Abstract
Overgrowth is rarely associated with chromosomal imbalances. Here we report on four children from two unrelated families presenting with overgrowth and a terminal duplication of the long arm of chromosome 15 diagnosed using cytogenetic and FISH studies. In both cases, chromosome analysis of the parents showed a balanced translocation involving 15q26.1-qter. Molecular and cytogenetic studies showed three copies of the insulin-like growth factor 1 receptor (IGF1R) gene. This finding suggests that overgrowth observed in our patients might be causally related to a dosage effect of the IGF1R gene, in contrast to severe growth retardation observed in patients with terminal deletion of 15q. The present observation emphasises the importance of chromosome analysis in patients with overgrowth and mental retardation. Moreover, it further delineates a specific phenotype related to trisomy 15q26.1-qter with macrosomia at birth, overgrowth, macrocephaly and mild developmental delay being the major clinical features.
Similar content being viewed by others
Introduction
Although overgrowth is observed in several genetic syndromes, its molecular basis is unravelled only in a small number of them including Beckwith-Wiedemann syndrome,1 Simpson-Golabi-Behmel syndrome2 and more recently Sotos syndrome.3 Overgrowth is rarely associated with chromosome imbalances. Presumably, a detailed study of these unusual cases can give us a clue for candidate chromosome regions or genes involved in the pathogenesis of overgrowth. Four partial trisomies including growth-related genes have been seen in overgrowth syndromes, including trisomy 4p16.3 and the FGFR3 gene,4 trisomy 11p15.5 and the IGF2 gene,5 trisomy 20p11.2 and the SSTR4 gene6 and trisomy 15q25-qter.7 Overgrowth has been reported as well in association with tetrasomy 15q25-qter.8 Observations of distal deletions of chromosome 15q including the IGF1R gene suggest that growth failure might be related to IGF1R hemizygosity.9 Here we report on four patients with overgrowth and trisomy 15q26.1-qter including the IGF1R gene.
Clinical reports
Family 1
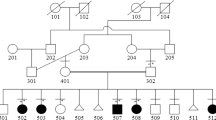
At least four patients were affected in family 1 (Figure 1a). Individual IV-5 was referred to us for genetic counselling because her daughter (V-4) and her niece (V-1) presented with mild developmental delay and macrocephaly. She had a sister and three brothers, two of which (IV-3 and IV-6) had mild mental retardation with IQ between 50 and 70, slight dysmorphism including an elongated face and macrocephaly. Their height was reported to be between 1.75 and 1.80 m. Their karyotypes were initially considered as normal and fragile X syndrome was excluded. New clinical evaluation was denied.
Her daughter (V-4) was born at term after an uneventful pregnancy. At birth, length was 51 cm (+1 SD), weight was 4000 g (+2 SD) and occipito-frontal circumference (OFC) was 37 cm (+2 SD). Developmental delay was noted at 10 months, she began to walk at 20 months and was able to speak at 4 years. At 7½ years, her height was 1.25 m (+1 SD), weight was 29 kg (+2 SD) and OFC was 57.2 cm (+4 SD). She interacted well with her environment but had learning difficulties. She was mildly dysmorphic with a large head and frontal bossing, an elongated face, slight midface retraction, a pointed chin, and small low-set ears. Wrists and ankles were large and thumbs were short. Cardiac examination, renal ultrasound and neuroimaging were normal. Ophthalmologic examination showed a visual acuity of 5/10 bilaterally.
