Abstract
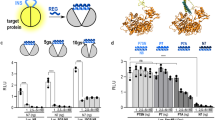
Regulation of protein function is vital for the control of cellular processes. Proteins are often regulated by allosteric mechanisms, in which effectors bind to regulatory sites distinct from the active sites and alter protein function. Intrasteric regulation, directed at the active site and thus the counterpart of allosteric control, is now emerging as an important regulatory mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Monod,J., Changeux,J. P. & Jacob,F. Allosteric proteins and cellular control systems. J. Mol. Biol. 6, 306–329 (1963).
Kemp,B. E. & Pearson,R. B. Intrasteric regulation of protein kinases and phosphatases. Biochim. Biophys. Acta 1094, 67–76 (1991).
Kemp,B. E. et al. in Protein Kinases (ed. Woodgett, J. R.) 30–67 (IRL, Oxford, 1994).
Knighton,D. R. et al. Structure of a peptide inhibitor bound to the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 414–420 (1991).
Hu,S.-H. et al. Insights into autoregulation from the crystal structure of twitchin kinase. Nature 369, 581–584 (1994).
Mayans,O. et al. Structural basis for activation of the titin kinase domain during myofibrillogenesis. Nature 395, 863–869 (1998).
Goldberg,J., Nairn,A. C. & Kuriyan,J. Structural basis for the auto-inhibition of calcium/calmodulin-dependent protein kinase I. Cell 84, 875–887 (1996).
Kissinger,C. R. et al. Crystal structures of human calcineurin and the human FKBP12-FK506-calcineurin complex. Nature 378, 641–644 (1995).
Kobe,B. et al. Giant protein kinases: domain interactions and structural basis of autoregulation. EMBO J. 15, 6810–6821 (1996).
Heierhorst,J. et al. Ca2+/S100 regulation of giant protein kinases. Nature 380, 636–639 (1996).
Hubbard,S. R., Wei,L., Ellis,L. & Hendrickson,W. A. Crystal structure of the tyrosine kinase domain of the human insulin receptor. Nature 372, 746–754 (1994).
Hubbard,S. R. Crystal structure of the activated insulin receptor tyrosine kinase in complex with peptide substrate and ATP analog. EMBO J. 16, 5572–5581 (1997).
Zhang,F., Strand,A., Robbins,D., Cobb,M. H. & Goldsmith,E. J. Atomic structure of the MAP kinase ERK2 at 2.3 Å resolution. Nature 367, 704–711 (1994).
Canagarajah,B. J., Khokhlatchev,A., Cobb,M. H. & Goldsmith,E. J. Activation mechanism of the MAP kinase ERK2 by dual phosphorylation. Cell 90, 859–869 (1997).
Barford,D., Das,A. K. & Egglof,M. P. The structure and mechanism of protein phosphatases: insights into catalysis and regulation. Annu. Rev. Biophys. Biomol. Struct. 27, 133–164 (1998).
Hof,P., Pluskey,S., Dhe-Paganon,S., Eck,M. J. & Shoelson,S. E. Crystal structure of the tyrosine phosphatase SHP-2. Cell 92, 441–450 (1998).
Bilwes,A. M., den Hertog,J., Hunter,T. & Noel,J. P. Structural basis for inhibition of receptor protein-tyrosine phosphatase-α by dimerization. Nature 382, 555–559 (1996).
Jiang,G. et al. Dimerization inhibits the activity of receptor-like protein-tyrosine phosphatase-α. Nature 401, 606–610 (1999).
Weiss, A. & Schlessinger, J. Switching signals on or off by receptor dimerization. Cell 94, 277–280 (1998).
Khan,A. R. & James,M. N. G. Molecular mechanisms for the conversion of zymogens to activate proteolytic enzymes. Protein Sci. 7, 815–836 (1998).
Freer,S. T., Kraut,J., Robertus,J. D., Wright,H. T. & Xuong,N. H. Chymotrypsinogen: 2.5-angstrom crystal structure, comparison with alpha-chymotrypsin, and implications for zymogen activation. Biochemistry 9, 1997–2009 (1970).
Kossiakoff,A. A., Chambers,J. L., Kay,L. M. & Stroud,R. M. Structure of bovine trypsinogen at 1.9 Å resolution. Biochemistry 16, 654–664 (1977).
Fehlhammer,H., Bode,W. & Huber,R. Crystal structure of bovine trypsinogen at 1–8 Å resolution. II. Crystallographic refinement, refined crystal structure and comparison with bovine trypsin. J. Mol. Biol. 111, 415–438 (1977).
Bode,W. & Huber,R. Natural protein proteinase inhibitors and their interactions with proteinases. Eur. J. Biochem. 204, 433–452 (1992).
Kobe,B. et al. Structural basis of autoregulation of phenylalanine hydroxylase. Nature Struct. Biol. 6, 442–448 (1999).
Carr,P. D., Verger,D., Ashton,A. R. & Ollis,D. L. Chloroplast NADP-malate dehydrogenase: structural basis of light-dependent regulation of activity by thiol oxidation and reduction. Structure 7, 461–475 (1999).
