Abstract
In humans, spontaneous DZ twinning is known to have a genetic basis. A prior investigation in the Flemish and Dutch population showed that the phenotype of ‘having DZ twins’ was consistent with an autosomal monogenic dominant model, with a gene frequency of 3.5% and a female-specific lifetime penetrance of 10%. Recessive, X-linked, polygenic and sporadic models were rejected. This study reports on a genome-wide scan of 14 Flemish families containing 57 mothers of spontaneous DZ twins. Two-point linkage analysis using the autosomal dominant model showed nine chromosomal regions with a LOD score around 1. After multipoint linkage analysis, including heterogeneity, three chromosomes continued to give high LOD scores. These regions were further haplotyped with additional markers at 1 cM distance. The multipoint analysis was not in favour of linkage of the DZ twinning trait in most candidate genes and other regions (LOD score < −2) under the genetic model of autosomal dominance.
To further evaluate the evidence for linkage given some uncertainty about the correct mode of inheritance of twinning susceptibility other models of inheritance were tested. Results of this analysis showed all models gave highest LOD scores under dominant models.
If heterogeneity among the families is taken into account, the peaks that were observed on chromosome 2, 7 and 18 could well contain a potential gene contributing to DZ twinning. These results give suggestive evidence that the mode of inheritance of DZ twinning is probably more complex than was originally expected.
Similar content being viewed by others
Introduction
The underlying biological mechanism for natural DZ twinning is multiple ovulation. Animal1, 2 and human models3, 4 have indicated a genetic basis of this trait. In humans, spontaneous DZ twinning varies across racial groups, also suggesting a genetic basis. We investigated the inheritance of DZ twinning in the Flemish and Dutch population by segregation analysis and found that the phenotype of ‘having DZ twins’ was consistent with an autosomal monogenic dominant model, with a gene frequency of 0.035 and a female-specific lifetime penetrance of 0.10. X-linked, recessive, polygenic and sporadic models were rejected.5 Sensitivity analysis showed that the autosomal dominant model was robust to the changing population prevalence and the loss of information due to the presence of same-sexed twin pairs of unknown zygosity. The penetrance of the nongene carrier was zero, indicating that in our population all DZ twin births were due to a genetic predisposition. Finally, this autosomal dominant model would be compatible with the autosomal codominant gene for multiple ovulation in Booroola merino sheep.1
Using the same approach, we found no heterogeneity between the pedigrees ascertained through a spontaneous DZ twin and those where the proband DZ twin was born after fertility treatment of the mother.6 Practically, this could mean that only those mothers, who carry the gene(s) predisposing to bear DZ twins (DZ-twinning gene), are at risk for having twins or higher order multiple births after infertility treatment, while those mothers, who do not carry the DZ-twin gene(s), would not be at risk. Therefore, identification of the DZ-twinning gene(s) should allow screening women at risk for bearing twins and higher order multiple births.
Methods
Subjects
Families with at least three DZ twins within their three-generation pedigree were selected and the pedigrees extended if they included meioses informative for linkage analysis. These families were identified through records from our previous genetic epidemiologic study using twins and their families in East Flanders, Belgium.5 The ascertainment was population based through the East Flanders Prospective Twin Survey (EFPTS).7 All cases of fertility treatments were excluded. For the mothers of proband twins, these data were identified from the obstetrical records. For all other mothers this was explicitly asked.
A total of 14 families were selected containing 57 mothers of spontaneous DZ twins and 11 mothers of twins with unknown zygosity. The 14 pedigrees included 359 individuals and varied in size from 13 to 49 persons. Blood was drawn from mothers of DZ twins, their first degree relatives and any other individuals in lineages connecting mothers of DZ twins. If informative persons were deceased, blood was drawn from as many children as possible in order to reconstitute the parental genotypes. Samples were collected from a total of 174 individuals, of whom 40 were mothers of DZ twins.
Zygosity of the proband twins was determined by placentation and differences in sex, blood groups and DNA fingerprinting in equivocal cases. If the twins were of different sex or had at least two different genetic markers they were considered as DZ. The probability of dizygosity given concordance of all markers was <0.01. Zygosity of the other twins was determined according to the same criteria as the proband twins if blood was available. If not, a validated zygosity questionnaire was taken by phone or by mail.8 For 11 mothers of twins no information of the zygosity could be obtained because one or both twins died at or around birth.
