Abstract
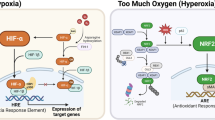
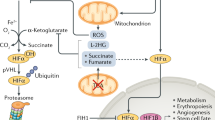
The past decade of research on hypoxic responses has provided a considerable understanding of how cells respond to hypoxic stress at the molecular level, thanks to the identification and molecular cloning of the hypoxia-inducible transcription factor, HIF-1α. Numerous target genes have since been identified to account for various aspects of the hypoxic response, including angiogenesis and glycolysis. Yet, fundamental questions remain regarding the mechanisms by which hypoxia controls cell proliferation, genetic instability, mitochondrial biogenesis, and oxidative respiration in cancer cells. Although the proto-oncoprotein c-Myc appears to be the diametrical opposite of HIF-1α in most of these processes, recent studies indicate that c-Myc is an integral part of the HIF-α–c-Myc molecular pathway in the hypoxic response. It has been shown that HIF-α engages with Myc by various mechanisms to achieve oxygen homeostasis for cell survival. This article focuses on the intricate roles of c-Myc in the hypoxic response, discusses various mechanisms controlling c-Myc activity by HIF-α for the regulation of hypoxia-responsive genes, and emphasizing the outcome of gene expression apparently dependent upon hypoxic conditions, cellular context, and gene promoter.
Similar content being viewed by others
Main
Solid tumors tend to develop hypoxia (oxygen deprivation), which arises out of the rapid cell proliferation that outstrips the supply of oxygen from the blood vessels. Consequently, tumor hypoxia activates hypoxia-inducible factor (HIF)-1α and HIF-2α,1, 2 two of the well-characterized hypoxia-inducible transcription factors critical for tumor angiogenesis, glycolysis, and metastasis.3, 4, 5 Under physiological conditions, hypoxia activates the ubiquitous HIF-1α, whereas HIF-2α expression is more tissue specific. Yet in human cancers of diverse origins, both HIF-1α and HIF-2α, referred to collectively hereinafter as HIF-α, are frequently overexpressed and activated.
Hypoxia-inducible factor-α is the regulatory subunit of the HIF heterodimeric complex paired with the aryl hydrocarbon receptor nuclear translocator (ARNT, also known as HIF-1β).6 They belong to the Per-ARNT-Sim (PAS) superfamily7 of transcription factors containing basic helix–loop–helix domains (Figure 1a). Whereas PAS domains confer target gene specificity through protein–protein interactions,8 the oxygen-dependent degradation domain, unique to HIF-α, mediates oxygen-dependent proteolysis.9 As such, despite constitutive transcription and translation of the HIF1A and EPAS1 genes (encoding HIF-1α and HIF-2α, respectively), HIF-α protein levels remain low in oxygenated conditions because of proteolysis via the ubiquitin–proteasome pathway.9, 10, 11, 12 HIF-α degradation requires the pVHL-containing E3 ubiquitin ligase,13, 14, 15 which recognizes two hydroxylated proline residues of HIF-α (P402 and P564 in HIF-1α; Figure 1a).16, 17, 18 HIF-α is hydroxylated by three prolyl hydroxylases, EglN1, EglN2, and EglN3 (better known as PHD2, PHD1, and PHD3, respectively), which sense oxygen tension and transduce oxygen signals through hydroxylation.19, 20, 21 Interestingly, HIF-1α is also subjected to asparaginyl hydroxylation at N803 (Figure 1a), which is located in the C-terminal transactivation domain that recruits the transcription co-activator p300/CBP.22 Thus, the asparaginyl hydroxylation hinders p300/CBP binding, thereby inhibiting HIF-α transactivation.23, 24, 25 By contrast, hypoxia inhibits hydroxylation by limiting oxygen availability, resulting in HIF-α stabilization, which triggers the HIF-α activation cascade involving dimerization with ARNT, recruitment of p300/CBP, and binding to the hypoxia-responsive element (HRE) in the promoter of hypoxia-responsive genes for transcriptional activation (Figure 1b).4