Her niece (V-1) was born at term after an uneventful pregnancy. At birth, length was 53 cm (+3 SD), weight was 3300 g (mean) and OFC 36 cm (+1 SD). Hypotonia was noticed in the first month of life. She was admitted at 4 months of life for progressive overgrowth with macrocephaly. She sat at age 8½ months, walked at age 2 years, and presented speech delay. At 6 years of age, her height was 127 cm (+3.5 SD), weight was 23 kg (+2 SD) and OFC was 57 cm (+4 SD). Dysmorphic features included a long triangular face, high, large and flat forehead, downslanting palpebral fissures, slight enophtalmia, small round and low set ears, and a large chin (Figure 2a). Hyperlaxity of fingers were noted, with long and slender hands and proximal implantation of the thumbs. There was a moderate psychomotor delay. Ophthalmologic and cardiac examination, neuroimaging and endocrine studies including growth hormone and 17 OH progesterone were normal. Diagnosis of Sotos syndrome was initially proposed. Her father's occipito-frontal circumference was 60 cm.
Family 2
Both siblings of family 2 were affected (Figure 1b). The parents originated from Antilles. The height and occipito-frontal circumference were respectively 163 and 57.5 cm in the mother and 190 and 58 cm in the father. The family history on the mother's side was unremarkable. The father had one half-sister, one half-brother and one brother. All of them presented with tall stature. The half-brother had developmental delay.
The pregnancy of the older sibling was uneventful and the baby was born at 38 weeks of gestation (WG). At birth, length was 49 cm (mean), weight 3590 g (+1.5 SD) and OFC 36 cm (+2 SD). She walked at 1½ years of age and was able to speak at 2½ years of age. She was referred to us at 6 years of age for overgrowth and psychomotor retardation. At this time, her height was 137 cm (+4 SD), weight was 32.4 kg (+4 SD) and OFC was 61 cm (+5 SD). Dysmorphic features including a long triangular face with high forehead, downslanting palpebral fissures, small round and low set ears, a flat nasal bridge, and a normal palate were noted (Figure 2b). Clinical examination of the extremities showed clinodactyly of the fifth left finger and pes planus and valgus. Hyperlaxity and a lumbar lordosis were also noticed. She was mildly retarded and needed special education at school. X-rays examination showed large epiphyses. Cardiac and renal ultrasound as well as neuroimaging showed no visceral malformations. Ophthalmologic examination showed a bilateral papillary coloboma. Endocrine studies including growth hormone and 17 OH progesterone were normal. IGF1 concentration was 166 ng/ml (N: 126–260). Metabolic studies including aminoacid and organic acid chromatography, isoelectric focusing of serum transferrin, very long chain fatty acids assay, mucopolysaccharides and oligosaccharides revealed normal results.
The second sibling was born at 37 WG after an uneventful pregnancy. At birth, length was 52 cm (+2.5 SD), weight 4120 g (+4 SD) and OFC 39 cm (+5 SD). She was hypotonic and suffered neonatal distress due to a pneumothorax. She was referred to us at 11 months of age for overgrowth and psychomotor retardation. At this time, her length was 80 cm (+3.5 SD), weight was 13.6 kg (+4 SD) and OFC was 51 cm (+4 SD). Dysmorphic features included a long face with high forehead and a prominent metopic suture, epicanthal folds, small, round and low set ears, a very flat nose and nasal bridge, and a normal palate (Figure 2c). Hyperlaxity and clinodactyly of the fifth finger were noticed. She sat at 11 months of age. Ophthalmologic examination showed hypermetropia. Ultrasound cardiac scan showed ductus arteriosus. Cerebral imaging and renal ultrasound were normal. Similar endocrine and metabolic studies performed in her sister were normal.
Cytogenetic and molecular studies
Family 1
High resolution chromosome analysis using R and G banding techniques reveals a balanced translocation between chromosomes 15 and 20 in individual IV-5 (Figure 3a). Chromosome painting with whole chromosome painting WCP15 and WCP20 showed additional material originating from chromosome 15 on chromosome 20p, but no material originating from chromosome 20 on chromosome 15q (Figure 3b). FISH analysis with subtelomeric probes of chromosome 20p (PAC 82O2) and 15q (PAC 124O5) confirmed that the translocation was reciprocal (not shown). This suggest that the size of the translocated 20p segment is too small to be detected using chromosome painting. Comparative genomic hybridization (CGH)10,11 was performed in V-1 and confirmed the excess of material from 15q26.1-qter but did not show evidence of loss of 20p material (Figure 3d). Further FISH analyses with various probes of chromosome 15q9,12,13,15 defined that the size of the 15q duplication ranged between 6 and 12 Mb (Table 1). In particular, YAC 802B4 was not duplicated. The karyotype of IV-5 was therefore 46,XX,t(15;20)(q26.1;p13). The same translocation was present in her healthy brother and in her mother. All affected patients are carriers of the same unbalanced karyotype with a der(20) resulting in trisomy 15q26.1-qter and a very subtle monosomy 20pter (Figure 3c).