Study protocols were reviewed and approved by the local Research Ethics Committee. Participation was voluntary and each patient gave informed consent.
Genotyping
For each subject, blood (20 ml) was drawn into EDTA containing tubes and stored at room temperature until white blood cell collection. White blood cell pellets were kept frozen until DNA extraction. A genome-wide screen was performed using markers at approximately 10 cM intervals from the set8A combo list from the Marshfield URL (http://research.marshfieldclinic.org/genetics/sets/combo.html), with some additional markers added to the set at certain chromosomal locations.9 The entire marker set consisted of 51 panels (400 markers), each containing markers pooled together according to size and fluorescent label (6-FAM, HEX and NED). Reaction conditions were standard for all markers used as described in the Marshfield PCR protocol (at the Marshfield web site). Each panel of pooled markers was electrophoresed on a 3700 DNA Analyser (PE Biosystems, Foster City, CA, USA).
Data analyses
Semiautomated sizing of alleles was carried out using the GeneScan 2.1. software (PE Biosystems, Foster City, CA, USA), and individual genotypes were assigned with the help of the Genotyper version 2.1 software (PE Biosystems, Foster City, CA, USA). To ensure accuracy of the genotypes, two individuals manually checked each genotype.
SLINK, a general simulation programme for linkage analysis10, 11 was used to conduct the initial power calculations. Two-locus linkage analysis was performed using MLINK10 and multipoint linkage analysis was performed by Simwalk2 (version 2.82),12 which gives multipoint LOD scores and heterogeneity LOD scores.
The phenotype of natural DZ twinning was defined as all mothers having spontaneously at least one pair of twins, of whom the birth weight of one of the children was 500 g. Specific genetic models tested for linkage to DZ twinning were (1) an autosomal dominant sex-limited model with a gene frequency of 0.035, sex-limited expression in females, overall penetrance of 0.10 and 0.013% phenocopies5 and (2) a sex-limited recessive model with a gene frequency of 0.50 and a penetrance of 0.05.4, 13 The carrier status of mothers of twins with unknown zygosity was assigned as unknown for the analysis.
The relationships of pedigree members were checked using Relative 1.10.14 Family structures were modified by adding dummy parents to any individuals who were indicated to be half-sibs based on the relationship checking. Mendelian inconsistencies found in the genotyping data using Pedcheck were sent back to the genotyping lab for error correction. MLINK was run on all chromosomes under two different models. Simwalk2 (version 2.82)12 haplotyping analysis was performed on all chromosomes that provided a LOD score of 1 or higher in MLINK. Simwalk2 (version 2.82) parametric analysis was performed on all chromosomes. Two markers on chromosomes 2 (D2S122 and D2S298) and one marker on chromosome 7 (D7S2423) and 18 (D21S369), respectively, were dropped because they showed an excess number of recombinations. The final data analysed included 14 families with 420 people and 350 autosomal markers.
Results
The simulation data are listed in Table 1. The simulations used a marker with 10 equally frequent alleles, which would be consistent with the results of a multipoint analysis. Runs were performed twice: once assuming no genetic heterogeneity between the families (α=1.0) and once assuming linkage at the same locus in only 75% of the families (α=0.75).
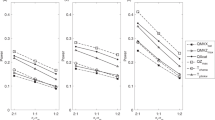
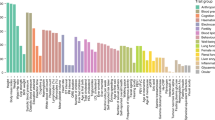
Overall, nine chromosomal regions showed a LOD score around one after two-point linkage analysis. After Simwalk2 parametric runs, including heterogeneity, three chromosomes continue to show high LOD scores. These three regions were further haplotyped with 5–12 additional markers per region at 1 cM distance. The results of the maximum multipoint LOD scores after haplotyping with these additional markers are shown in Table 2. Figure 1 gives an overview of the multipoint LOD scores for all chromosomes, respectively, with and without genetic heterogeneity between families. This multipoint analysis was not in favour of linkage of the DZ twinning trait in most regions (LOD score <−2) under the genetic model of autosomal dominance (see black lines in Figure 1).
LOD score graphs of all chromosomes with and without genetic heterogeneity. The vertical axis of each plot shows the multipoint LOD score ranging from −5 to +2.5. The black lines indicate the multipoint LOD scores without genetic heterogeneity, the grey lines with heterogeneity. The horizontal line on each subfigure (LOD=0) denotes the different markers along each chromosome. Note that the black line is often not visible because it frequently has values under the −5.