Schematic representation of the HIF-1 molecule and HIF-mediated transcriptional activation of hypoxia-responsive genes. (a) HIF-1 is a heterodimer composed of HIF-1α and ARNT. Both subunits contain bHLH (basic helix–loop–helix), PAS (Per-ARNT-Sim), and TA (transactivation) domains. The PAS domain comprises PAS-A and PAS-B subdomains. In addition, HIF-1α harbors an ODD (oxygen-dependent degradation) domain. Four indicated HIF-1α amino-acid residues are subject to post-translational modification. (b) In normoxia, HIF-α is unstable and therefore incapable of transactivation. Under hypoxia, however, HIF-α accumulates, leading to dimerization with ARNT, recruitment of p300/CBP, and binding to the HRE (hypoxia-responsive element) in the promoter of hypoxia-responsive genes, as indicated, for transcriptional activation
These remarkable advances have smoothed the way for the identification of an ever-expanding body of hypoxia-responsive genes that account for many aspects of tumor hypoxia, including angiogenesis, glycolysis, low pH, and cell survival. Yet, unanswered questions remain regarding how hypoxia alters cell proliferation, induces genetic instability, and reprograms cell metabolism and mitochondrial biogenesis, which are critical for tumor survival and progression. Recent studies, however, have begun to shed new light on these questions by unraveling the intricate role of the proto-oncoprotein c-Myc in relation to HIF-α in adaptation to the tumor microenvironment.26, 27, 28, 29
c-Myc plays a central role in a transcription factor network that regulates cellular growth, differentiation, apoptosis, and metabolism.30, 31 It is frequently overexpressed in human cancers because of genetic rearrangements such as gene amplifications and chromosomal translocation. Deregulated c-Myc expression has been shown to drive vasculogenesis, reduce cell adhesion, and promote metastasis.32 c-Myc can activate and repress transcription of its target genes. When in a binary complex with its partner Max, it activates transcription by binding to an E-box consensus sequence. However, Max also pairs with the c-Myc antagonist Mad1 or Max interactor 1 (Mxi1) to occupy the same E-box element for gene repression. In addition, the c-Myc–Max can be recruited by Miz1 for gene repression to core promoter sequences that lack the E-box sequence. Other transcription factors including Sp1 can also tether c-Myc to core promoter sequences.32 Although HIF-α and c-Myc share common target genes, such as those involved in glycolysis and angiogenesis, they apparently have opposing effects on cell proliferation, mitochondrial biogenesis, and DNA repair. How both HIF-1α and HIF-2α employ diverse mechanisms by either collaborating with or counteracting c-Myc to mediate adaptive responses to hypoxia, why HIF-1α differs functionally from HIF-2α in engaging with c-Myc, and how HIF-1α and HIF-2α sometimes have opposite effects on c-Myc are discussed below. These interesting yet complex findings suggest that target gene promoter, cellular context, and hypoxic environment determine the way by which HIF-α engages with c-Myc and consequently the outcome of target gene expression.
HIF-1α Regulates Cell Cycle and DNA Repair Genes by Counteracting c-Myc Activities Through c-Myc Displacement
The c-Myc ternary repressive complex containing Miz1 is known to repress the cyclin-dependent kinase inhibitor gene CDKN1A (encoding p21cip1) for cell-cycle progression.33, 34 By contrast, hypoxia upregulates CDKN1A gene in an HIF-1α-dependent way.35, 36 Furthermore, HIF-1α expression is sufficient to stimulate CDKN1A expression in normoxia, leading to cell-cycle arrest.26, 37 Of particular interest is that such gene upregulation, in contrast with the HIF-α activation cascade (Figure 1b), is independent of HIF-1α DNA-binding and transactivation domains.26 In fact, the N-terminal portion of HIF-1α that harbors PAS domains (Figure 1a) is sufficient to induce CDKN1A expression and cell-cycle arrest, suggesting alternative mechanisms for HIF-1α activation of the CDKN1A gene. Indeed, HIF-1α activates CDKN1A gene expression by displacing the inhibitory c-Myc from binding to the CDKN1A proximal promoter, resulting in gene derepression (Figure 2a). Likewise, another cyclin-dependent kinase inhibitor gene CDKN1B (encoding p27kip1), also suppressed by c-Myc, can be upregulated by hypoxia by a similar mechanism.28, 37 Thus, HIF-1α antagonizes repressive c-Myc activity for gene activation. It should be noted that unlike c-Myc displacement from the MSH2 promoter (see below), the biochemistry of c-Myc displacement by HIF-1α in the CDKN1A and CDKN1B gene promoters has not been well characterized.
