
The family-based analysis of the polymorphisms related to the studied cytokines is illustrated in Tables 2 and 3. The multi-allelic analysis, assuming an additive model for which every cytokine gene acts on the phenotype, evidenced a significant association of IgAN with IFNγ intron-1 microsatellite polymorphism located at position 1349–1373 (P=0.03). No significant association was found between other cytokine gene polymorphisms and the disease (Table 2).
We identified seven different alleles for IFNγ intron-1 microsatellite polymorphism with a heterozygosity rate of 69% (126 out of 183). We, then, used a bi-allelic mode of analysis to evaluate how these different IFNγ alleles influenced the disease. The bi-allelic analysis, using the o option for unknown disease prevalence in the population, evidenced a significant association between the 13-CA repeat polymorphism and IgAN (P=0.006; Bonferroni P-value=0.04), suggesting that this allele was preferentially transmitted to the affected individuals (Table 3). It was transmitted 32 times from a heterozygous parent to the index patient and remained untransmitted 15 times. These results were obtained from 40 informative families that had at least one heterozygous parent. The total sample was in Hardy–Weinberg equilibrium.
We genotyped an additional case–control cohort (174 cases and 185 healthy controls). The clinical characteristics, at the time of renal biopsy, of the IgAN patients analyzed (121 men, 53 women) were mean age 28±10 years, mean serum creatinine 1.26±0.76 mg/dl, mean daily proteinuria value 1.12±1.28 g/day and mean serum creatinine clearance (according to Cockcroft and Gault formula) 94±32 ml/min; arterial hypertension was present in 40.3% of the patients. These baseline demographic and clinical findings did not show a statistically significant difference when compared to the source population (n=437). The overall comparison of CA repeat distribution between IgAN patients and controls did not give a statistically significant deviation. Although up to seven alleles for the CA repeat within intron 1 of IFNγ have been found, two of them (12 and 13-CA repeat) account for 87% of alleles in both groups. The discordant results between the familial and case–control studies suggest a possible genetic imprinting of the IFNγ intron 1 polymorphism, but a more detailed familial analysis considering the transmission of IFNγ CA repeats from mother and father separately does not evidence a possible parent-of-origin effect.
We moved on to study the influence of IFNγ intron-1 13-CA repeat polymorphism on the progression of the disease. Clinical findings and laboratory data at the time of renal biopsy in 174 IgAN patients with/without occurrence of this polymorphism were analyzed. No association was found between the presence/absence of risk genotype and gender, age at onset of the disease and at the time of renal biopsy, serum creatinine, renal function, daily proteinuria and histological lesions. The renal survival was also evaluated in 166 IgAN patients identified as progressor and non-progressor. Cumulative renal survival rates at 15 years from renal biopsy were 47.8% in homozygous and 46.5% in heterozygous patients for the 13-CA repeat allele versus 44.1% of the non-carriers; no significant difference was observed in the renal survival rates between IgAN patients bearing and non-bearing the IFNγ 13-CA repeat allele; log-rank test, χ2=0.005, P=0.99 (Figure 1a).
The 15 years renal survival in IgAN patients in respect to IFNγ gene polimorphisms. The outcome measures were represented by the doubling of baseline serum creatinine value and end-stage renal disease. Patients at risk were the number of cases in observation at each time. (a) Renal survival in IgAN patients bearing (+) and non-bearing (−) the IFNγ 13-CA repeat allele. No significant difference was observed (log-rank test, χ2=0.005, P=0.99). (b) Renal survival in homozygous subjects for 13-CA repeat/+874A haplotype, 12-CA repeat/+874T haplotype and heterozygous subjects for these haplotypes. No significant difference was observed (log-rank test, χ2=0.745, P=0.69).
The direct sequencing of the PCR products, obtained from genotyping the IFNγ intron-1 microsatellite, evidenced a strict association between the 13-CA repeat allele and the +874A variant of the +874T/A SNP (χ2=57.4, P<0.0001).
Renal survival was not different between homozygous subjects for 13-CA repeat/+874A haplotype (32 of 166), homozygous subjects for 12-CA repeat/+874T haplotype (31 of 166) and heterozygous subjects for these haplotypes (58 of 166); log-rank test, χ2=0.75, P=0.69 (Figure 1b). The same results were obtained considering only +874T/A SNP genotype (data not shown).
