Abstract
Usher syndrome type 3 (USH3) is an autosomal recessive disorder characterised by the association of post-lingual progressive hearing loss, progressive visual loss due to retinitis pigmentosa and variable presence of vestibular dysfunction. Because the previously defined transcripts do not account for all USH3 cases, we performed further analysis and revealed the presence of additional exons embedded in longer human and mouse USH3A transcripts and three novel USH3A mutations. Expression of Ush3a transcripts was localised by whole mount in situ hybridisation to cochlear hair cells and spiral ganglion cells. The full length USH3A transcript encodes clarin-1, a four-transmembrane-domain protein, which defines a novel vertebrate-specific family of three paralogues. Limited sequence homology to stargazin, a cerebellar synapse four-transmembrane-domain protein, suggests a role for clarin-1 in hair cell and photoreceptor cell synapses, as well as a common pathophysiological pathway for different Usher syndromes.
Similar content being viewed by others
Introduction
Combined deafness and blindness in adults is most frequently caused by Usher syndromes (USH). These account for more than half of the dual sensory deficit, with a prevalence of 1/10 000 in the age group of 30–50.1 Sensorineural hearing loss and retinitis pigmentosa (RP) characterise this group of autosomal recessive hereditary disorders. Three USH types are distinguished clinically. Postlingual progressive hearing loss and variable vestibular function characterise USH3 (MIM 276902).2 Progressive RP with variable age of onset occurs in all USH types. While USH1 and USH2 map to at least 10 distinct loci,3,4,5,6,7,8,9,10,11 only one locus for USH3 has been reported so far.12,13 Four of the USH1 and the USH2A genes have been identified. Mutations in the unconventional myosin MYO7A cause USH1B (MIM 27690314) and in two unique cases also atypical USH phenotype similar to that of USH3.15 Mutations in the PDZ-domain-containing protein, harmonin, cause USH1C (MIM 276904)16,17. PDZ-domains are protein–protein interaction modules that allow the binding to and clustering of specific membrane-associated proteins, such as receptors and ion channels.18 Cadherin23 and protocadherin15 mutations have recently been shown to underlie USH1D (MIM 601067) and USH1F (MIM 602083), respectively.19,20,21,22 Most cadherins are integral membrane glycoproteins that mediate the calcium-dependent formation of cell–cell adhesion,23 while protocadherins are thought to be involved in a variety of functions, including neural development, neural circuit formation and formation of the synapse.24 Finally, mutations in Usherin, a gene that encodes a basement membrane protein, cause USH2A (MIM276901)25.
Joensuu et al.26 have recently identified three mutations (a missense, a nonsense and a 3 bp deletion) in exons 2 and 3 of a newly identified gene that maps to 3q24, approximately 100 kb proximal to the previously defined USH3A linkage interval in 3q25.1. These mutations established this gene as the USH3A gene. Northern blot analysis and reverse-transcription PCR indicated the expression of the USH3A gene in several tissues, including retina. Two predicted transmembrane domains were identified in its deduced protein product, which at the time did not show similarity to any known protein.26
In this study, we characterise new human and mouse USH3A transcripts (AF495717; AF495718; AF495719 and AF495720), identify three additional USH3 mutations, analyse the USH3A expression pattern and by in situ hybridisation localise transcripts of this gene to mouse cochlear hair cells and spiral ganglion cells. We redefine the USH3A protein, name it clarin-1 and affiliate it to a new four transmembrane domain (4TM) vertebrate-specific protein family. Furthermore, based on sequence similarities to stargazin, a well-studied member of this hyperfamily, we suggest a role for clarin-1 in the hair cells synaptic junctions.
Materials and Methods
USH3 families and controls
Blood samples were drawn by venipuncture after obtaining informed consent in accordance with the guidelines of the Tel Aviv University Helsinki Committee. Three of the Jewish Ashkenazi USH3 families included in this study, were referred to us through the Center for Deaf–Blind Persons in Tel Aviv as part of a larger study on the genetics of Usher syndromes in Israel. The fourth Jewish Ashkenazi USH3 family living in the US has one affected and two unaffected siblings whose grandparents originate from Eastern Europe. The family was ascertained by one of us (JGF). The USH3 Jewish Yemenite and the Spanish families have been described elsewhere.27,28 Clinical diagnosis of affected members in these families is compatible with the USH3 phenotype and haplotype segregation analysis does not exclude linkage to the USH3A locus. Control DNA samples from unrelated Jewish Ashkenazi and Yemenite individuals, were provided by the National Laboratory for the Genetics of Israeli Populations. A total of 51 grandparents in families from the Centre d'Etude du Polymorphisme Humain (CEPH) were also studied as controls.