HIF-1α regulates hypoxia-responsive gene expression through c-Myc displacement. (a) c-Myc acts as a repressor by binding to Miz1 at the initiator element (INR) of the CDKN1A gene in normoxia. Hypoxia-induced HIF-1α displaces c-Myc from the promoter, resulting in gene de-repression. (b) Activating c-Myc drives gene transcription (as indicated) through interaction with the transcription factor Sp1 that is bound directly to the GC-rich region of the promoter and direct binding to an E-box element in the intron. Hypoxic stabilization of HIF-1α induces selective c-Myc displacement and, in turn, gene repression. It is noteworthy that c-Myc binding to the E-box remains intact under hypoxia in the CDC25A and NBS1 genes
In the absence of Miz1, c-Myc forms an activating complex with Max for gene activation.38 Would HIF-1α counteract an activating c-Myc, thereby resulting in gene repression? Very few genes have been reported to be downregulated by hypoxia, and the underlying mechanisms hitherto have been elusive. Evidently, hypoxia-induced gene repression can hardly be explained by the direct role of HIF-1α in transcriptional activation through binding to the HRE (Figure 1b). However, several studies have shown that c-Myc displacement accounts for hypoxic repression of c-Myc–activated genes including the DNA repair genes MSH2, MSH6,27 and NBS139 and the cell-cycle gene CDC25A;40 that is, these c-Myc–activated genes are inhibited by hypoxia resulting from c-Myc displacement (Figure 2b). Although all of these genes share a common mechanism for gene repression, noticeable differences have been observed that distinguish one from another. For example, hypoxic downregulation of mismatch repair genes MSH2 and MSH6 has the distinction of p53 dependence. Consistently, c-Myc displacement in the MSH2 and MSH6 promoters requires wild-type p53. By contrast, hypoxic downregulation of NBS1 and CDC25A genes is p53 independent. Unlike the regulation of DNA repair genes, HIF-1α expression alone neither inhibits CDC25A expression nor induces c-Myc displacement, despite the requirement of HIF-1α for mediating hypoxia-induced gene repression and c-Myc displacement. Biochemical and promoter analyses are needed to identify the binding partners responsible for all these subtle distinctions.
A more intriguing question, however, is how HIF-1α displaces an activating c-Myc from the target gene promoter. As for the CDKN1A upregulation, HIF-1α uses its N-terminal portion, the PAS-B subdomain (Figure 1a) in particular, for gene repression.27, 39 HIF-1α competes with c-Myc for binding to the transcription factor Sp1 that directly interacts with the MSH2 promoter. Such competition is consistent with a tenuous interaction between HIF-1α and c-Myc in the cytosol.26 Accordingly, c-Myc interacts with Sp1 for gene activation in normoxia where HIF-1α levels are low. Under hypoxia, HIF-1α occupies Sp1 via c-Myc displacement, resulting in gene inactivation (Figure 2b).
The requirement of Sp1 for c-Myc displacement raises the question as to whether c-Myc displacement occurs when c-Myc is bound directly to an E-box element. Interestingly, although both NBS1 and CDC25A genes harbor an E-box element in their introns, no significant changes in c-Myc binding to the E-box have been detected when gene expression is suppressed.39, 40 Therefore, it appears that HIF-1α selectively targets c-Myc bound indirectly to DNA via another transcription factor for displacement (Figure 2b). Such a selective mechanism may explain the relatively weak suppression (approximately two- to threefold) of these genes by hypoxia, because of the remnant c-Myc-activating activity bound to the E-box. Furthermore, it stands to reason that HIF-1α might displace c-Myc from Sp1 more readily than from an E-box element, assuming a stronger binding affinity to the DNA sequence. However, more quantitative analyses of c-Myc binding to the E-box element in c-Myc-activated genes such as ODC, CCND2, and E2F1 (encoding ornithine decarboxylase 1, cyclin D2, and E2F transcription factor 1, respectively) have shown decreased c-Myc promoter occupancy under hypoxia,28 suggesting the occurrence of c-Myc displacement from the E-box element, albeit to a lesser extent. Furthermore, HIF-1α also induces a shift in the heterocomplex formation from activating c-Myc–Max to repressive Mad1–Max or Mxi1–Max, thereby further contributing to the inactivation of c-Myc target genes.28, 41
HIF-2α, in Contrast with HIF-1α, Promotes Cell Proliferation by Enhancing c-Myc Activities Through Stimulation of Max Binding
Although both HIF-1α and HIF-2α follow the same HIF-α activation cascade (Figure 1b) for gene activation, studies have shown that HIF-2α possesses divergent (patho)physiological functions despite striking similarities to HIF-1α in amino-acid sequence and protein structure.42, 43, 44, 45, 46, 47 At the molecular level, several HIF-2α preferentially regulated genes have been identified, including CCND1, EPO, POU5F1, TGFA, and VEGFA (encoding cyclin D1, erythropoietin, octamer-binding transcription factor 4, transforming growth factor α, vascular endothelial growth factor A, respectively).46, 48, 49 However, how HIF-2α specifically targets these genes remains unclear, although the involvement of ETS transcription factors in HIF-2α target gene selection has been suggested.50
Thus, given the high percentage of identity in amino-acid sequence particularly to the HIF-1α PAS-B domain, does HIF-2α also engage in c-Myc displacement? Interestingly, HIF-2α is not involved in the c-Myc counteraction because of its inability to compete with c-Myc for Sp1 binding.39 Accordingly, HIF-2α fails to repress any of the foregoing DNA repair genes.27, 39 What distinguishes HIF-2α from HIF-1α in c-Myc displacement is a specific phosphorylation of HIF-2α at codon T324 (equivalent to HIF-1α T322; Figure 1a) by protein kinase D1, which prevents HIF-2α from competing for Sp1 binding. Conversely, HIF-1α, which is not subjected to phosphorylation at T322, is capable of c-Myc displacement.