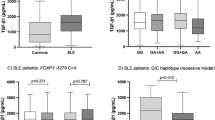
The in vitro PBMC production of IFNγ after mitogen stimulation was significantly lower in homozygous subjects for the 13-CA repeat/+874A haplotype than that in homozygous subjects for the 12-CA repeat/+874T haplotype (P=0.028, Mann–Whitney test) (Figure 2).
IFNγ production differences seem largely due to polymorphisms associated with the CA microsatellite. The +874T/A polymorphism occurs in an NF-κB binding site. To assess the functional role of these two variants, we transfected HK2 cells with IFNγ +874A-pGL2 and IFNγ +874T-pGL2 promoter constructs. HK2 cells were stimulated with an NF-κB activator (sCD40L) in the presence or absence of a specific inhibitor (CAPE). We observed a statistically significant difference in NF-κB binding activity of IFNγ +874A-pGL2 and IFNγ +874T-pGL2 promoter constructs (P=0.0007) (Figure 3).
HK2 cells were transiently transfected with the IFNγ +874A-pGL2 and IFNγ +874T-pGL2 promoter constructs and the luciferase activity was measured. Cells were stimulated with an NF-κB activator (sCD40L) in the presence or absence of a specific inhibitor (CAPE). HK2 cells transfected with an expression vector containing the luciferase cDNA under the control of three NF-κB consensus sequences (pLucNF-κB) were used as positive control. A statistically significant difference in the NF-κB binding affinity, due to +874T/A IFNγ SNP (rs2430561), is evidenced (P=0.0007). *sCD40L versus basal (P<0.003).
Discussion
Our family-based study shows an association between IgAN and 13-CA repeat allele of IFNγ intron-1 microsatellite. This genotype seems to confer susceptibility to the disease, but to have no influence on its progression. We demonstrate an absolute association between the 13-CA repeat and the +874A variant of the +874T/A SNP directly adjacent to the 5′ end of the microsatellite region. Homozygous subjects for 13-CA repeat and +874A allele show an in vitro statistically significant lower production of IFNγ than the homozygous subjects for 12-CA allele and +874T in response to mitogen stimulation. This SNP is situated within a putative binding site for the transcription factor NF-κB.8, 9, 10 Transient transfection experiments clearly show that +874T/A variants modulate differently NF-κB binding activity. As this protein plays an important role in the transcriptional regulation of IFNγ gene,24 the +874T and +874A alleles could be responsible for high and low IFNγ production, respectively.
The present study is the first family-based study that investigates the association between cytokine polymorphisms and IgAN. This disease has a multifactorial etiology including environmental influences and a complex genetic background. The genetic analysis of cytokine polymorphisms in complex diseases has traditionally focused on case–control association studies.25 Most study designs involve too small sample sizes lacking the statistical power to detect the correct gene effects. Another principal limitation of these studies is represented by the possibility that a positive association is biologically irrelevant because of population admixture.26 During the last decade, family-based association studies have challenged the case–control design. In the absence of both linkage and association between marker and disease locus, marker alleles are transmitted randomly from parents to offspring. This study design prevents errors of the first order, arising when investigators enroll cases and possibly mismatched unrelated controls. In addition, the case–control approach fails to detect an association when a parent-of-origin effect occurs. We find an association using a familiar approximation but not in a case–control analysis. We do not detect genetic imprinting in the transmission of the IFNγ CA polymorphism to IgAN patients. However, we will better investigate this hypothesis enrolling more case-parent-triads, since this effect was observed in another genetic immune complex mucosal disorder.27
In the present study, FBAT software was used to evaluate allele transmission.19, 20, 21, 22 This is the best approach to accommodate bi- and multi-allelic markers and to analyze the IgAN patients' different pedigree constellations available in our DNA genomic bank. Masutani et al13 in a traditional case–control study firstly described a significantly higher incidence of IFNγ 13-CA repeat allele in Japanese IgAN patients, with influence on the development of the disease. In complex diseases, it is particularly important to replicate findings before definite conclusions can be drawn. The present family-based study explored this possible genetic association in a completely independent patient group. Great relevance and strength was given to this finding, as a preferential transmission of IFNγ 13-CA repeat allele has been observed in Caucasian IgAN patients using a family-based association study. Moreover, our study shows a strict association of 13-CA repeat allele with +874A genotype and low in vitro production of IFNγ that has relevance in conferring susceptibility to IgAN, but not to its progression. This may be due to possible differences in the treatment of patients, when ACE-inhibitors were not available. However, this result is similar to that obtained by Masutani et al, 13 who demonstrated an impact on disease progression only for IL-4 B1 allele polymorphism and B1/B1 genotype.
