Abstract
TGF-β-receptor 2 (TGFBR2) gene defects have been recently associated with Marfan syndrome (MFS) with prominent cardio-skeletal phenotype in patients with negative fibrillin-1 (FBN1) gene screening. Four mutations have been identified to date in five unrelated families. We screened TGFBR2 gene by direct automated sequencing in two adult patients diagnosed with MFS according to Ghent criteria, and in one girl clinically suspected as affected on the basis of a major cardiovascular criterion and skeletal involvement, all proven not to carry mutations in the exon–intron boundaries of FBN1 gene. We identified two novel and one known TGFBR2 gene mutations in the three unrelated probands. The D446N was identified in a 4-year-old girl with de novo disease characterized by severe cardiovascular disease and skeletal involvement. The M425V and R460H mutations were identified in two familial, autosomal dominant MFSs, both characterized by major cardio-skeletal signs and absence of major ocular signs. The mutation R460H has been recently reported in a family with thoracic aortic aneurysms and dissection. The three mutations are absent in 192 controls and affect evolutionarily conserved residues of the serine/threonine kinase domain (exon 5). Our data support the recently reported association between TGFBR2 gene and MFS without major ocular signs (MFS2). The number of genotyped cases however is too low to confirm that major ocular signs are characteristically absent in MFS2. Accordingly, all patients proven or suspected to be affected by MFS with negative FBN1 gene screening could benefit from rapid investigation of the TGFBR2 gene.
Similar content being viewed by others
Introduction
Marfan syndrome (MFS, MIM #154700) is an autosomal dominant disorder of connective tissue with cardiovascular, skeletal and ocular abnormalities. More than 90% of MFS cases are caused by mutations in the fibrillin-1 gene (FBN1, chr. 15q21.1).1 A second locus (MFS2) (MIM #154705) was found to map at 3p25–p24.22 after the identification of a family with autosomal dominant MFS not linked to FBN1 gene.3 A 3p24.1 chromosomal breakpoint disrupting the transforming growth factor beta type II receptor (TGFBR2) gene in a Japanese individual with MFS led to consider TGFBR2 as a gene underlying association with MFS at the MFS2 locus.4 Then, a mutation disturbing the TGFBR2 canonical splicing site consensus sequence was identified in affected members of the French MFS family originally found not to be associated with FBN1. Three additional missense TGFBR2 mutations have been identified in four unrelated probands.4 In the families reported by Mizuguchi et al,4 major ocular signs were absent: the phenotype was characterized by major cardio-skeletal signs.
TGFBR2 gene (chr.3p225) is a putative tumor suppressor gene implicated in several malignancies, and now also proven as associated with inherited connective tissue disorders.4, 6 The TGFBR2 gene consists of seven exons and six introns, and encodes the human II TGF-β receptor (70/80 kDa). This receptor belongs to the serine–threonine kinase family of cell surface receptors, which regulate several cellular processes, including proliferation, cell cycle arrest, apoptosis, differentiation and formation of extracellular matrix.7, 8
We report two novel and one known TGFBR2 gene missense mutations in unrelated adults with familial autosomal dominant MFS characterized by major cardioskeletal signs, and in one child with a sporadic phenotype characterized by major cardiovascular signs and skeletal involvement. In the three probands, the screening of the exon–intron boundaries of the FBN1 gene was negative.
Materials and methods
All patients were referred to the Interdisciplinary Group for MFS (GISM) for suspected MFS and underwent multidisciplinary clinical and noninvasive instrumental study (cardiological, ophthalmological, orthopedic, radiological). Lumbo-sacral magnetic resonance imaging (MRI) for the detection of dural ectasia was only performed in patient #1. The aortic root diameter was evaluated with 2D echocardiography by experienced adult and pediatric echocardiographists. The calculation of the aortic root ratio (ARR) was based on the Roman et al9 criteria.
The diagnostic criteria for MFS were based on Ghent nosology in adults.10 The criterion for screening TGFBR2 gene was the negative screening of all exon–intron boundaries of the FBN1 gene in adult patients with proven or suspected MFS and in children with partial expression of MFS, and suspected to meet the full criteria at an older age.