Family 1: chromosome analysis and FISH studies. (a) Balanced translocation t(15;20)(q26.1;p13). Abnormal chromosomes are on the right. (b) Whole painting of chromosome 15 (grey) and 20 (white) of the balanced translocation. The small size of the translocated 20p segment prevents its visualization. (c) Unbalanced translocation der(20)t(15;20). (d) CGH showing trisomy 15q26.1qter. The monosomy 20p is not detected.
Family 2
High-resolution chromosome analysis of the older sibling was performed on cultured peripheral blood lymphocytes using R and G banding techniques. These studies showed additional chromosome material of unknown origin on the long arm of chromosome 13 in all cells examined (Figure 4c). Fluorescence in situ hybridisation (FISH) analysis with a chromosome 15 painting probe was performed and confirmed that the additional material originated from chromosome 15 (Figure 4b). FISH analysis with subtelomeric probes of chromosome 13q (PAC 163C9) and 15q (PAC 124O5) showed one signal of chromosome 13q and three signals of chromosome 15q (not shown). CGH confirmed the excess of material from bands 15q26.1-qter, but did not show evidence of loss of 13q material (Figure 4d). Chromosome analysis of the parents showed that this unbalanced rearrangement resulted from the segregation of a paternal balanced translocation. The same unbalanced karyotype was found in the younger sister. Further FISH analyses with various probes of chromosome 15q12,13 defined that the size of the 15q duplication ranged between 6 and 12 Mb since YAC 802B4 was not duplicated (Table 1). Similarly, we showed that the size of the 13q deletion was small, ranging from 80 Kb to 2.5 Mb. The karyotype of the father was therefore 46,XY,t(13;15)(q34;q26.1) (Figure 4a) and the children karyotypes were 46,XX,der(13)t(13;15)(q34;q26.1)pat.
Family 2: chromosome analysis and FISH studies. (a) Balanced translocation t(13;15)(q34;q26.1). Abnormal chromosomes are on the right. (b) Whole painting of chromosome 13 (grey) and 15 (white) of the balanced translocation. The size of the translocated 13q segment prevents its visualization. (c) Unbalanced translocation der(13)t(13;15). (d) CGH showing trisomy 15q26.1qter. The monosomy 13q is not detected.
IGF1R gene
FISH studies using an IGF1R probe (BAC 654A16) showed one signal on each 15q and one on 20p for proband (V-1) of family 1 (Figure 5a), and one signal on each 15q and one on 13q for proband of family 2 (Figure 5b). Molecular studies using an intragenic polymorphic marker of the IGF1R gene performed in the proband of family 2 were fully informative. They showed three alleles of the IGF1R gene, two of which being of paternal origin (Figure 5c) as expected. The polymorphic marker was identified from the published genomic sequence of IGF1R (http://genome.ucsc.edu/).
IGF1R: molecular and FISH studies. (a) FISH experiments with BAC 654A16 containing the IGF1R gene showed an additional signal at the end of the der(20) in proband of family 1. (b) FISH experiments with BAC 654A16 showed an additional signal at the end of the der(13) in proband of family 2. (c) Proband of family 2: an intragenic polymorphic microsatellite of the IGF1R gene (primers 5′-GAGTATGAACAGGTGGCATTAG-3′ and 5′-ACATAGCCTTCATCCAACTTCC-3′) showed three alleles of the gene of which two are of paternal origin. F: Father, M: Mother, P: Proband.