IFNγ gene intron-1 polymorphisms influence immune complex disease susceptibility characterized by an imbalance of the immunoregolatory systems, as reported in recently published papers.27, 28, 29, 30 The balance between Th1- and Th2-type cytokine production modulates the immune response in renal disease like primary glomerulonephritis.31 Controversial results have been published on Th1/Th2 polarization of nephritogenic immune responses in IgAN.32, 33, 34, 35 The different involvement of a Th1 or Th2 immune response reported in IgAN may be due to both the polymorphous clinical phenotype and the variation of the cytokine profile at different times or stages of the disease in a genetically susceptible host. As the +874A allele significantly correlates with an in vitro lower IFNγ production, the association between this genotype and IgAN development is suggestive of a predominance of the Th2 immune response. Hyperfunction of Th2 cells and cytokine polarity are linked to a more nephritogenic pattern of IgA1 glycosylation in the animal model, and the decreased glycosylation of IgA1 elicited by Th2 cytokines is blunted in vitro by the addition of IFNγ.36 There is evidence of a preferential Th2-mediated immune response involvement as a consequence of airway inflammation,3 and a recent report described tonsillar IgA1 as a possible source of hypoglycosylated IgA1 in the serum of IgAN patients.37 In conclusion, an unbalanced Th2 cytokine production in response to upper respiratory tract infections is a significant pathogenic factor in human IgAN, as it may lead to abnormalities in IgA1 glycosylation and formation of circulating IgA1–IgG (anti-IgA1) immune complexes. However, the pathogenesis of IgAN implies that multiple mechanisms might act concomitantly to determine and to affect disease phenotype.
References
Maloy KJ, Powrie F : Regulatory T cells in the control of immune pathology. Nat Immunol 2001; 2: 816–822.
Podolsky DK : Mucosal immunity and inflammation. V. Innate mechanisms of mucosal defense and repair: the best offense is a good defense. Am J Physiol 1999; 277: G495–G499.
Neurath MF, Finotto S, Glimcher LH : The role of Th1/Th2 polarization in mucosal immunity. Nat Med 2002; 8: 567–573.
Weiner HL : Oral tolerance: immune mechanisms and the generation of Th3-type TGF-β-secreting regulatory cells. Microbes Infect 2001; 3: 947–954.
Akbari O, Dekruyff RH, Umetsu DT : Pulmonary dentritic cells producing Il-10 mediate tolerance induced by respiratory exposure to antigen. Nat Immunol 2001; 2: 725–731.
Warzocka K, Bienvenu J, Coiffier B, Salles G : Mechanism of action of tumor necrosis factor and lymphotoxin ligand–receptor system. Eur Cytokine Netw 1995; 6: 83–96.
Pravica V, Asderakis A, Perrey C, Hajeer A, Sinnott PJ, Hutchinson IV : In vitro production of IFNγ correlates with CA repeat polymorphism in the human IFNγ gene. Eur J Immunogenet 1999; 26: 1–3.
Laundy GJ, Spink CF, Keen LJ, Wood NA, Bidwell JL : A novel polymorphism in the human interleukin-13 (IL-13) promoter. Eur J Immunogenet 2000; 27: 53–54.
Grainger DJ, Heathcote K, Chiano M et al.: Genetic control of the circulating concentration of transforming growth factor type beta 1. Hum Mol Genet 1999; 8: 93–97.