In Table 3, the multipoint LOD scores at known candidate gene regions are summarized assuming no heterogeneity and a dominant mode of inheritance. With the exception of chorionic gonadotropin alpha chain (FSHA) on chromosome 6, our data were not in favour of linkage of the genes in the FSH pathway,15, 16, 17 the gene encoding peroxisome proliferator-activated receptor-gamma (PPARG) on chromosome 3,18 methylenetetrahydrofolate reductase (MTHFR)19 on chromosome 1 and bone morphogenetic protein receptor type IB (BMPR1B) on chromosome 4.20
Our results demonstrating DZ twinning is inherited as a dominant model contradicted the sex-limited recessive model with a gene frequency of 0.50 and a penetrance of 0.05 suggested previously.4, 13 We therefore reanalysed our data using this model of inheritance, but could not find any positive 2-point LOD score higher than what was found with the dominant model.
To more completely evaluate the evidence for linkage given some uncertainty about the correct mode of inheritance of twinning susceptibility we further computed LOD scores over a grid of recurrence risk values ranging from 1.3 to 1.7 for dominant and recessive models of inheritance. Results of this analysis showed all models gave highest LOD scores under dominant models. We found some increase of linkage evidence with maximal LOD scores of 1.67 on chromosome 7 for a model with penetrances of 0.16, 1.37 (actually was a decrease from original model of 1.41) on chromosome 2 for a model with penetrance of 0.05 and 1.91 on chromosome 18 for a model with penetrance of 0.05.
We also explored linkage under several models incorporating parity and maternal age (Table 2). Parity and maternal age at the twin's birth from mothers of DZ twins in the multiplex families and from mothers of monozygotic twins in EFPTS (for twins born between 1970 and 2000) were used to generate penetrances. Initial exploration indicated a reasonable grouping was two parity groups (para 1 or 2, and para 3 or more) and three maternal age groups (1=under 25, 2=25–34, and 3=over 34). Probability density function values were generated for the three maternal age groups in the multiplex families (genetic cases) and the EFPTS (phenocopies) by parity groups. The resulting ratio was then used to calculate the final penetrances for each maternal age/parity group. Penetrances were computed for a twinning prevalence of 0.007 and gene frequencies of 0.01, 0.02, 0.035, and 0.1, and for a prevalence of 0.01 and gene frequencies of 0.01 and 0.035. A total of eight liability classes (male, unknown maternal age and parity at twin birth, and the 6 age/parity classes) were used for parametric linkage analysis. Files were arranged in standard LINKAGE format and prepared for analysis with Mega2.21 We analysed markers on chromosomes 2, 7, and 18 with Simwalk2 as described in the previous section.
Discussion
The genome-wide linkage scan is now a standard approach to deciphering the genetic basis of traits in the absence of good candidate genes. It has proven most effective for the localization of genes underlying monogenic traits, and since our segregation analysis suggested a major dominant genetic effect in DZ twinning, we adopted this method of analysis. We estimated that, using all 14 families, the power to detect linkage was sufficient and complex segregation analysis5 showed that the phenotype of ‘having DZ twins’ was consistent with an autosomal monogenic dominant model. However, we could not detect any positive LOD score higher than 2. The microsatellite markers provided adequate coverage across all autosomes, with an average distance between the markers of 9.8 cM and the largest gap being 33 cM. If we assume no heterogeneity among the families, there was complete exclusion of linkage to DZ twinning under genetic models of autosomal dominance across most of the chromosomes (Figure 1), as a LOD score of −2 (odds of 100 against 1) is considered significant evidence against linkage.22
DZ twinning results from the release and subsequent fertilization of multiple oocytes and survival of both embryos. Variation in spontaneous DZ twinning could result from differences in multiple ovulation frequency and/or embryo survival. Mothers of spontaneous DZ twins have a higher incidence of multiple ovulations and several studies report an increased concentration of follicle-stimulating hormone (FSH) during their menstrual cycle.15, 16 FSH release is controlled by a complex feedback mechanism from the ovary. Genes in the pathway controlling the synthesis, release and activity of FSH are therefore good candidates for increased DZ twinning frequencies.17 With the exception of chorionic gonadotropin alpha chain (FSHA) on chromosome 6, our data are not in favour of linkage of genes in the FSH pathway, assuming no heterogeneity and a dominant mode of inheritance. This confirms previous findings.13, 23
Busjahn et al18 found linkage of DZ twinning to the gene encoding PPARG on chromosome 3. The authors suggested that intrauterine selection may be responsible for their findings. Methylenetetrahydrofolate reductase (MTHFR)19 on chromosome 1 has also be linked to DZ twinning, probably by influencing the proliferation of rapidly dividing embryonic and maternal cells. We could not replicate these findings as results in both these regions were not in favour of linkage with LOD scores of –3.5 (odds of 5000 against 1) under the dominant model. This also confirms the results of others.24, 25
The most important perceived limitation of the LOD score method is that the mode of inheritance and the genetic parameters must be specified in order to use the method. Segregation analysis may sometimes conclude to a wrong model, which would result in a reduction of power when conducting linkage analysis.26 The use of the dominant model, found in our segregation analysis, could therefore be erroneous and lead to reduced power to detect linkage with a true marker.