Interestingly, what is remarkable is not what HIF-2α cannot do, but what it can. It has been uncovered that HIF-2α's ‘hidden talent’ is to enhance c-Myc activities by stimulating the interaction with Max.28 As a result, HIF-2α promotes, rather than inhibits, cell-cycle progression in hypoxic cells by further inhibiting the expression of cyclin-dependent kinase inhibitor genes CDKN1A and CDKN1B (Figure 3a) and by augmenting the expression of cell-cycle genes CCND2 and E2F1 (Figure 3b). Moreover, the effects of HIF-2α on gene expression correlate with increased c-Myc target-gene promoter occupancy in hypoxic cells, and HIF-2α facilitates formation of c-Myc–Max complex through binding to Max. Evidently, further molecular and biochemical characterization of HIF-2α interactions with Max should provide a mechanistic understanding of the functional difference between HIF-1α and HIF-2α in cell-cycle regulation. It will also be interesting to determine whether these opposing effects on cell cycle by HIF-1α and HIF-2α occur in the same tumor cell or if one dominates the other depending on their expression levels.
HIF-2α, in contrast with HIF-1α, enhances c-Myc activity. (a and b) Hypoxia-induced HIF-2α increases c-Myc binding to its partners (Miz1, Sp1, and Max), resulting in further repression of CDKN1A and CDKN1B when c-Myc is repressive (a) and enhanced transcription of CCND2 and E2F1 when c-Myc is activating (b)
HIF-α Controls Mitochondrial Biogenesis by Inhibiting c-Myc-Mediated Transcription Through the Induction of c-Myc Degradation and Mxi1 Expression
In contrast with the opposing effects on c-Myc by HIF-1α and HIF-2α, the two HIF-α isoforms have been reported to work in the same direction to inhibit mitochondrial biogenesis.29 Peroxisome proliferators-activated receptor γ coactivator 1β is a transcription factor that is regulated by c-Myc and implicated in mitochondrial biogenesis. Study has shown that HIF-α overexpression in a pVHL-deficient renal carcinoma cell line inhibits PGC-1β expression, resulting in decreased mitochondrial DNA content, mitochondrial mass, and oxygen consumption.29 Both HIF-1α and HIF-2α are involved in the transcriptional repression of PPARGC1B (encoding PGC-1β). In contrast to the direct role of HIF-1α in c-Myc target gene repression (Figure 2b), the HIF-α isoforms inhibit PPARGC1B expression in two indirect ways (Figure 4a). On the one hand, HIF-α transcriptionally activates the MXI1 gene (encoding Mxi1 that competes for Max binding).29, 41 On the other hand, HIF-α contributes to c-Myc proteolysis under severe or prolonged hypoxia via the ubiquitin–proteasome pathway without affecting MYC mRNA levels. Such a double-punch mechanism of c-Myc inactivation has also been shown to be responsible for transcriptional repression of other c-Myc-activated genes such as CAD (encoding carbamoyl-phosphate synthetase 2/aspartate trans-carbamylase/dihydroorotase) and ODC1. Therefore, both HIF-α isoforms induce c-Myc degradation and MXI1 gene expression to inhibit c-Myc activity.