Eskdale J, Gallagher G, Verweij CL, Keijsers V, Westendorp RG, Huizinga TW : Interleukin 10 secretion in relation to human IL-10 locus haplotypes. Proc Natl Acad Sci,USA 1998; 95: 9465–9470.
Kroeger KM, Carville KS, Abraham LJ : The −308 tumor necrosis factor-alpha promoter polymorphism effects transcription. Mol Immunol 1997; 34: 391–399.
Awad MR, El-Gamel A, Hasleton P, Turner DM, Sinnott PJ, Hutchinson IV : Genotypic variation in the transforming growth factor-beta1 gene: association with transforming growth factor-beta1 production, fibrotic lung disease, and graft fibrosis after lung transplantation. Transplantation 1998; 27: 1014–1020.
Masutani K, Miyake K, Nakashima H et al.: Impact of Interferon-γ and Interleukin-4 gene polymorphisms on development and progression of IgA nephropathy in Japanese patients. Am J Kidney Dis 2003; 41: 371–379.
Pravica V, Perrey C, Stevens A, Lee JH, Hutchinson IV : A single nucleotide polymorphism in the first intron of the human IFN-γ gene: absolute correlation with a polymorphic CA Microsatellite marker of high IFN-γ production. Hum Immunol 2000; 61: 863–866.
Rossouw M, Nel HJ, Cooke GS, van Helden PD, Hoal EG : Association between tuberculosis and a polymorphic NFκB binding site in interferon γ gene. Lancet 2003; 361: 1871–1872.
Di Paolo S, Schena A, Stallone G et al.: Captopril enhances transforming growth factor (TGF)-beta1 expression in peripheral blood mononuclear cells: a mechanism independent from angiotensin converting enzyme inhibition? A study in cyclosporine-treated kidney-transplanted patients. Transplantation 2002; 74: 1710–1715.
Jang WH, Yang YI, Yea SS et al.: The −238 tumor necrosis factor-alpha promoter polymorphism is associated with decreased susceptibility to cancers. Cancer Lett 2001; 10: 41–46.
Spielman RS, McGinnis RE, Ewens WJ : Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am J Hum Genet 1993; 52: 506–516.
Horvath S, Xu X, Laird NM : The Family Based Association Test Method: strategies for studying general genotype–phenotype associations. Euro J Hum Genet 2001; 9: 301–306.
Lunetta KL, Farone SV, Biederman J, Laird NM : Family-based tests of association and linkage using unaffected sibs, covariates and interaction. Am J Hum Genet 2000; 66: 605–614.
Rabinowitz D, Laird NM : A unified approach to adjusting association tests for population admixture with arbitrary pedigree structure and arbitrary missing marker information. Hum Hered 2000; 50: 211–233.
Lake SL, Blacker D, Laird NM : Family-based tests of association in presence of linkage. Am J Hum Genet 2000; 67: 1515–1525.
Lange C, Laird NM : Power calculations for a general class of family-based association tests: dichotomous traits. Am J Hum Genet 2002; 71: 575–584.
Rengarajan J, Szabo SJ, Glimcher LH : Transcriptional regulation of Th1/Th2 polarization. Immunol Today 2000; 21: 479–483.
Bidwell J, Keen L, Gallagher G et al.: Cytokine gene polymorphism in human disease: on-line databases. Genes Immun 1999; 1: 3–17.
Cardon LR, Palmer LJ : Population stratification and spurious allelic association. Lancet 2003; 361: 598–604.
Rueda B, Martinez A, Lopez-Nevot MA et al.: A functional variant of IFNγ gene is associated with coeliac disease. Genes Immun 2004; 5: 517–519.
Tsiavou A, Hatziagelaki E, Chaidaroglou A, Koniavitou K, Degiannis D, Raptis SA : Correlation between intracellular interferon-gamma (IFN-gamma) production by CD4+ and CD8+ lymphocytes and IFN-gamma gene polymorphism in patients with type 2 diabetes mellitus and latent autoimmune diabetes of adults (LADA). Cytokine 2005; 31: 135–141.
Cantor MJ, Nickerson P, Bernstein CN : The role of cytokine gene polymorphisms in determining disease susceptibility and phenotype in inflammatory bowel disease. Am J Gastroenterol 2005; 100: 1134–1142.