RNA expression
The expression of Ush3a was determined by in situ hybridisation and reverse-transcription (RT)–PCR in mice at embryonic day (E) 16, postnatal day (P)0, 5, 10, 15, 20, and 30. For RT–PCR, cochleae were dissected out of the temporal bones. Total RNA was prepared from cochleae with TRI Reagent (Sigma). Genomic DNA was removed from all RNA samples using DNA-free (Ambion). Total RNA was purified using phenol-chloroform extraction (PCI; Gibco) and phase lock gel tubes (Eppendorf), followed by an isopropanol precipitation. RT reactions were performed using Expand Reverse Transcriptase (Roche) with Homo-Oligomeric DNA d(T)12-18 and Random Hexamer (Amersham Pharmacia Biotech). cDNA was amplified from the RT reactions using primers (F) 5′-GGTCCAAGCCATCCCCGTA-3′ AND (R) 5′- CCTCCTGCTTCTGTTATTTTCC-3′. As a control, primers spanning the last intron of Myo6 (F) 5′-CTGGTGGTATGCCCATTTTGA-3′ and (R) 5′-TCGCTTTGCATAAGGCATTTCTA-3′ were used.29
For in situ hybridisation, cochleae were dissected from surrounding tissue and fixation was performed by immersion in 4% paraformaldehyde in PBS for 12–16 h at 4°C. The samples were processed without decalcification until P10. From P10 onwards, the samples were dehydrated in graded concentrations of methanol, from 25–100%, followed by decalcification (to facilitate dissection) in 8% formic acid in 100% methanol for up to 4 days. We generated digoxygenin-labelled cRNA probes in a standard transcription reaction (Roche Molecular Biochemicals). We performed whole mount in situ hybridisation as described,30 with modifications. Finally, we visualised in situ reaction product by cryoprotecting whole cochleae with sucrose, embedding the tissue in OCT, and cryostat sectioning (10 μm). Antisense and sense probes were in vitro transcribed from a linearised vector, pPCR-Script AMP SK(+) cloning vector (Stratagene), containing a partial mouse Ush3a cDNA (corresponding to exons 2 and 3) by using either T3 or T7 RNA polymerase. No in situ hybridisation signal was detected with the control sense RNA probe. Our use of animals was approved by the Tel Aviv University Animal Care and Use Committee (11-00-65).
Mutation detection and analysis
Using the primers detailed in Table 2 we amplified fragments encompassing USH3A exons from genomic DNA templates of USH3 patients and their family members. PCR products were gel-purified, sequenced using dye terminators of Big-Dyes kits (Perkin-Elmer/Applied Biosystems) and analysed on an ABI 3700 sequencer (Perkin-Elmer/Applied Biosystems). Sequence comparisons were performed by the use of Sequencher 4.1 software from GeneCodes Corporation. The Jewish Ashkenazi N48K mutation was screened by StuI digestion (16 h at 37°C) followed by agarose (2%) gel analysis. The 23 bp deletion (found in patients of Yemenite origin) was screened by PCR amplification of a 227 bp fragment followed by FMC's MetaPhor gel (4%) electrophoresis. The Y63X mutation (found in a patient of Spanish origin) was screened by SSCP; a 207-bp fragment of exon 0 was PCR amplified by primers ex0F2 (5-ATCAAAGCCACTGTCCTCTG-3) and ex0R (5-CTGGGAAGAGTCTGCCTAAA-3), and the alleles were separated on 0.7×MDE gels, for 18 h at 5 W, at room temperature and were visualised by silver staining.31
Computational analyses and sequence annotation
Genomic sequences were retrieved from four NCBI (National Center for Biotechnology Information) Human clones: AC020636, AC036109, AC011103, AC078816. Fragments were assembled by Sequencher 4.1 software from GeneCodes Corporation. Complete sequence annotation was performed using the GESTALT workbench.32 Sequence similarity searches performed by the use of the BLAST33 and Blimps34 programs. Sequences were aligned using the BlockMaker34 and MACAW35 programs; PHYLIP ProtDist36 PHYLIP – Phylogeny Inference Package (Version 3.2 Cladistics 5: 164–166) and CLUSTAL W programs37 were used to compute trees and their significance (bootstrap) values. TM regions were predicted using the PHDhtm38 and TMHMM programs.39 Primers were designed using the Oligo primer analysis software (http://medprobe.com/is/oligo/html).
Results
Genomic analysis and transcripts
Since no mutations were identified within the described USH3A transcripts26 in patients from other USH3 families, we assumed that there might be additional uncharacterised exons of this gene. Therefore, we assembled and reconstructed a 500 kb genomic interval spanning the published USH3A gene and the previous USH3A linkage interval, defined by D3S3413 and D3S1279 (Figure 1A). Based on gene predictions and alignment with human and mouse ESTs, we designed primers that allowed the amplification of a new human USH3A transcript. This longer transcript begins with a newly identified exon 0, continues with exon 2, has exons 3 and 4 transcribed together with their intervening intron, but does not include the previously identified exons 1 and 1b26 (Figure 1B).
(A) Schematic representation of physical and transcript maps covering the USH3A region comprised of partly overlapping NCBI (National Center for Biotechnology Information) clones. Polymorphic markers and sequence-tagged sites are indicated on the bar below the chromosome. Pink markers define the borders of the old USH3A linkage interval. The red marker is the newly identified polymorphic intragenic microsatellite located within the USH3A's 3′-UTR. Colored arrows below the clones indicate the position and orientation of the UCRP pseudogene (light blue), the USH3A (red) and partially overlapping USHARF (green) genes. Below these arrows is an enlarged view of an approximate 66 kb segment, showing the genomic structure of the USH3A gene and the first two exons of the USHARF gene. Blue boxes and lines indicate coding exons, while grey boxes and lines indicate non-coding regions. Light blue and green lines above the exons indicate overlapping regions between USH3A and, respectively, UCRP and USHARF. (B) Alternative transcripts of the USH3A gene. The first two pale colored transcripts were defined previously by Joensuu et al. (2001).26 (C) The first two exons of the USHARF gene. (D) Genomic structure of the mouse Ush3a gene and its three newly defined in frame alternative transcripts. Marked by black arrows is the position of the primers that were used for the amplification of the (RT)–PCR and the in situ hybridisation probe. Previously known USH3A human and mouse ESTs and mRNAs, amplified from several different tissues are listed in Table 3.