The clinical and genetic program for MFS has been approved by the local Ethical Committee. Patients received detailed information on the MFS and related diagnostic work-up, likelihood of mutation identification, methods of analysis of the genes and related sensitivity. Written informed consent was obtained from all probands or, in case of age <18 years, from their parents, as well as from all relatives who underwent clinical and genetic screening.
DNA isolation and PCR
Genomic DNA was isolated from peripheral blood leukocytes by the QIAmp DNA blood kit (Qiagen Inc., Valencia, CA, USA). The exon–intron boundaries of the FBN1 gene had previously been screened using DHPLC.11 The analysis documented the absence of mutations. The seven exons and flanking regions of the TGFBR2 were amplified using previously reported intron-specific primers,8 with the exception of three amplicons (exons 1, 2 and 5) for which the two primers were newly designed using the Amplify 1.0 software:
- 1F::
-
5′TCCGGGAAGGCGCCGTCCGCT3′,
- 1R::
-
5′CGACTGTCAAGCGCAGCGGAGAG3′;
- 2F::
-
5′GCTGCCTGGCAGTTGGATAAT3′,
- 2R::
-
5′ACACTGACTGTGTGTACTATG3′;
- 5F::
-
5′AATGATGGGCCTCACTGTCT3′,
- 5R::
-
5′ACACAATGATGCTGGTCCAC3′.
Mutational analysis: The analysis was performed by direct automated sequencing. Direct sequencing analysis was performed using the BigDye Terminator Cycle sequencing kit V 3.1 (Applied Biosystems) on an ABI 3100 Genetic Analyzer, following the manufacturer's directions. Amplicon including exon 3 was amplified using a proof-reading Taq Polymerase (Phusion, Finnzymes, Finland) to avoid false-positive heterozygous Adel in the known instability site (A)10 of exon 3.
Controls were constituted of 384 chromosomes from 192 unrelated control individuals without Marfan phenotype. In all, 55 control amplicons including exon 3 were sequenced after amplification with the proof-reading Taq polymerase.
Family screening: Alive relatives were clinically and genetically screened for the mutation identified in the corresponding probands using direct automated sequencing.
Results
Case no. 1
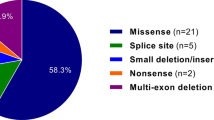
The proband is a 27-year-old male, whose father was affected by MFS and died suddenly for autopsy-proven aortic dissection at the age of 40 years. His mother died at the age of 46 for mammary carcinoma. The patient was first diagnosed with MFS in his infancy, when a prominent pectus excavatum led him to the clinical attention. Since then, several cardiological clinical and echocardiographic evaluations were repeated. His phenotype is now characterized by aortic dilation (BSA 1.98; aortic root=41 mm, ARR=1.28; the entire aorta was evaluated with echodoppler: the size and course were normal), mitral valve prolapse with mild incompetence, mild tricuspid valve incompetence and major skeletal signs, including extremely severe pectus excavatum, pes planus, arachnodactily with wrist and thumb signs, scoliosis and left lumbar gibbus; ocular and nervous systems did not show major signs. The patient showed mild myopia and thoracic striae distensae. The Ghent criteria for diagnosis were fulfilled, independently on the family history. In this patient, we identified the M425V mutation of the TGFBR2 (Figure 1). The mutation affects a highly conserved residue of the serine/threonine kinase domain of the TGFBR2, and is absent in 192 healthy controls (Figure 1). A younger sister of the proband is phenotypically and genetically nonaffected.
Genomic structure of the TGFBR2 gene: the three mutations found in present series of patients with non-FBN1-related MFS (below), and known mutations (above) previously identified in MFS type II (blue), colon–rectal cancer type 6 (pink) and LDAS (orange); chromatograms showing the three novel heterozygous mutations (arrows); alignments showing the high evolutionally conserved residues; pedigrees of the three families.