Reanalysing our data using a sex-limited recessive model with a gene frequency of 0.50 and a penetrance of 0.054, 13 did not reveal any positive two-point LOD score higher than what was found with the dominant model.
We further did a grid search over a range of possible values for the overall recurrence risk of twinning ranging from 1.3 to 1.7 and penetrances within those recurrence risks for recessive and dominant mode of inheritance. Overall, the dominant models continued to give the highest LOD scores within the same regions and assuming heterogeneity among the families. Some increase of linkage evidence was found for chromosomes 7 and 18, with a maximal LOD score of 1.67 for a model with penetrance of 0.16 and 1.91 for a model with penetrance of 0.05, respectively. At a population level, known risk factors for the birth of spontaneous DZ twins are maternal age and parity.4, 27 We therefore calculated some penetrances based on maternal age and parity at twin birth. Data from the multiplex families were used as genetic cases and mothers of monozygotic twins in the East Flanders registry as sporadic cases. There was, however, not much distinction between the two groups, so not much gain in power could be expected. We used various values of gene frequency to calculate the final penetrance values, and a minimal gain in linkage signal was obtained with maximum LOD scores of 1.51, 1.36, and 1.99 for the chromosomes 2, 7 and 18, respectively (Table 2).
We have focused on the Flemish population in hopes of reducing heterogeneity among families and increasing the detectable effects of any particular locus. Despite this, we could not find any region with a LOD score above 2 that could be linked to the trait of DZ twinning, defined as ‘having DZ twins’ and assuming no heterogeneity. Although a study like this cannot exclude mutations with very small effects, we can conclude from our results that it is unlikely that there exists mutations with large effects in the majority of our families, and that the mode of inheritance is probably more complex than was originally expected.
If, however, the mode of inheritance is complex and heterogeneity among the families is taken into account, then the peaks that we observe on chromosome 2, 7, and 18 could well contain a potential gene contributing to DZ twinning. The maximum LOD scores found with the different models tested are all located near or within the same regions as compared to the original model and the proportion of linked pedigrees also seem to be constant through the different models. Further work is necessary to support these findings.
References
Montgomery GW, Crawford AM, Penty JM et al: The ovine Booroola fecundity gene (FecB) is linked to markers from a region of human chromosome 4q. Nat Genet 1993; 4: 410–414.
Davis GH, McEwan JC, Fennessy PF, Dodds KG, Farquhar PA : Evidence for the presence of a major gene influencing ovulation rate on the X chromosome of sheep. Biol Reprod 1991; 44: 620–624.
Lewis CM, Healey SC, Martin NG : Genetic contribution to DZ twinning. Am J Med Genet 1996; 61: 237–246.
Bulmer MG : The Biology of Twinning in Man. Oxford: Clarendon, 1970.
Meulemans WJ, Lewis CM, Boomsma DI et al: Genetic modelling of dizygotic twinning in pedigrees of spontaneous dizygotic twins. Am J Med Genet 1996; 61: 258–263.
Meulemans WJ, Lewis CM, Boomsma DI et al: Segregation analysis of dizygotic twinning in pedigrees of dizygotic twins conceived after artificial induction of ovulation. Am J Hum Genet 1994; 55: A158.
Loos R, Derom C, Vlietinck R et al: The East Flanders Prospective Twin Survey (Belgium): a population-based register. Twin Res 1998; 1: 167–175.