Johansson,K. et al. Structural basis for light activation of a chloroplast enzyme: the structure of sorghum NADP-malate dehydrogenase in its oxidized form. Biochemistry 38, 4319–4326 (1999).
Lin,K., Hwang,P. & Fletterick,R. J. Distinct phosphorylation signals converge at the catalytic center in glycogen phosphorylase. Structure 5, 1511–1523 (1997).
Johnson,L. N., Barford,D., Owen,D. J., Noble,M. E. & Garman,E. F. From phosphorylase to phosphorylase kinase. Adv. Second Messenger Phosphoprotein Res. 31, 11–28 (1997).
Kobe,B. Autoinhibition by an internal nuclear localization signal revealed by the crystal structure of mammalian importin alpha. Nature Structur. Biol. 6, 388–397 (1999).
Cignolani,G., Petosa,C., Weis,K. & Muller,C. W. Structure of importin-beta bound to the IBB domain of importin-alpha. Nature 399, 221–229 (1999).
Moroianu,J., Blobel,G. & Radu,A. The binding site of karyopherin alpha for karyopherin beta overlaps with a nuclear localization sequence. Proc. Nat. Acad. Sci. USA 93, 6572–6576 (1996).
Conti,E., Uy,M., Leighton,L., Blobel,G. & Kuriyan,J. Crystallographic analysis of the recognition of a nuclear localization signal by the nuclear import factor karyopherin alpha. Cell 94, 193–204 (1998).
Gorlich,D., Henklein,P., Laskey,R. A. & Hartmann,E. A 41 amino acid motif in importin-alpha confers binding to importin-beta and hence transit into the nucleus. EMBO J. 15, 1810–1817 (1996).
Weis,K., Ryder,U. & Lamond,A. I. The conserved amino-terminal domain of hSRP1 alpha is essential for nuclear protein import. EMBO J. 15, 1818–1825 (1996).
Xu,W., Harrison,S. C. & Eck,M. J. Three-dimensional structure of the tyrosine kinase c-Src. Nature 385, 595–602 (1997).
Sicheri,F., Moarefi,I. & Kuriyan,J. Crystal structure of the Src family tyrosine kinase Hck. Nature 385, 602–653 (1997).
Williams,J. C. et al. The 2.35 Å crystal structure of the inactivated form of chicken Src: a dynamic molecule with multiple regulatory interactions. J. Mol. Biol. 274, 757–775 (1997).
Moarefi,I. et al. Activation of the Src-family tyrosine kinase Hck by SH3 domain displacement. Nature 385, 650–653 (1997).
Chinkers,M. & Garbers,D. L. The protein kinase domain of the ANP receptor is required for signalling. Science 245, 1392–1394 (1989).
Morgan,D. O. Principles of CDK regulation. Nature 374, 131–134 (1995).
Hurley,J. H., Dean,A. M., Sohl,J. L., Koshland,D. E. Jr & Stroud,R. M. Regulation of an enzyme by phosphorylation at the active site. Science 249, 1012–1016 (1991).
Wang,H. et al. NMR solution structure of the 21 kDa chaperone protein DnaK substrate binding domain: a preview of chaperone-protein interaction. Biochemistry 37, 7929–7940 (1998).
Morshauer,R. C. et al. High-resolution solution structure of the 18 kDa substrate-binding domain of the mammalian chaperone protein Hsc70. J. Mol. Biol. 289, 1387–1403 (1999).
Park,H.-W., Boduluri,S. R., Moomaw,J. F., Casey,P. J. & Beese,L. S. Crystal structure of protein farnesyltransferase at 2.25 Angstrom resolution. Science 275, 1800–1804 (1997).
Nicholls,A., Sharp,K. A. & Honig,B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank J. Heierhorst, I. Jennings and other members of the protein research groups at St. Vincent's Institute for discussions; T. Teh for comments on the manuscript; and P. Carr for unpublished data. The work was supported by the National Health and Medical Research Council, Wellcome Trust, Australian Research Council and National Heart Foundation; B.K. is a Wellcome Senior Research Fellow in Medical Science in Australia; B.E.K. is an NHMRC Fellow.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kobe, B., Kemp, B. Active site-directed protein regulation . Nature 402, 373–376 (1999). https://doi.org/10.1038/46478
Issue Date:
DOI: https://doi.org/10.1038/46478
This article is cited by
-
Smart polymer chemosensors: signal-amplification systems with allosterism
Polymer Journal (2021)
-
Allosteric signal-amplification sensing with polymer-based supramolecular hosts
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2019)
-
Autophosphorylation of CaMKK2 generates autonomous activity that is disrupted by a T85S mutation linked to anxiety and bipolar disorder
Scientific Reports (2015)
-
Chemical approaches to study metabolic networks
Pflügers Archiv - European Journal of Physiology (2013)
-
Plasma membrane calcium ATPase 4b inhibits nitric oxide generation through calcium-induced dynamic interaction with neuronal nitric oxide synthase
Protein & Cell (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.