Case no. 2
The proband is a 24-year-old female, nonprofessional basketball athlete, who was found to have an aortic root dilation (BSA=1.58, aortic root=36 mm, ARR=1.31; the entire aorta was evaluated with echodoppler: the size and course were normal) and mitral valve prolapse with mild regurgitation. Scoliosis (>20 grades) was documented by multiple column X-rays other than the orthopedic evaluations, spondylolisthesis (L5-S1), arm span to height ratio=1.07, arachnodactily, positive wrist and thumb signs, pectus excavatum, pes cavus, joint hypermobility and highly arched palate, with crowding of the teeth characterized her skeletal phenotype. The proband showed multiple and severe striae distensae at the thorax and upper arm level. The ocular system was not involved. Lumbo-sacral MRI did not show dural ectasia. Her younger brother, a professional basketball athlete, did not show any aortic and mitral, as well as skeletal, abnormalities. The proband was found to carry the R460H mutation (Figure 1), while her brother was a noncarrier. In the family history, the deceased father and his sister (aunt of the proband) were described as affected by MFS with major cardiovascular and skeletal signs (documented in clinical reports). Ocular involvement was present only in the aunt. Both died for aortic root dissection in their middle age (37 and 45 years, respectively). The aunt deceased during surgery for aortic root repair. Their skeletal phenotype was characterized by scoliosis, pectus excavatum, arachnodactily (wrist and thumb signs are not detailed in the medical reports) and pes cavus, with a major severity in the father than in the aunt of the proband. No data are available on the lung and integumental system. The paternal grandfather deceased suddenly at the same age of the father. This patient fulfills the diagnostic criteria for MFS independently on the family history.
Case no. 3
The proband is a 4-year-old girl who was found to have aortic root dilation, aortic valve incompetence, pulmonary dilation and mitral valve prolapse immediately after birth, and skeletal habitus suggestive of MFS and skeletal involvement (facial appearance, dolicocephaly, joint hypermobility and arachnodactily). At present (4 years and 10 months), the BSA is 0.7, the aortic root is 31 mm (ARR=1.8) and her skeletal signs also include the pectus carinatum. She does not show major ocular signs. Facial abnormalities such as the cleft palate, bifid uvula or with broad implant, hypertelorism, craniosynostosis and congenital heart disease, in particular patent ductus arteriosum, were excluded by the referral pediatricians after discussion on differential diagnosis with Loeys–Dietz syndrome.6 Although in this girl Ghent criteria for MFS were not fulfilled, the combination of major cardiovascular signs with skeletal involvement led to molecular genetic analysis. The proband was found to carry the de novo D446N mutation (Figure 1). The follow-up will fully clarify the final phenotype associated with this mutation. Both parents and a younger brother are clinically and genetically nonaffected.
Polymorphisms: Probands 1 and 2 carried the intron 2 (+7A>G) and intron 3 (−4T>A) heterozygous polymorphisms.
Discussion
Our results support recent data demonstrating TGFBR2 gene as a candidate for FBN1-negative patients diagnosed with MFS according to Ghent's criteria.10 The recent identification of the four mutations, as well as the description of the clinical profile with major cardiovascular and skeletal involvement but no major ocular signs,4 adds a new likelihood of genotyping patients with non-FBN1 gene-related MFS. This is particularly relevant when the screening of all exon–intron boundaries of the FBN1 gene is negative in adult patients who fulfill diagnostic Ghent's criteria for MFS, as well as in patients with cardio-skeletal signs of MFS without major ocular signs, and in children who show incomplete phenotype. In both our adult patients with autosomal dominant disease, major cardiovascular and skeletal signs, and absence of major ocular signs characterized the clinical phenotype. In spite of the absence of Ectopia Lentis in the families described by Mizuguchi et al4 and in our two adult patients, the absence of major ocular signs in the MFS2 phenotype requires further cases for confirmation. In our third patient, the cardiovascular phenotype was very severe since the age of four months: an age-dependant ocular phenotype expression cannot be ruled out, and the de novo mutation does not allow comparison of the phenotype among affected relatives.
The clinical problem of differential diagnosis between MFS and MASS syndrome (myopia, mitral valve prolapse, mild aortic dilatation, and skin and skeletal manifestations (MIM #157700))12 in these patients is particularly difficult. We excluded a MASS syndrome in our patients because the skeletal system was severely affected and the father of patient #1 as well as both father and aunt of our patient #2 deceased for aortic root dissection at 40, 37 and 45 years of age, respectively. Furthermore, she has no myopia. Finally, in the 4-year-old girl, the cardiovascular involvement was severe since her first month of life.