The USH3A gene shows partial exon overlap (but no protein overlap) with the 5′ end of the UCRP pseudogene26 in an opposite polarity, and in the same polarity with the 5′ end of a newly characterised gene, USHARF (USH3A Alternative Reading Frame, ms in preparation) (Figure 1C). By EST assembly and PCR amplification from mouse inner ear cDNA library we also defined three alternative transcripts of the Ush3A orthologue (Figure 1D). One of these transcripts (AF495719) has the same exon-intron structure as the longest human mRNA, and its protein product shares 88% identity with the product of the corresponding human transcript (AF495717). A longer mouse transcript (AF495718) has an extra exon coding for additional 18 amino acids (Figure 3C).
(A) Unrooted tree reflecting sequence relations between protein products of 12 of the 14 identified clarins. Sequence similarity searches revealed 14 members of the 4TM USH3A gene family, 12 of which, with the most completed sequences, are presented in this tree. These include HsClarin-1 (USH3A) and its two predicted paralogues HsClarin-2 (LOC166908 product of XM_068256) and HsClarin-3 (LOC119467 product of XM_058398), that map to 4p15.33 and 10q26.2 respectively; MmClarin-1 (Ush3a) and its two predicted paralogues MmClarin-2, deduced from TblastN results against Celera's mouse genomic assembly, and MmClarin-3 (product of Celera's annotated transcript mCP9472) that map to mouse chromosomes 5 and 7 respectively; three predicted Fugu gene products, TrClarin-1 (JGI_31106), TrClarin-2 (JGI_10159) and TrClarin-3 (JGI_29906) that was corrected according to TblastN and Block analysis results; Three predicted Tetraodon genes that were also reconstructed from TblastN results against Genoscope Tetraodon genomic traces and named TnClarin1,TnClarin-2 and TnClarin-3. Prediction of a partial mRNA of a bovine gene (BF044503) were also included in the analyses of the USH3 gene family but are not shown in the figure. Block multiple alignment analysis34 of the clarins identified several conserved sequence motifs covering about 95% of the sequence of clarin-1 and 80% or more of the sequences of other family members. Both PHYLIP and CLUSTAL W, used to compute trees and their significance (bootstrap) values, programs gave identical topologies. Included branch points have 70% and higher bootstrap values. (B) Sequence logo of the two regions (indicated in C by green arrows) that are similar between the clarins and members of the IPR000729 protein family. In the logo, the height of each amino acid is scaled in bits of information and is proportional to its degree of conservation.34 Black lines under the logos indicate TM prediction. (C) Alignment of clarin-1 and its three orthologues including MmClarin1, TrClarin1 (JGI_31106) and TnClarin1, which are boxed in red on the phylogenetic tree (A). Newly identified and previously described26 USH3A mutations are indicated, respectively, by red and purple arrows. The starting position of each exon is indicated above the sequence. Black and grey boxes indicate TM positions according to, respectively, PHDhtm analysis of the four aligned proteins and TMHMM analysis of clarin-1. Boxed in blue is the putative N- glycosylation site. Green arrows indicate the positions of regions, where clarins share similarity with members of the IPR000729 protein family. Human and mouse clarin-1 harbour in their C-terminal region a TNV signature that might serve as a PDZ-binding motif. However this signature is not conserved among other members of the clarins. Gaps in the aligned sequence originate from the full alignment of all 12 clarins presented in the tree (A).
Expression patterns
By PCR amplification from cDNA or cDNA libraries we confirmed the previously reported expression of USH3A in retina and skeletal muscle. In addition, we have amplified USH3A transcripts from human testis and olfactory epithelium cDNA. The later observation might be related to previously reported expression of two other Usher genes, myosin VIIA and harmonin, in the olfactory tissue.40,41,42,43 Yet, despite the observation of anomalies in olfactory cilia, no chemosensory deficits have been confirmed in Usher patients.44
By PCR amplification of mouse inner ear cDNA and by whole mount in situ hybridisation on mouse cochleae we also demonstrate the expression of Ush3a transcripts in the auditory sensory organ. A developmental expression profile of Ush3a was performed both by PCR amplification of mouse inner ear cDNA and whole mount in situ hybridisation on mouse cochleae. Ush3a transcripts were detected as early as E16, the earliest age tested, and at all postnatal stages examined (P0, P5, and P10) (Figure 2A,B). Hybridisation with an Ush3a RNA probe on whole cochleae detected mRNA expression in the sensory epithelium at E16 (Figure 2C,D), P0, P5, P10, P15, P20 (n=4 for each age) (data not shown). Sectioning through these cochleae revealed specific hybridisation in the inner and outer hair cells of the organ of Corti (Figure 2E,G). No hybridisation was detected in the supporting cells of the organ of Corti, including the Deiters' cells, the pillar cells, and the Hensen cells. The only other hybridisation found in the cochlea was in the spiral ganglion cells (Figure 2F) containing the primary neurons that innervate the cochlear sensory epithelia.