HIF-α inhibits the expression of c-Myc target genes in two independent ways, while deregulated c-Myc cooperates with HIF-1α for the activation of hypoxia-responsive genes. (a) Chronic hypoxia induces HIF-α-dependent c-Myc degradation and transcriptional upregulation of the c-Myc antagonist gene encoding Mxi1, which forms a complex with Max and recruits Sin3A for the repression of c-Myc target genes as indicated. (b) Deregulated c-Myc cooperates with HIF-1α under hypoxia to augment transcription of genes, as indicated, through binding to E-box and HRE, respectively
Although the contribution of HIF-α to c-Myc degradation is not well understood, the above studies support the notion that c-Myc can be regulated at different levels under different hypoxic conditions and perhaps in different cell types. Treatment with mild hypoxic conditions at 0.5–1% oxygen levels for less than 24 h leads to no significant change in c-Myc protein levels;26, 27, 28 however, severe hypoxic conditions such as 0.1% oxygen or prolonged treatment result in c-Myc degradation without affecting MYC mRNA levels.29, 41 Yet, transcriptional downregulation of MYC gene by hypoxia has also been suggested to be a possible mechanism controlling the expression of AFP gene (encoding α-fetoprotein),51 which is activated by c-Myc. Furthermore, given MYC gene being a transcriptional target of the β-catenin/T-cell factor-4 complex, such mechanism apparently accounts for the derepression of CDKN1A gene in hypoxic colon cancer cells, where HIF-1α disrupts the complex formation by competing for β-catenin binding, thereby reducing c-Myc levels.52 Therefore, c-Myc activity can be affected by various hypoxic conditions.
In c-Myc-Deregulated Cells, HIF-1α Cooperates with c-Myc to Enhance Common Target Gene Expression
In human cancers, c-Myc deregulation is mostly a result of genetic rearrangements that disable homeostatic mechanisms controlling c-Myc protein levels. Thus, c-Myc may no longer be subjected to hypoxic regulation as discussed above. It has been shown that in a Burkitt lymphoma cell line where c-Myc levels are controlled by tetracycline, HIF-1α cooperates with, rather than antagonizes, c-Myc activities to enhance the expression of shared target genes including HK2 (encoding hexokinase 2), PDK1 (encoding pyruvate dehydrogenase kinase isozyme 1), and VEGFA (Figure 4b).53 HK2 catalyzes ATP-dependent phosphorylation of glucose to form glucose-6-phosphate in the first step of glycolysis, whereas PDK1 inhibits mitochondrial respiration by inactivating pyruvate dehydrogenase, which converts pyruvate to acetyl-CoA. It is interesting to note that although coexpression of HIF-1α and c-Myc enhances glucose metabolism, HK2 is the only glycolytic gene to be upregulated in this system. The expression of LDHA, another glycolytic gene best known to be responsive to HIF-1α or c-Myc, is not further enhanced by the two. Consistent with previous reports that the PDK1 gene is a target of HIF-1α,54, 55 c-Myc augments PKD1 expression in the presence of HIF-1α for lactate production. In addition, HIF-1α and c-Myc also cooperate to activate the VEGFA gene. Such collaboration is associated with discrete and/or overlapping binding of HIF-1α and c-Myc to the target gene promoters. It remains unclear, however, what determines the interaction and/or cooperation between HIF-1α and c-Myc in the promoters of these common target genes. Whether such collaboration occurs for enhanced gene expression in other c-Myc deregulated tumor cells is yet to be determined. Furthermore, biochemical characterization of additional interacting proteins may help reconcile the differences between cooperative and antagonistic effects (discussed above) of HIF-1α on c-Myc.
Concluding Remarks
The revelation of the intricate roles of c-Myc in hypoxic responses has shed light on some of the fundamental aspects of tumor growth and progression. Although c-Myc is regarded as a central transcriptional hub in the control of growth and proliferation through binding to several thousand genomic loci,56 its activities are subjugated under hypoxic conditions by HIF-α to serve the interests of tumor cells for adaptation and survival. Various mechanisms are employed to control c-Myc activities in an attempt to adjust to hypoxic conditions by balancing the needs for cell proliferation and metabolism, angiogenesis, DNA repair, and mitochondrial biogenesis.57, 58, 59 Regardless of c-Myc cooperation with or counteraction by HIF-α, the identification of such HIF-α–c-Myc molecular pathway might have just opened the door to delve further into the cellular responses to the hypoxic microenvironment.
Abbreviations
- ARNT:
-
aryl hydrocarbon receptor nuclear translocator
- bHLH:
-
basic helix–loop–helix
- HIF:
-
hypoxia-inducible factor
- HRE:
-
hypoxia-responsive element
- INR:
-
initiator element
- ODD:
-
oxygen-dependent degradation
- PAS:
-
Per-ARNT-Sim
References
Wang GL, Jiang BH, Rue EA, Semenza GL . Hypoxia-inducible factor 1 is a basic-helix–loop–helix-PAS heterodimer regulated by cellular O2 tension. Proc Natl Acad Sci USA 1995; 92: 5510–5514.