Spriewald BM, Witzke O, Wassmuth R et al.: Distinct tumour necrosis factor alpha, interferon gamma, interleukin 10, and cytotoxic T cell antigen 4 gene polymorphisms in disease occurrence and end stage renal disease in Wegener's granulomatosis. Ann Rheum Dis 2005; 64: 457–461.
Schena FP, Gesualdo L, Grandaliano G, Montinaro V : Progression of renal damage in human glomerulonephritides: is there sleight of hand in winning the game? Kidney Int 1997; 52: 1439–1457.
Lim SC, Zheng S, Kim YS et al.: Th1/Th2 predominamnce and proinflammatory cytokines determine the clinicopathological severity of IgA nephropathy. Nephrol Dial Transplant 2001; 16: 269–275.
Lim SC, Yoon HJ, Kim YS et al.: Clinicopathological correlation of intrarenal cytokine and chemokines in IgA nephropathy. Nephrology 2003; 8: 21–27.
Shu KH, Lee SH, Cheng CH, Wu MJ, Lian JD : Impact of Interleukin-1 receptor antagonist and tumor necrosis factor-alfa gene polymorphism on IgA nephropathy. Kidney Int 2000; 58: 783–789.
Ebihara I, Hirayama K, Yamamoto S, Muro K, Yamagata K, Koyama A : Th2 predominance at the single-cell level in patients with IgA nephropathy. Nephrol Dial Transplant 2001; 16: 1783–1789.
Chintalacharuvu SR, Nagy NU, Sigmund N, Nedrud JG, Amm ME, Emancipator SN : T cell cytokines determine the severity of experimental IgA nephropathy by regulating IgA glycosylation. Clin Exp Immunol 2001; 126: 326–333.
Itoh A, Iwase H, Takatani T et al.: Tonsillar IgA1 as a possible source of hypoglycosylated IgA1 in the serum of IgA nephropathy patients. Nephrol Dial Transplant 2003; 18: 1108–1114.
Acknowledgements
This study was supported by the 5th European Framework Programme (QLG1-CT-2000-00464), the Ministero dell'Istruzione, Universita' e Ricerca PRIN 2001-067748, L.488/92 Cluster 03 and FIRB 2001-RBNE013JYM, and CEGBA (Centro di Eccellenza Genomica in Campo Biomedico ed Agrario). We thank Paola Pontrelli, Michele Ursi and Giuseppe Grandaliano for their help in IFNγ variant functional study. We are also indebted to Mariella Mastrolonardo for the editorial assistance of the manuscript.
Author information
Authors and Affiliations
Consortia
Corresponding author
Additional information
Related world web site
Human cytokine gene nucleotide sequence alignments: http://www.bris.ac.uk/pathandmicro/services/cytokine2.htm
PubMed search engine, primed to search for cytokine gene polymorphisms: http://www.bris.ac.uk/pathandmicro/services/GAI/cytokine4.htm
FBAT Software package is freely available at web address: www.biostat.harvard.edu/~fbat/default.htlm
PBAT Software package is freely available at web address: http://www.biostat.harvard.edu/~clange/default.htm
Rights and permissions
About this article
Cite this article
Schena, F., Cerullo, G., Torres, D. et al. Role of interferon-γ gene polymorphisms in susceptibility to IgA nephropathy: a family-based association study. Eur J Hum Genet 14, 488–496 (2006). https://doi.org/10.1038/sj.ejhg.5201591
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201591
Keywords
This article is cited by
-
Omics are Getting Us Closer to Understanding IgA Nephropathy
Archivum Immunologiae et Therapiae Experimentalis (2023)
-
Effect of IFN-γ +874 T/A polymorphism on clinical manifestations of dengue: a meta-analysis
Journal of Genetics (2021)
-
T cells in IgA nephropathy: role in pathogenesis, clinical significance and potential therapeutic target
Clinical and Experimental Nephrology (2019)
-
Associations between functional polymorphisms and response to biological treatment in Danish patients with psoriasis
The Pharmacogenomics Journal (2018)
-
Evidence for genetic association of TBX21 and IFNG with systemic lupus erythematosus in a Chinese Han population
Scientific Reports (2016)