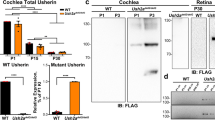
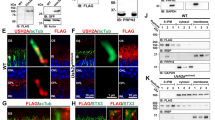
Expression of Ush3a in mouse cochlea by reverse transcriptase (RT)–PCR and in situ hybridisation. (A) Expression of Ush3a in mouse cochlea by RT–PCR at E16 and P0, P5, and P10. RNA derived from cochlear ducts was used to amplify a 605 bp fragment spanning exons 2–4 of the Ush3a gene by RT–PCR. Amplifications were carried out with (+) and without (−) RT on cochlear RNA, on mouse genomic DNA (G), and a water control (W). (B) A known cochlear-expressing gene, myosin VI (Myo6),64 amplified with primers spanning an intron from the same RNA preparation. (C–G) In situ hybridisation on mouse cochlea was done using digoxigenin-labelled sense and antisense riboprobes for Ush3a, followed by cryosectioning to confirm specific cell patterns. Expression was shown in the hair cells of the cochlea and in spiral ganglion cells at all ages tested. (C) Whole mount of mouse cochlea at E16 reveals specific labelling of sensory epithelium by the Ush3a antisense probe (arrow). Bar indicates area of cross-section for (E). (D) Absence of labelling with the Ush3a sense probe in a whole mount cochlea. (E) Cross section of cochlea in C showing a portion of the organ of Corti and specific labelling of the Ush3a probe only in the inner (IHC) and outer hair cells (OHC). Hybridisation is absent in the supporting cells, including the pillar cells (PC), Deiters' cells (DC), and Hensen's cells (HC). Scale bar, 10 μm. Left corresponds to the peripheral end of the section bar in C. (F) The region containing the spiral ganglion (SG) cells is shown. No staining was seen in any other regions of the cochlea upon sectioning. Scale bar, 50 μm. (G) Cross section of cochlea showing hair cells, supporting cells, and spiral ganglion in D demonstrating lack of hybridisation with sense probe. Scale bar, 10 μm. All experiments with mice were carried out with the approval of the Tel Aviv University Animal Care and Use Committee (11-00-65).
Gene product and protein structure
The open reading frame of the newly defined USH3A transcript is expected to encode a 232 amino acid protein with four transmembrane domains, which we named clarin-1, after the clarity of sensory perception allowed by the intact protein (Figure 3C). Sequence database searches by BLAST33 identified two additional clarin-1 human paralogues, clarin-2 and -3 (Figure 3A). The clarins were found to have respective orthologues in mouse as well as in fish (Takifugu rubripes [Fugu] and Tetraodon nigroviridis), all encoding small 224–284 amino acid proteins (Figure 3A). A partial clarin-1 mRNA was also identified in chicken. No orthologues were identified in the genomes of prokaryotes, yeast, plants, nematodes and insects. Four transmembrane-domains (TMs), conserved sequence motifs and a single glycosylation consensus site between TM1 and TM2 characterise the clarin family members (Figure 3C).
In a wider context the clarins appear to belong to a large hyperfamily of small integral membrane glycoproteins with four transmembrane domains. These include the tetraspannins45 (InterPro families PR00218 and PS00930) as well as the PMP22/EMP/MP20 and claudin family (InterPro family IPR000729) that perform diverse membrane transport, transduction and cell–cell interaction and scaffolding functions. Yet, BLAST searches did not provide a sequence similarity basis for a relationship between clarin-1 and any members of this hyperfamily. To examine this question further, it was necessary to employ more sensitive search routines, capable of detecting subtle sequence similarities. First, we asked whether any of the clarins contained protein motifs already included in the BLOCKS database, but the results were negative. Therefore, using the newly discovered group of 12 clarin sequences, we defined novel clarin-specific Blocks motifs. These were subsequently analysed by the Local Alignment of Multiple Alignments (LAMA) method,46 looking for remote but statistically significant similarities between the clarin Blocks and all others. Two of the clarin Blocks, in and around the first and fourth putative TM domains generated a hit with two of the 4TM proteins Blocks (Figure 3B,C). Pairwise Smith-Waterman alignments47 then allowed us to delineate the broad similarity relations between the clarins and representative members of several human 4TM families (Figure 5).
Schematic relations of clarins to other four-transmembrane-domain proteins. Each sequence family is colour coded as follows: clarins: red, tetraspanins: purple, connexins: blue, claudins: orange and brown, EMPs and PMP22: turquoise, MP20 and calcium channel γ subunit-like proteins (CACNGs): green, CACNG1 and CACNG6: dark green. Claudins, EMPs, PMP22, MP20 and CACNGs all belong to the IPR000729 protein family. Marked with a black star are genes that were previously shown to be involved in forms of deafness or retinal diseases. Distances between all sequences were calculated by aligning each sequence pair. The presented unrooted tree was calculated from the resulting distance matrix using the Neighbor-Joining method.34 Sequence SwissProt/TREMBL accessions are clarin-1 (USH3A): this work, clarin-2: XP_068256.1, clarin-3: XP_058398.1, CACNG1: CCG1_HUMAN, CACNG2 (stargazin): CCG2_HUMAN, CACNG3: CCG3_HUMAN, CACNG4: CCG4_HUMAN, CACNG5: CCG5_HUMAN, CACNG6: CCG6_HUMAN, CACNG7: CCG7_HUMAN, MP20: LMIP_HUMAN, PM22 : PM22_HUMAN, EMP1: EMP1_HUMAN, EMP2: EMP2_HUMAN, claudin 1: CLD1_HUMAN, claudin 2: CLD2_HUMAN, claudin 3: CLD3_HUMAN, claudin 5: CLD5_HUMAN, claudin 8: CLD8_HUMAN, claudin 10: CLDA_HUMAN, claudin 11: CLDB_HUMAN, claudin 12: CLDC_HUMAN, claudin 14: CLDE_HUMAN, claudin 15: CLDF_HUMAN, connexin 43: CXA1_HUMAN, connexin 50: CXA8_HUMAN, connexin 32: CXB1_HUMAN, connexin 26: CXB2_HUMAN, connexin 31: CXB3_HUMAN, connexin 30: CXB6_HUMAN, CD82: CD82_HUMAN, CD53: CD53_HUMAN, CD9: CD9_HUMAN, CD63: CD63_HUMAN, TM4SF6: T4S6_HUMAN, TM4SF9: T4S9_HUMAN, peripherin (RDS) : RDS_HUMAN, ROM1: ROM1_HUMAN, TNE6: TNE6_HUMAN, uroplakinA: UPKA_HUMAN.