Tian H, McKnight SL, Russell DW . Endothelial PAS domain protein 1 (EPAS1), a transcription factor selectively expressed in endothelial cells. Genes Dev 1997; 11: 72–82.
Harris AL . Hypoxia—a key regulatory factor in tumour growth. Nat Rev Cancer 2002; 2: 38–47.
Semenza GL . Targeting HIF-1 for cancer therapy. Nat Rev Cancer 2003; 3: 721–732.
Giaccia A, Siim BG, Johnson RS . HIF-1 as a target for drug development. Nat Rev Drug Discov 2003; 2: 803–811.
Wang GL, Semenza GL . Purification and characterization of hypoxia-inducible factor 1. J Biol Chem 1995; 270: 1230–1237.
Huang ZJ, Edery I, Rosbash M . PAS is a dimerization domain common to Drosophila period and several transcription factors. Nature 1993; 364: 259–262.
Zelzer E, Wappner P, Shilo BZ . The PAS domain confers target gene specificity of Drosophila bHLH/PAS proteins. Genes Dev 1997; 11: 2079–2089.
Huang LE, Gu J, Schau M, Bunn HF . Regulation of hypoxia-inducible factor 1alpha is mediated by an O2- dependent degradation domain via the ubiquitin–proteasome pathway. Proc Natl Acad Sci USA 1998; 95: 7987–7992.
Huang LE, Arany Z, Livingston DM, Bunn HF . Activation of hypoxia-inducible transcription factor depends primarily upon redox-sensitive stabilization of its alpha subunit. J Biol Chem 1996; 271: 32253–32259.
Pugh CW, O'Rourke JF, Nagao M, Gleadle JM, Ratcliffe PJ . Activation of hypoxia-inducible factor-1; definition of regulatory domains within the alpha subunit. J Biol Chem 1997; 272: 11205–11214.
Kallio PJ, Wilson WJ, O'Brien S, Makino Y, Poellinger L . Regulation of the hypoxia-inducible transcription factor 1alpha by the ubiquitin–proteasome pathway. J Biol Chem 1999; 274: 6519–6525.
Maxwell PH, Wiesener MS, Chang GW, Clifford SC, Vaux EC, Cockman ME et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 1999; 399: 271–275.
Ohh M, Park CW, Ivan M, Hoffman MA, Kim TY, Huang LE et al. Ubiquitination of hypoxia-inducible factor requires direct binding to the beta-domain of the von Hippel-Lindau protein. Nat Cell Biol 2000; 2: 423–427.
Tanimoto K, Makino Y, Pereira T, Poellinger L . Mechanism of regulation of the hypoxia-inducible factor-1alpha by the von Hippel-Lindau tumor suppressor protein. EMBO J 2000; 19: 4298–4309.
Ivan M, Kondo K, Yang H, Kim W, Valiando J, Ohh M et al. HIFalpha targeted for VHL-mediated destruction by proline hydroxylation: implications for O2 sensing. Science 2001; 292: 464–468.
Jaakkola P, Mole DR, Tian YM, Wilson MI, Gielbert J, Gaskell SJ et al. Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 2001; 292: 468–472.
Yu F, White SB, Zhao Q, Lee FS . HIF-1alpha binding to VHL is regulated by stimulus-sensitive proline hydroxylation. Proc Natl Acad Sci USA 2001; 98: 9630–9635.
Epstein AC, Gleadle JM, McNeill LA, Hewitson KS, O'Rourke J, Mole DR et al. C. elegans EGL-9 and mammalian homologs define a family of dioxygenases that regulate HIF by prolyl hydroxylation. Cell 2001; 107: 43–54.
Bruick RK, McKnight SL . A conserved family of prolyl-4-hydroxylases that modify HIF. Science 2001; 294: 1337–1340.
Kaelin WG . Proline hydroxylation and gene expression. Annu Rev Biochem 2005; 74: 115–128.
Arany Z, Huang LE, Eckner R, Bhattacharya S, Jiang C, Goldberg MA et al. An essential role for p300/CBP in the cellular response to hypoxia. Proc Natl Acad Sci USA 1996; 93: 12969–12973.