Discussion
Here, we report on two families with overgrowth and partial trisomy of chromosome 15q26.1-qter including the IGF1 receptor gene. In both cases, the associated monosomy is small in size.
The phenotype of distal trisomy 15q, with a breakpoint being usually between 15q15 and 15q22, is well delineated. It includes mental retardation, growth deficiency, microdolichocephaly, peculiar facial features, skeletal abnormalities, and cardiovascular defects.14 Most of these cases result from the malsegregation of a parental translocation. In a few cases, the trisomy is confined to chromosome 15q25-qter and a different phenotype is observed, including macrosomia at birth, overgrowth, macrocephaly and craniosynostosis.7 Also, mosaicism for an unstable interstitial duplication of chromosome band 15q25 was recently described as a susceptibility factor for panic and phobic disorders.15 The segment duplicated on 15q in our patients is distal to this region as shown using YAC 802B4 which is located near its distal border.
Reports of six relevant patients with 15q25 or 26-qter trisomy and of the four present cases are summarised in Table 2.16,1718,7 Patients IV-3 et IV-6 of family 1 were not included because there was not enough of clinical data available. One report with cloverleaf skull anomaly was also excluded because of insufficient data.19 From this analysis, it appears that 8 out of 10 cases (six families) are familial. All cases result from an unbalanced segregation of a parental translocation with a breakpoint in 15q25 or q26 and the breakpoint on the other chromosome located in the terminal band (except in one case). Interestingly, breakpoints found in our family 2 are similar to those found in the family reported by Zollino.7 Seven out of 10 cases presented with overgrowth between +2 SD and +6 DS. None of the patients presented with a size below the normal limit. Interfamilial variability in overgrowth is observed. There is also intrafamilial variability, as shown in our family 1 and the family reported by Zollino.7 The severity of overgrowth does not seem linked to the parental origin of the translocation. Indeed, overgrowth was noted in 3 out of 5 cases of paternal origin, and 4 out of 5 cases of maternal origin. Macrosomia was present at birth in 7 out of 8 informative patients, birth length ranging from 49 to 59 cm and weight from 3.3 to 5 kg. If growth velocity seems to decelerate with age in some observations,7 tall stature is still present in adulthood in others.3 The adult size varies from 1.78 to 2.10 m. Macrocephaly was even more frequent than overgrowth and was ⩾4 SD in 8 out of 10 cases. In one patient, macrocephaly is present without overgrowth.18 Craniosynostosis was associated in 3 out of 10 cases. Various cardiac defects or kidney malformations were found respectively in 3 out of 10 and 2 out of 9 cases, and scoliosis in 4 out of 10 cases. Psychomotor delay was a consistent feature and ranged from mild to severe. Distinctive facies found in most patients include a long and triangular face, downslanting palpebral fissures, and abnormal low-set ears. Finger abnormalities were often noted, such as arachnodactyly, flexed or tapering fingers.
It is well known that the insulin-like growth factors and their receptors are implicated in pre- and postnatal development.20 The IGF1R is a transmembrane heterotetramer, which mediates the mitogenic effect of IGF1 and IGF2. Distal deletions of chromosome 15q are associated with severe growth retardation.9,21,22,2324 It has been suggested that growth failure might be related to IGF1R hemizygosity. On the contrary, in the present observations, overgrowth might well be related to a dosage excess of IGF1R gene.