In order to find out which of the 4TM proteins is most highly related to the clarins, we performed further pairwise alignments with accurate statistical testing (Figure 6). This highlighted the calcium channel gamma subunit proteins (CACNG) as best clarin sequence matches. Of these CACNG2 (stargazin)48,49 was the only protein showing significant matches (P<0.001) to all three clarins. It is noteworthy that like in some USH3 cases,2 stargazin mutation appears to affect also the inner ear vestibular function.48,49
(A) Significance values of sequence similarity scores (P values) between the three human clarin paralogues and some other four transmembrane-domain proteins. P values were calculated using the PRSS program47 with 1000 shuffled sequences and the program's default parameters. Changing the shuffling method from uniform to window did not change the results significantly (not shown). Pink squares, blue triangles and yellow circles in this figure represent P values of comparisons with clarin-1, clarin-2 and clarin-3 respectively. Bars indicate the geometric means of the three. (B) Alignment of the first 58 amino acids of clarins-1 clarin-2 and stargazin. Boxed in red is the putative N-glycosylation site. Average positions of the inferred first extracellular loop and N-glycosylation site were found to be more similar between the clarins and the γ subunits calcium channel CACNG1-5, which include stargazin, than between the clarins and the claudins and other members of the IPR000729 protein family (data not shown).
USH3A mutations
Three novel USH3 mutations were identified among 10 affected individuals from six USH3A families (Table 1 and Figure 4). A 143T>G substitution expected to cause a N48K missense mutation was found in six affected individuals from four unrelated families of Eastern European Jewish origin (Figure 4A). Shared microsatellite and SNP haplotypes on carrier chromosomes (data not shown) suggest the existence of a founder effect for this mutation. A 189C>A substitution, expected to cause a Y63X nonsense mutation, was found in a homozygous state in three affected individuals from a non-consanguineous Spanish family (Figure 4B; Table 1). A 23 bp deletion, spanning nucleotides 187–209 downstream from the first methionine codon, was found in a homozygous state in two affected individuals from a non-consanguineous family of Yemenite Jewish origin (Figure 4C).
USH3 families and mutations. (A) Four Eastern-European Jewish USH3 families. Below the pedigrees are parts of exon 0 chromatograms of DNA from non-carrier (top) and affected (bottom) individuals. Marked with an arrow is the 143T>G substitution, which is expected to cause an N48K missense mutation. Five patients from families 2–4 are homozygous for the 143T>G substitution while the patient from family 1 carries this substitution only on one allele. The second USH3A mutation of this individual was not detected. This result might be due to mutation in unexplored control regions or to a gross deletion, which in hemizygous state is hard to detect by the standard methods used in this work. (B) An USH3 family of Spanish origin. Below are parts of exon 0 chromatograms of non-carrier and affected individuals. Marked with an arrow is the A 189C>A substitution that is expected to cause a Y63X nonsense mutation. (C) An USH3 family of Yemenite Jewish origin. Below is FMC's MetaPhor agarose-gel electrophoresis of USH3A exon 0 fragments containing the 187–209 23 bp, which were amplified from genomic DNA samples of all family members.
Discussion
Affiliation of clarin-1 to the newly defined clarin family and the definition of several motifs characterizing all members of this family indicate that the full-length coding sequence of the clarin-1 has now been determined. The additional 18aa encoded by the longest mouse transcript as well as additional protein segments encoded by the long transcripts of Trclarin-1 and Hsclarin-2 elongate the putative first extracellular loop (Figure 3C). On the other hand, Trclarin-3 and Tnclarin-3 seem to be missing the C-terminal TM domain, indicated by a dashed blue line in Figure 3C. The fact that no orthologues were identified in the genomes of prokaryotes, yeast, plants, nematodes and insects, suggests that this gene family is limited to vertebrates.
The single missense USH3A mutation newly identified in this study, N48K, disrupts the molecule's only N-glycosylation consensus site (Figure 3C), and may render clarin-1 incapable of proper intracellular trafficking and plasma membrane insertion.50 Both other newly identified mutations, the Y63X and the 187–209del23 bp mutations, are expected to result in truncated proteins. Thus, these three mutations likely render clarin-1 functionally inactive, and account for the disease in most studied USH3A patients (Figure 4). Of the previously identified missense mutations,26 one (M120K under the new enumeration) leads to the replacement of a hydrophobic methionine residue by a charged lysine inside or very near to the border of TM2, and another (IL153,154M) results in the shortening of TM3 by one residue. Because the four TMs are rather well conserved, such mutations may also be functionally deleterious.
USH3 is progressive, i.e. at birth both vision and audition are much less severely impaired than later in life. This implies that the role played by clarin-1 in the hair cell and retina may display at least a measure of functional redundancy. The best candidates for serving as substitute for clarin-1 may be its two orthologues, clarin-2 and clarin-3. However, it is also possible that other 4TM proteins might subserve this function. Future studies will be needed to clarify the mechanism of time-dependent loss of such potential redundancy.