Lando D, Peet DJ, Whelan DA, Gorman JJ, Whitelaw ML . Asparagine hydroxylation of the HIF transactivation domain a hypoxic switch. Science 2002; 295: 858–861.
Lando D, Peet DJ, Gorman JJ, Whelan DA, Whitelaw ML, Bruick RK . FIH-1 is an asparaginyl hydroxylase enzyme that regulates the transcriptional activity of hypoxia-inducible factor. Genes Dev 2002; 16: 1466–1471.
Hewitson KS, McNeill LA, Riordan MV, Tian YM, Bullock AN, Welford RW et al. Hypoxia inducible factor (HIF) asparagine hydroxylase is identical to factor inhibiting HIF (FIH) and is related to the cupin structural family. J Biol Chem 2002; 277: 26351–26355.
Koshiji M, Kageyama Y, Pete EA, Horikawa I, Barrett JC, Huang LE . HIF-1alpha induces cell cycle arrest by functionally counteracting Myc. EMBO J 2004; 23: 1949–1956.
Koshiji M, To KK, Hammer S, Kumamoto K, Harris AL, Modrich P et al. HIF-1alpha induces genetic instability by transcriptionally downregulating MutSalpha expression. Mol Cell 2005; 17: 793–803.
Gordan JD, Bertout JA, Hu CJ, Diehl JA, Simon MC . HIF-2alpha promotes hypoxic cell proliferation by enhancing c-myc transcriptional activity. Cancer Cell 2007; 11: 335–347.
Zhang H, Gao P, Fukuda R, Kumar G, Krishnamachary B, Zeller KI et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 2007; 11: 407–420.
Dang CV . c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol Cell Biol 1999; 19: 1–11.
Grandori C, Cowley SM, James LP, Eisenman RN . The Myc/Max/Mad network and the transcriptional control of cell behavior. Annu Rev Cell Dev Biol 2000; 16: 653–699.
Adhikary S, Eilers M . Transcriptional regulation and transformation by Myc proteins. Nat Rev Mol Cell Biol 2005; 6: 635–645.
Herold S, Wanzel M, Beuger V, Frohme C, Beul D, Hillukkala T et al. Negative regulation of the mammalian UV response by Myc through association with Miz-1. Mol Cell 2002; 10: 509–521.
Seoane J, Le HV, Massague J . Myc suppression of the p21(Cip1) Cdk inhibitor influences the outcome of the p53 response to DNA damage. Nature 2002; 419: 729–734.
Carmeliet P, Dor Y, Herbert JM, Fukumura D, Brusselmans K, Dewerchin M et al. Role of HIF-1alpha in hypoxia-mediated apoptosis, cell proliferation and tumour angiogenesis. Nature 1998; 394: 485–490.
Goda N, Ryan HE, Khadivi B, McNulty W, Rickert RC, Johnson RS . Hypoxia-inducible factor 1alpha is essential for cell cycle arrest during hypoxia. Mol Cell Biol 2003; 23: 359–369.
Mack FA, Patel JH, Biju MP, Haase VH, Simon MC . Decreased growth of Vhl−/− fibrosarcomas is associated with elevated levels of cyclin kinase inhibitors p21 and p27. Mol Cell Biol 2005; 25: 4565–4578.
Adhikary S, Marinoni F, Hock A, Hulleman E, Popov N, Beier R et al. The ubiquitin ligase HectH9 regulates transcriptional activation by Myc and is essential for tumor cell proliferation. Cell 2005; 123: 409–421.
To KK, Sedelnikova OA, Samons M, Bonner WM, Huang LE . The phosphorylation status of PAS-B distinguishes HIF-1alpha from HIF-2alpha in NBS1 repression. EMBO J 2006; 25: 4784–4794.
Hammer S, To KK, Yoo YG, Koshiji M, Huang LE . Hypoxic suppression of the cell cycle gene CDC25A in tumor cells. Cell Cycle 2007; 6: 1919–1926.
Corn PG, Ricci MS, Scata KA, Arsham AM, Simon MC, Dicker DT et al. Mxi1 is induced by hypoxia in a HIF-1-dependent manner and protects cells from c-Myc-induced apoptosis. Cancer Biol Ther 2005; 4: 1285–1294.
Tian H, Hammer RE, Matsumoto AM, Russell DW, McKnight SL . The hypoxia-responsive transcription factor EPAS1 is essential for catecholamine homeostasis and protection against heart failure during embryonic development. Genes Dev 1998; 12: 3320–3324.