Also, a gene for macrocephaly is likely to be located in this region. Interestingly a large family with autosomal recessive syndrome of macrocephaly, multiple epiphyseal dysplasia and distinctive facies was recently reported and a gene locus was identified in 15q26.24 The aggrecan gene (AGC1), a chondroitin sulphate proteoglycan that was a possible candidate gene, was excluded.25
In conclusion, we report on four further cases of overgrowth associated with trisomy 15q26.1-qter including the IGF1R locus. The present report gives further evidence for a specific phenotype related to trisomy 15q25 or 26-qter with macrosomia at birth, overgrowth, macrocephaly and mild developmental delay being the major clinical features. We suggest that overgrowth might be causally related to a dosage excess of IGF1R gene. Interestingly, overgrowth has also been reported in patients with tetrasomy of chromosome 15q25-qter,8 as well as in some patients with larger trisomy 15q22qter.26,27 Mainly, this report emphasizes the importance of chromosome analysis in the investigation of patients with overgrowth and mental retardation. Further studies of similar cases will provide a valuable opportunity for elucidating the molecular basis of overgrowth.
References
Li M, Squire JA, Weksberg R . Molecular genetics of Wiedemann-Beckwith syndrome Am J Med Genet 1998 79: 253–259
Neri G, Gurrieri F, Zanni G, Lin A . Clinical and molecular aspects of the Simpson-Golabi-Behmel syndrome Am J Med Genet 1998 79: 279–283
Kurotaki N, Imaizumi K, Harada N et al. Haploinsufficiency of NSD1 causes Sotos syndrome Nat Genet 2002 30: 365–366
Partington MW, Fagan K, Soubjaki V, Turner G . Translocations involving 4p16.3 in three families: deletion causing the Pitt-Rogers-Danks syndrome and duplication resulting in a new overgrowth syndrome J Med Genet 1997 34: 719–728
Elliott M, Maher ER . Syndrome of the month: Beckwith-Wiedemann syndrome J Med Genet 1994 31: 560–564
Faivre L, Viot G, Prieur M et al. Apparent Sotos syndrome (cerebral gigantism) in a child with trisomy 20p11.2-p12.1 mosaicism Am J Med Genet 2000 91: 273–276
Zollino M, Tiziano F, Di Stefano C, Neri G . Partial duplication of the long arm of chromosome 15: Confirmation of a causative role in craniosynostosis and definition of a 15q25-qter trisomy syndrome Am J Med Genet 1999 87: 391–394
Rowe AG, Abrams L, Qu Y, Chen E, Cotter PD . Tetrasomy 15q25→qter: Cytogenetic and molecular characterization of an analphoid supernumerary marker chromosome Am J Med Genet 2000 93: 393–398
Peoples R, Milatovich A, Francke U . Hemizygosity at the insulin-like growth factor I receptor (IGF1R) locus and growth failure in the ring chromosome 15 syndrome Cytogenet Cell Genet 1995 70: 228–234
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, Pinkel D . Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors Science 1992 258: 818–821
Lapierre JM, Cacheux V, Da Silva F, Collot N, Hervy N, Wiss J, Tachdjian G . Comparative genomic hybridization: technical development and cytogenetic aspects for routine use in clinical laboratories Ann Genet 1998 41: 56–62
Kingsley K, Wirth J, van der Maarel S, Freier S, Ropers HH, Haaf T . Complex FISH probes for the subtelomeric regions of all human chromosomes: comparative hybridization of CEPH YACs to chromosomes of the Old World monkey Presbytis cristata and great apes Cytogenet Cell Genet 1997 78: 12–19
Knight SJ, Lese CM, Precht KS et al. An optimized set of human telomere clones for studying telomere integrity and architecture Am J Hum Genet 2000 67: 320–332
Chandler K, Schrander-Strumpel CTRM, Engelen J, Theunissen P, Fryns JP . Partial trisomy 15q: report of a patient and literature review Genet Couns 1997 8: 91–97
Gratacos M, Nadal M, Martin-Santos R et al. A polymorphic genomic duplication on human chromosome 15 is a susceptibility factor for panic and phobic disorders Cell 2001 106: 367–379