Other small 4TM proteins have previously been shown to underlie deafness and retinal diseases. Mutations in four different connexins (26, 30, 31, and 43) and in claudin-14 underlie several forms of deafness (Hereditary Hearing Loss Homepage: http://www.uia.ac.be/dnalab/hhh/). Mutations in connexin 50 cause congenital cataracts51 and mutations in human peripherin/RDS and mouse ROM were shown to disrupt photoreceptor morphogenesis, leading to retinitis pigmentosa.45 Also, mouse mutations in Cacng2 (stargazin) cause defects not only in cerebellum but also in inner ear.52 Yet, such information cannot help define more accurately the underlying mechanism for USH3, because the above listed genes participate in diverse cellular pathways, including the assembly and maintenance of gap junctions, tight junctions and synaptic junctions.53,54,55
Because stargazin has been shown to play a key role in the shaping and maintenance of cerebellar synapses,55 our protein sequence analysis is suggestive of a synaptic role for clarin-1 as well. The case is strengthened by corroborative evidence: (a) Stargazin has been shown to interact with PDZ domain-containing proteins of the Membrane Associated Guanylate Kinase type (MAGUK), serving as intracellular anchors essential for the integrity of synaptic densities.55 Intriguingly, one form of the Usher syndromes (USH1C) is caused by mutations in another PDZ protein harmonin. This may indicate that clarin-1 and harmonin are part of the same synapse-formation pathway, despite the absence of a clear PDZ-binding consensus56 shared by all the clarins (Figure 3C). (b) Stargazin was shown to be involved in controlling the expression and mobilisation of amino-hydroxyl-methyl-isoxazole propionate (AMPA) glutamate receptors to the post-synaptic cleft in cerebellar granule cells. That cochlear hair cells also utilize glutamate (or a highly similar compound) as a rapid excitatory neurotransmitter57 seems relevant. (c) Stargazin and other members of the CACNG family have been suggested to also play a role of forming protein–protein contacts across the synaptic cleft.58 In bridging two cellular membranes they resemble other 4TM proteins such as the gap junction-forming connexins and the tight junction-forming claudins. That clarin-1 is expressed in inner ear, both pre- and post-synaptically, is consistent with possible clarin–clarin homophylic interactions, which might work in the hair cell synapse. Thus, we would like to propose that clarin-1 has a role in the excitatory ribbon synapse junctions between hair cells and cochlear ganglion cells,59 and presumably also in analogous synapses within the retina.
We have previously reported a possible epistatic interaction between the USH3A locus and the MYO7A gene,27 whereby two USH3A haploidentical patients of a Yemenite family showed different USH phenotypes, the patient with the more severe, USH1-like phenotype was also found to be a carrier for a complex rearrangement (3260T>C and 3266delG) in the tail of myosin VIIA.27 Both patients are shown here to be homozygous for the 23 bp deletion in USH3A exon 0 (Figure 4C). While the double heterozygous mother and siblings are healthy, in the context of a homozygous USH3A null mutation the presence of one null mutation in the MYO7A tail (the out of phase one base deletion in the complex rearrangement) mimics haploinsufficiency, and illustrates a departure from the monogenic model.
Since MYO7A expression in the inner ear is restricted to sensory hair cells,60,61,62 including the sensory synaptic region,62 the present observation that Ush3a is expressed in mouse sensory hair cells, and the implication of a synaptic role, are compatible with a clarin-1/MYO7A interaction. Furthermore, while myosin VIIA was localised only in pre-synaptic cells, MyRIP, a most recently defined MYO7A-interacting protein,63 is present in both pre- and post-synaptic cells, similar to clarin-1. The proposed clarin-1/MYO7A interaction is also consistent with a pathophysiological continuum between USH1 and USH3A, and is relevant to their treatment. Access to Ush3 mutant models should allow testing of these hypotheses and contributing to a better understanding of the role of clarin-1 in the retina and inner ear.
References
Petit C . Usher syndrome: from genetics to pathogenesis Annu Rev Genomics Hum Genet 2001 2: 271–297
Pakarinen L, Karjalainen S, Simola KO, Laippala P, Kaitalo H . Usher's syndrome type 3 in Finland Laryngoscope 1995 105: 613–617
Kaplan J, Gerber S, Bonneau D et al. A gene for Usher syndrome type I (USH1A) maps to chromosome 14q Genomics 1992 14: 979–987
Kimberling W, Smith RJ . Gene mapping of the Usher syndromes Otolaryngol Clin North Am 1992 25: 923–934
Smith RJ, Lee EC, Kimberling WJ et al. Localization of two genes for Usher syndrome type I to chromosome 11 Genomics 1992 14: 995–1002
Wayne S, Der Kaloustian VM, Schloss M et al. Localization of the Usher syndrome type ID gene (Ush1D) to chromosome 10 Hum Mol Genet 1996 5: 1689–1692
Chaib H, Kaplan J, Gerber S et al. A newly identified locus for Usher syndrome type I, USH1E, maps to chromosome 21q21 Hum Mol Genet 1997 6: 27–31