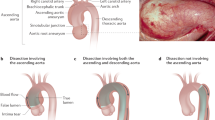
Two additional phenotypes are allelic at the same locus: the Loeys–Dietz aortic aneurysm syndrome (LDS MIM #609192) with five mutations reported to date in the TGFBR2 gene6 and the thoracic aortic aneurysms and dissection (TAAD) with two mutations of the TGFBR2 gene in four unrelated families.13 LDS is clinically characterized by ascending aortic aneurysm and dissection, hypertelorism, bifid uvula and/or cleft palate, generalized arterial tortuosity and additional multiple system signs, including craniosynostosis, structural brain abnormalities, mental retardation, congenital heart diseases and aneurysms with dissection throughout the arterial tree.6 Although tortuosity of the aorta and neck vessels was evaluated only with echodoppler imaging, and not with CT scan with contrast and 3D reconstruction, our findings do not support the diagnosis of LDS in the two adult patients because they fulfill Ghent criteria for the diagnosis of MFS and because none of them showed the craniofacial and neurocognitive traits reported in LDS none had hypertelorism, congenital heart disease, in particular patent ductus arteriosum, bifide or broad implanted uvula, and cleft palate. In case of clinical need for further progression of aortic root dilation and indication to surgery, the CT scan with contrast and 3D reconstruction will be performed. TAAD may occur as isolated phenotype in patients without MFS or may be part of MFS: analogously to the second locus for MFS (MFS2), the TAAD2 locus had been mapped to 3p24–2514 and mutations of the candidate TGFBR2 gene have been recently identified in four families with TAAD.13 In our patients #1 and 2, the major cardiovascular and skeletal signs plus the cutaneous (both) and ocular (pt #1) signs led to the diagnosis of MFS. Therefore (analogously to several other diseases that are allelic at the same locus), MFS2, LDS and TAAD are all caused by TGFBR2 gene defects: the major clinical message is that aortic aneurysms and dissections are the common marker of all known phenotypes associated to date to TGFBR2 gene defects. The clinical advantage in patients with MFS and LDS is that they can be suspected as affected on the basis of their physical phenotypes. Vice versa, patients with isolated TAAD are usually diagnosed when the first dissection occurs in the family and therefore is almost impossible to protect the proband.
The three mutated residues identified in our patients, M425V, D446N and R460H, are located in the serine/threonine kinase domain of the TGFBR2. The potential functional roles of mutations affecting serine/threonine kinase domains involve signal signaling and transduction mechanisms. Mutations in this domain have been associated with both connective tissue disorders and cancer. The fact that a germinal TGFBR2 mutation T315M has been associated with certain types of cancer (colon cancer type 6 (MIM+190182)15) raises unexpected problems on the potential additional risk of cancer in MFS patients carriers of TGFBR2 mutations. Whether patients with cancer, carriers of germinal TGFBR2 mutations, may present Marfan-like traits, or vice versa, is unknown since most studies have been focused on somatic mutations. The mutation R460H has been reported in one of the families with TAAD described by Pannu et al.14 Patients of this family did not meet the diagnostic criteria of MFS. This observation confirms the phenotypic heterogeneity of diseases caused by TGFBR2 gene defects, and joins a similar observation for many other monogenic disorders, especially involving the cardiovascular system (ie LMNA gene, MIN *150330).
Our results support prior data showing that TGFBR2 gene mutations are associated with MFS without major ocular signs. Accordingly, patients with major cardio-skeletal phenotype who are diagnosed with MFS on the basis of Ghent criteria and proven not to carry defects in exons and flanking regions of the FBN1 gene could benefit from rapid investigation of the TGFBR2 gene. Available scientific data, however, are still nonsufficient to exclude from TGFBR2 gene-screening patients with ocular signs, both major and involvement.
GeneBank accession numbers: TGFBR2 cDNA, NM_003242; TGFBR2 coding region, NT_022517.
Accession codes
References
Loeys B, De Backer J, Van Acker P et al: Comprehensive molecular screening of the FBN1 gene favors locus homogeneity of classical Marfan syndrome. Hum Mutat 2004; 24: 140–146.