Kristoffersson ULF, Bergwall B . Partial trisomy 15(q25qter) in two brothers Hereditas 1984 100: 7–10
Van Allen MI, Siegel-Bartelt J, Feigenbaum A, Teshima IE . Craniosynostosis associated with partial duplication of 15q and deletion 2q Am J Med Genet 1992 43: 688–692
Wieczorek D, Engels H, Viersbach R, Henke B, Schwanitz G, Passarge E . Analysis of a familial three way translocation involving chromosomes 3q, 6q, and 15q by high resolution banding and fluorescent in situ hybridisation (FISH) shows to different unbalanced karyotypes in sibs J Med Genet 1998 35: 545–553
Pedersen C . Partial trisomy 15 as a result of an unbalanced 12/15 translocation in a patient with a cloverleaf skull anomaly Clin Genet 1976 9: 378–380
Baker J, Lui JP, Robertson EJ, Efstratiadis A . Role of insulin-like growth factors in embryonic and postnatal growth Cell 1993 75: 73–82
de Lacerda L, Carvalho JA, Stannard B et al. In vitro and in vivo responses to short-term recombinant human insulin-like growth factor-1 (IGF-1) in a severely growth-retarded girl with a ring chromosome 15 and deletion of a single allele for the type I IGF receptor gene Clin Endocrinol 1999 51: 541–550
Siebler T, Lopaczynski W, Terry CL et al. Insulin-like growth factor I receptor expression and function in fibroblasts from two patients with deletion of the distal long arm of chromosome 15. J Clin Endocrinol Metab 1995 80: 3447–3457
Tamura T, Tohma T, Ohta T et al. Ring chromosome 15 involving deletion of the insulin-like growth factor 1 receptor gene in a patient with features of Silver-Russell syndrome Clin Dysmorphol 1993 2: 106–113
Roback EW, Barakat AJ, Dev VJ, Mbikay M, Chrétien M, Butler MG . An infant with deletion of the distal long arm of chromosome 15 (q26.1→qter) and loss of insulin-like growth factor 1 receptor gene Am J Med Genet 1991 38: 74–79
Bayoumi R, Saar K, Lee YA et al. Localisation of a gene for an autosomal recessive syndrome of macrocephaly, multiple epiphyseal dysplasia, and distinctive facies to chromosome 15q26 J Med Genet 2001 38: 369–373
Gregoire MJ, Boue J, Junien C, Pernot C, Gilgenkrantz S, Zergollern L . Duplication of 15q22→15qter and its phenotypic expression Hum Genet 1981 59: 429–433
Goldstein DJ, Ward RE, Nichols WC, Palmer CG . Familial t(8;15)(p23.3;q22.3): report of two cases with dup(15)(q22.3→ qter) J Med Genet 1987 24: 684–687
Acknowledgements
We thank Pr B Le Marec for his clinical expertise and help with family 1. M Picq, C Ozilou and JM Lapierre are acknowledged for their excellent technical work and Y Deris for his photographic assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Faivre, L., Gosset, P., Cormier-Daire, V. et al. Overgrowth and trisomy 15q26.1-qter including the IGF1 receptor gene: report of two families and review of the literature. Eur J Hum Genet 10, 699–706 (2002). https://doi.org/10.1038/sj.ejhg.5200879
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200879
Keywords
This article is cited by
-
Genome wide noninvasive prenatal testing detects microduplication of the distal end of chromosome 15 in a fetus: a case report
Molecular Cytogenetics (2022)
-
Molecular characterization and evaluation of complex rearrangements in a case of ring chromosome 15
Molecular Cytogenetics (2017)
-
Chromosomal microarray analysis in clinical evaluation of neurodevelopmental disorders-reporting a novel deletion of SETDB1 and illustration of counseling challenge
Pediatric Research (2016)
-
Copy-number variation associated with congenital anomalies of the kidney and urinary tract
Pediatric Nephrology (2015)
-
Array-based comparative genomic hybridization identifies a high frequency of copy number variations in patients with syndromic overgrowth
European Journal of Human Genetics (2010)