Kimberling WJ, Weston MD, Moller C et al. Localization of Usher syndrome type II to chromosome 1q Genomics 1990 7: 245–249
Hmani M, Ghorbel A, Boulila-Elgaied A et al. A novel locus for Usher syndrome type II, USH2B, maps to chromosome 3 at p23-24.2 Eur J Hum Genet 1999 7: 363–367
Pieke-Dahl S, Moller CG, Kelley PM et al. Genetic heterogeneity of Usher syndrome type II: localisation to chromosome 5q J Med Genet 2000 37: 256–262
Mustapha M, Chouery E, Torchard-Pagnez D et al. A novel locus for Usher syndrome type I, USH1G, maps to chromosome 17q24-25 Hum Genet 2002 110: 348–350
Sankila EM, Pakarinen L, Kaariainen H et al. Assignment of an Usher syndrome type III (USH3) gene to chromosome 3q Hum Mol Genet 1995 4: 93–98
Joensuu T, Hamalainen R, Lehesjoki AE, de la Chapelle A, Sankila EM . A sequence-ready map of the Usher syndrome type III critical region on chromosome 3q Genomics 2000 63: 409–416
Weil D, Blanchard S, Kaplan J et al. Defective myosin VIIA gene responsible for Usher syndrome type 1B Nature 1995 374: 60–61
Liu XZ, Hope C, Walsh J et al. Mutations in the myosin VIIA gene cause a wide phenotypic spectrum, including atypical Usher syndrome Am J Hum Genet 1998 63: 909–912
Bitner-Glindzicz M, Lindley KJ, Rutland P et al. A recessive contiguous gene deletion causing infantile hyperinsulinism, enteropathy and deafness identifies the Usher type 1C gene Nat Genet 2000 26: 56–60
Verpy E, Leibovici M, Zwaenepoel I et al. A defect in harmonin, a PDZ domain-containing protein expressed in the inner ear sensory hair cells, underlies Usher syndrome type 1C Nat Genet 2000 26: 51–55
Kornau HC, Seeburg PH, Kennedy MB . Interaction of ion channels and receptors with PDZ domain proteins Curr Opin Neurobiol 1997 7: 368–373
Di Palma F, Holme RH, Bryda EC et al. Mutations in Cdh23, encoding a new type of cadherin, cause stereocilia disorganization in waltzer, the mouse model for Usher syndrome type 1D Nat Genet 2001 27: 103–107
Bork JM, Peters LM, Riazuddin S et al. Usher syndrome 1D and nonsyndromic autosomal recessive deafness DFNB12 are caused by allelic mutations of the novel cadherin-like gene CDH23 Am J Hum Genet 2001 68: 26–37
Alagramam KN, Yuan H, Kuehn MH et al. Mutations in the novel protocadherin PCDH15 cause Usher syndrome type 1F Hum Mol Genet 2001 10: 1709–1718
Ahmed ZM, Riazuddin S, Bernstein SL et al. Mutations of the protocadherin gene PCDH15 cause Usher syndrome type 1F Am J Hum Genet 2001 69: 25–34
Nollet F, Kools P, van Roy F . Phylogenetic analysis of the cadherin superfamily allows identification of six major subfamilies besides several solitary members J Mol Biol 2000 299: 551–572
Suzuki ST . Recent progress in protocadherin research Exp Cell Res 2000 261: 13–18
Eudy JD, Weston MD, Yao S et al. Mutation of a gene encoding a protein with extracellular matrix motifs in Usher syndrome type IIa Science 1998 280: 1753–1757
Joensuu T, Hamalainen R, Yuan B et al. Mutations in a novel gene with transmembrane domains underlie Usher syndrome type 3 Am J Hum Genet 2001 69: 673–684
Adato A, Kalinski H, Weil D et al. Possible interaction between USH1B and USH3 gene products as implied by apparent digenic deafness inheritance Am J Hum Genet 1999 65: 261–265
Espinos C, Najera C, Millan JM et al. Linkage analysis in Usher syndrome type I (USH1) families from Spain J Med Genet 1998 35: 391–398
Ahituv N, Sobe T, Robertson NG et al. Genomic structure of the human unconventional myosin VI gene Gene 2000 261: 269–275
Wilkinson DG, Nieto MA . Detection of messenger RNA by in situ hybridization to tissue sections and whole mounts Methods Enzymol 1993 225: 361–373
Bassam BJ, Caetano-Anolles G, Gresshoff PM . Fast and sensitive silver staining of DNA in polyacrylamide gels Anal Biochem 1991 196: 80–83
Glusman G, Lancet D . GESTALT: a workbench for automatic integration and visualization of large-scale genomic sequence analyses Bioinformatics 2000 16: 482–483
Altschul SF, Madden TL, Schaffer AA et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs Nucleic Acids Res 1997 25: 3389–3402
Henikoff S, Henikoff JG, Alford WJ, Pietrokovski S . Automated construction and graphical presentation of protein blocks from unaligned sequences Gene 1995 163: GC17–GC26
Schuler GD, Altschul SF, Lipman DJ . A workbench for multiple alignment construction and analysis Proteins 1991 9: 180–190
Felsenstein J . PHYLIP–Phylogeny Inference Package(Version 3.2) Cladistics 1989 164–166
Thompson JD, Higgins DG, Gibson TJ, Clustal W . improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice Nucleic Acids Res 1994 22: 4673–4680
Rost B, Fariselli P, Casadio R . Topology prediction for helical transmembrane proteins at 86% accuracy Protein Sci 1996 5: 1704–1718
Krogh A, Larsson B, von Heijne G, Sonnhammer EL . Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes J Mol Biol 2001 305: 567–580
Arden GB, Fox B . Increased incidence of abnormal nasal cilia in patients with retinitis pigmentosa Nature 1979 279: 534–536
Sahly I, El-Amraoui A, Abitbol M, Petit C, Dufier JL . Expression of myosin VIIA during mouse embryogenesis Anat Embryol (Berl) 1997 196: 159–170
Wolfrum U, Liu X, Schmitt A, Udovichenko IP, Williams DS . Myosin VIIa as a common component of cilia and microvilli Cell Motil Cytoskeleton 1998 40: 261–271