Collod G, Babron MC, Jondeau G et al: A second locus for Marfan syndrome maps to chromosome 3p2.4–p25. Nat Genet 1994; 8: 264–268.
Boileau C, Jondeau G, Babron MC et al: Autosomal dominant Marfan-like connective-tissue disorder with aortic dilation and skeletal anomalies not linked to the fibrillin gene. Am J Hum Genet 1993; 53: 46–54.
Mizuguchi T, Collod-Beroud G, Akiyama T et al: Heterozygous TGFBR2 mutations in Marfan syndrome. Nat Genet 2004; 36: 855–860.
Mathew S, Murty VVVS, Cheifetz S et al: Transforming growth factor receptor gene TGFBR2 maps to human chromosome band 3p22. Genomics 1994; 20: 114–115.
Loeys BL, Chen J, Neptune ER et al: A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet 2005; 37: 275–281.
Annes JP, Munger JS, Rifkin DB : Making sense of latent TGFbeta activation. J Cell Sci 2003; 116: 217–224.
ten Dijke P, Hill CS : New insights into TGF-beta-Smad signaling. Trends Biochem Sci 2004; 29: 265–273.
Roman MJ, Devereux RB, Kramer-Fox R, Spitzer MC : Comparison of cardiovascular and skeletal features of primary mitral valve prolapse and Marfan syndrome. Am J Cardiol 1989; 63: 317–321.
De Paepe A, Devereux R, Dietz H, Hennekam R, Pyeritz R : Revised diagnostic criteria for the Marfan Syndrome. Am J Med Genet 1996; 62: 417–426.
Arbustini E, Grasso M, Ansaldi S et al: Identification of 62 novel and 12 known mutations in FBN1 gene in 81 unrelated probands with Marfan syndrome and other fibrillinopathies. Hum Mutat 2005-0059.R2,www3.interscience.willey.com/cgi-bin/Jabout/38515/MutationsBrief2005.html.
Dietz HC, McIntosh I, Sakai LY et al: Four novel FBN1 mutations: significance for mutant transcript level and EGF-like domain calcium binding in the pathogenesis of Marfan syndrome. Genomics 1993; 17: 468–475.
Pannu H, Fadulu VT, Chang J et al: Mutations in transforming growth factor-beta receptor type II cause familial thoracic aortic aneurysms and dissections. Circulation 2005; 112: 513–520.
Hasham SN, Willing MC, Guo DC et al: Mapping a locus for familial thoracic aortic aneurysms and dissections (TAAD2) to 3p24–25. Circulation 2003; 107: 3184–3190.
Lu SL, Kawabata M, Imamura T et al: HNPCC associated with germline mutation in the TGF-β type II receptor gene. Nat Genet 1998; 19: 17–18.
Acknowledgements
We thank the patients and families for their participation. This work was supported by Grants from National Ministry of Health, Grants ‘Ricerche Finalizzate’ 1997–2000, ‘Ricerche Correnti’ 2001–2004, IRCCS Policlinico S Matteo, Pavia (Italy). Grant for Marfan Sindrome, Fondazione Banca Regionale Europea, Milano.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Disabella, E., Grasso, M., Marziliano, N. et al. Two novel and one known mutation of the TGFBR2 gene in Marfan syndrome not associated with FBN1 gene defects. Eur J Hum Genet 14, 34–38 (2006). https://doi.org/10.1038/sj.ejhg.5201502
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201502
Keywords
This article is cited by
-
Cellular and molecular mechanisms of thoracic aortic aneurysms
Nature Reviews Cardiology (2009)
-
Prevalence data on all Ghent features in a cross-sectional study of 87 adults with proven Marfan syndrome
European Journal of Human Genetics (2009)
-
A new sporadic case of early-onset Loeys-Dietz syndrome due to the recurrent mutation p.R528C in theTGFBR2 gene substantiates interindividual clinical variability
Journal of Applied Genetics (2009)
-
Evaluation of resequencing on number of tag SNPs of 13 atherosclerosis-related genes in Thai population
Journal of Human Genetics (2008)
-
Absence of TGFBR2 mutations in patients with spontaneous spinal CSF leaks and intracranial hypotension
The Journal of Headache and Pain (2008)