Peeters H, Van Gestel S, Vlietinck R et al: Validation of a telephone zygosity questionnaire in twins of known zygosity. Behav Genet 1998; 28: 159–163.
Jawaheer D, Seldin MF, Amos CI et al: A genomewide screen in multiplex rheumatoid arthritis families suggests genetic overlap with other autoimmune diseases. Am J Hum Genet 2001; 68: 927–936.
Ott J : Computer-simulation methods in human linkage analysis. Proc Natl Acad Sci USA 1989; 86: 4175–4178.
Weeks D, Ott J, Lathrop G : SLINK: a general simulation program for linkage analysis. Am J Hum Genet 1990; 47: A204.
Sobel E, Lange K : Descent graphs in pedigree analysis: applications to haplotyping, location scores, and marker-sharing statistics. Am J Hum Genet 1996; 58: 1323–1337.
Montgomery GW, Duffy DL, Hall J et al: Mutations in the follicle-stimulating hormone receptor and familial dizygotic twinning. Lancet 2001; 357: 773–774.
Göring H, Ott J : Verification of sib relationship without knowledge of parental genotypes. Am J Hum Genet 1995; 57: A192.
Gilfillan CP, Robertson DM, Burger HG et al: The control of ovulation in mothers of dizygotic twins. J Clin Endocrinol Metab 1996; 81: 1557–1562.
Nylander PP : Serum levels of gonadotrophins in relation to multiple pregnancy in Nigeria. J Obstet Gynaecol Br Commonw 1973; 80: 651–653.
Lockwood GM, Muttukrishna S, Ledger WL : Inhibins and activins in human ovulation, conception and pregnancy. Hum Reprod Update 1998; 4: 284–295.
Busjahn A, Knoblauch H, Faulhaber HD et al: A region on chromosome 3 is linked to dizygotic twinning. Nat Genet 2000; 26: 398–399.
Hasbargen U, Lohse P, Thaler CJ : The number of dichorionic twin pregnancies is reduced by the common MTHFR 677C-->T mutation. Hum Reprod 2000; 15: 2659–2662.
Fabre S, Pierre A, Pisselet C et al: The Booroola mutation in sheep is associated with an alteration of the bone morphogenetic protein receptor-IB functionality. J Endocrinol 2003; 177: 435–444.
Mukhopadhyay N, Almasy L, Schroeder M, Mulvihill W, DE W : Mega2, a data-handling program for facilitating genetic linkage and association analyses. Am J Hum Genet 1999; 65: A436.
Ott J : Analysis of Human Genetic Linkage. Baltimore: Johns Hopkins University Press, 1999.
Derom C, Groenen P, Vlietinck R : Follicle-stimulating-hormone receptor and twinning. Lancet 2001; 357: 230–231, author reply 231–232.
Duffy D, Montgomery G, Treloar S et al: IBD sharing around the PPARG locus is not increased in dizygotic twins or their mothers. Nat Genet 2001; 28: 315.
Montgomery GW, Zhao ZZ, Morley KI et al: Dizygotic twinning is not associated with methylenetetrahydrofolate reductase haplotypes. Hum Reprod 2003; 18: 2460–2464.
Dizier MH, Babron MC, Clerget-Darpoux F : Conclusion of LOD-score analysis for family data generated under two-locus models. Am J Hum Genet 1996; 58: 1338–1346.
Campbell D : Natural factors influencing multiple gestation; in Blickstein I, Keith LG (eds): Multiple pregnancy: epidemiology, gestation & perinatal outcome. London: New York, Taylor & Francis, 2005, pp 87–93.
Acknowledgements
This research was supported by the Fund for Scientific Research – Flanders (G.0408.99) and Twins, a non-profit Association for Scientific Research in Multiple Births. We thank all families for their enthusiastic cooperation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Derom, C., Jawaheer, D., Chen, W. et al. Genome-wide linkage scan for spontaneous DZ twinning. Eur J Hum Genet 14, 117–122 (2006). https://doi.org/10.1038/sj.ejhg.5201522
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201522
Keywords
This article is cited by
-
A major QTL at the LHCGR/FSHR locus for multiple birth in Holstein cattle
Genetics Selection Evolution (2021)
-
Genetic control of multiple births in low ovulating mammalian species
Mammalian Genome (2012)