Marietta J, Walters KS, Burgess R et al. Usher's syndrome type IC: clinical studies and fine-mapping the disease locus Ann Otol Rhinol Laryngol 1997 106: 123–128
Seeliger M, Pfister M, Gendo K et al. Comparative study of visual, auditory, and olfactory function in Usher syndrome Graefes Arch Clin Exp Ophthalmol 1999 237: 301–307
Hemler ME . Specific tetraspanin functions J Cell Biol 2001 155: 1103–1107
Pietrokovski S . Searching databases of conserved sequence regions by aligning protein multiple-alignments Nucleic Acids Res 1996 24: 3836–3845
Pearson WR, Lipman DJ . Improved tools for biological sequence comparison Proc Natl Acad Sci U S A 1988 85: 2444–2448
Letts VA, Valenzuela A, Kirley JP et al. Genetic and physical maps of the stargazer locus on mouse chromosome 15 Genomics 1997 43: 62–68
Noebels JL, Qiao X, Bronson RT, Spencer C, Davisson MT . Stargazer: a new neurological mutant on chromosome 15 in the mouse with prolonged cortical seizures Epilepsy Res 1990 7: 129–135
van Geest M, Lolkema JS . Membrane topology and insertion of membrane proteins: search for topogenic signals Microbiol Mol Biol Rev 2000 64: 13–33
White TW, Goodenough DA, Paul DL . Targeted ablation of connexin50 in mice results in microphthalmia and zonular pulverulent cataracts J Cell Biol 1998 143: 815–825
Letts VA, Felix R, Biddlecome GH et al. The mouse stargazer gene encodes a neuronal Ca2+-channel gamma subunit Nat Genet 1998 19: 340–347
Shibata Y, Kumai M, Nishii K, Nakamura K . Diversity and molecular anatomy of gap junctions Med Electron Microsc 2001 34: 153–159
Heiskala M, Peterson PA, Yang Y . The roles of claudin superfamily proteins in paracellular transport Traffic 2001 2: 93–98
Chen L, Chetkovich DM, Petralia RS et al. Stargazing regulates synaptic targeting of AMPA receptors by two distinct mechanisms Nature 2000 408: 936–943
Hillier BJ, Christopherson KS, Prehoda KE, Bredt DS, Lim WA . Unexpected modes of PDZ domain scaffolding revealed by structure of nNOS-syntrophin complex Science 1999 284: 812–815
Ottersen OP, Takumi Y, Matsubara A et al. Molecular organization of a type of peripheral glutamate synapse: the afferent synapses of hair cells in the inner ear Prog Neurobiol 1998 54: 127–148
Tomita S, Nicoll RA, Bredt DS . PDZ protein interactions regulating glutamate receptor function and plasticity J Cell Biol 2001 153: F19–F24
Wagner HJ . Presynaptic bodies (“ribbons”): from ultrastructural observations to molecular perspectives Cell Tissue Res 1997 287: 435–446
Hasson T, Heintzelman MB, Santos-Sacchi J, Corey DP, Mooseker MS . Expression in cochlea and retina of myosin VIIa, the gene product defective in Usher syndrome type 1B Proc Natl Acad Sci U S A 1995 92: 9815–9819
Hasson T, Gillespie PG, Garcia JA et al. Unconventional myosins in inner-ear sensory epithelia J Cell Biol 1997 137: 1287–1307
El-Amraoui A, Sahly I, Picaud S et al. Human Usher 1B/mouse shaker-1: the retinal phenotype discrepancy explained by the presence/absence of myosin VIIA in the photoreceptor cells Hum Mol Genet 1996 5: 1171–1178
El-Amraoui A, Schonn JS, Kussel-Andermann P et al. MyRIP, a novel Rab effector, enables myosin VIIa recruitment to retinal melanosomes EMBO Rep 2002 1818
Avraham KB, Hasson T, Sobe T et al. Characterization of unconventional MYO6, the human homologue of the gene responsible for deafness in Snell's waltzer mice Hum Mol Genet 1997 6: 1225–1231
Acknowledgements
We are grateful to all patients and their family members who participated in this study. We would also like to thank Ronna Hertzano for the preparation of the mouse inner ear cDNA. This work was funded by an Infrastructure grant of the Israeli Ministry of Science Culture and Sports, the Crown Human Genome Center at The Weizmann Institute of Science, the Alfried Krupp Foundation and by the Finnish Eye and Tissue Bank Foundation, the Finnish Eye Foundation, the Maud Kuistila Memorial Foundation, the Oskar Oflund Foundation, Finnish State grant TYH9235, the European Commission (QLG2-CT-1999-00988) (KB Araham) and by the Foundation Fighting Blindness. JS Beckman holds the, Hermann Mayer professorial chair and D Lancet holds the Ralf and Lois Silver professorial chair.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Adato, A., Vreugde, S., Joensuu, T. et al. USH3A transcripts encode clarin-1, a four-transmembrane-domain protein with a possible role in sensory synapses. Eur J Hum Genet 10, 339–350 (2002). https://doi.org/10.1038/sj.ejhg.5200831
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200831
Keywords
This article is cited by
-
Usher syndrome IIIA: a review of the disorder and preclinical research advances in therapeutic approaches
Human Genetics (2022)
-
The genetic and phenotypic landscapes of Usher syndrome: from disease mechanisms to a new classification
Human Genetics (2022)
-
Modeling and Preventing Progressive Hearing Loss in Usher Syndrome III
Scientific Reports (2017)
-
Genetic characterization and disease mechanism of retinitis pigmentosa; current scenario
3 Biotech (2017)
-
Emerging Gene Therapies for Genetic Hearing Loss
Journal of the Association for Research in Otolaryngology (2017)