Compernolle V, Brusselmans K, Acker T, Hoet P, Tjwa M, Beck H et al. Loss of HIF-2alpha and inhibition of VEGF impair fetal lung maturation, whereas treatment with VEGF prevents fatal respiratory distress in premature mice. Nat Med 2002; 8: 702–710.
Kondo K, Klco J, Nakamura E, Lechpammer M, Kaelin WG . Inhibition of HIF is necessary for tumor suppression by the von Hippel-Lindau protein. Cancer Cell 2002; 1: 237–246.
Scortegagna M, Ding K, Oktay Y, Gaur A, Thurmond F, Yan LJ et al. Multiple organ pathology, metabolic abnormalities and impaired homeostasis of reactive oxygen species in Epas1−/− mice. Nat Genet 2003; 35: 331–340.
Covello KL, Kehler J, Yu H, Gordan JD, Arsham AM, Hu CJ et al. HIF-2alpha regulates Oct-4: effects of hypoxia on stem cell function, embryonic development, and tumor growth. Genes Dev 2006; 20: 557–570.
Gruber M, Hu CJ, Johnson RS, Brown EJ, Keith B, Simon MC . Acute postnatal ablation of Hif-2alpha results in anemia. Proc Natl Acad Sci USA 2007; 104: 2301–2306.
Raval RR, Lau KW, Tran MG, Sowter HM, Mandriota SJ, Li JL et al. Contrasting properties of hypoxia-inducible factor 1 (HIF-1) and HIF-2 in von Hippel-Lindau-associated renal cell carcinoma. Mol Cell Biol 2005; 25: 5675–5686.
Rankin EB, Biju MP, Liu Q, Unger TL, Rha J, Johnson RS et al. Hypoxia-inducible factor-2 (HIF-2) regulates hepatic erythropoietin in vivo. J Clin Invest 2007; 117: 1068–1077.
Aprelikova O, Wood M, Tackett S, Chandramouli GV, Barrett JC . Role of ETS transcription factors in the hypoxia-inducible factor-2 target gene selection. Cancer Res 2006; 66: 5641–5647.
Mazure NM, Chauvet C, Bois-Joyeux B, Bernard MA, Nacer-Cherif H, Danan JL . Repression of alpha-fetoprotein gene expression under hypoxic conditions in human hepatoma cells: characterization of a negative hypoxia response element that mediates opposite effects of hypoxia inducible factor-1 and c-Myc. Cancer Res 2002; 62: 1158–1165.
Kaidi A, Williams AC, Paraskeva C . Interaction between beta-catenin and HIF-1 promotes cellular adaptation to hypoxia. Nat Cell Biol 2007; 9: 210–217.
Kim JW, Gao P, Liu YC, Semenza GL, Dang CV . HIF-1 and dysregulated c-Myc cooperatively induces VEGF and metabolic switches, HK2 and PDK1. Mol Cell Biol 2007; 27: 7381–7393.
Kim JW, Tchernyshyov I, Semenza GL, Dang CV . HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab 2006; 3: 177–185.
Papandreou I, Cairns RA, Fontana L, Lim AL, Denko NC . HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metab 2006; 3: 187–197.
Zeller KI, Zhao X, Lee CW, Chiu KP, Yao F, Yustein JT et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc Natl Acad Sci USA 2006; 103: 17834–17839.
Koshiji M, Huang LE . Dynamic balancing of the dual nature of HIF-1alpha for cell survival. Cell Cycle 2004; 3: 853–854.
To KK, Koshiji M, Hammer S, Huang LE . Genetic instability: the dark side of the hypoxic response. Cell Cycle 2005; 4: 881–882.
Gordan JD, Thompson CB, Simon MC . HIF and c-Myc: sibling rivals for control of cancer cell metabolism and proliferation. Cancer Cell 2007; 12: 108–113.
Acknowledgements
This work is supported in part by an NIH Grant CA084563 from the National Cancer Institute. I thank K Kraus for excellent editorial assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by N Chandel
Rights and permissions
About this article
Cite this article
Huang, L. Carrot and stick: HIF-α engages c-Myc in hypoxic adaptation. Cell Death Differ 15, 672–677 (2008). https://doi.org/10.1038/sj.cdd.4402302
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402302
Keywords
This article is cited by
-
USP25 promotes pathological HIF-1-driven metabolic reprogramming and is a potential therapeutic target in pancreatic cancer
Nature Communications (2022)
-
Mild chronic hypoxia-induced HIF-2α interacts with c-MYC through competition with HIF-1α to induce hepatocellular carcinoma cell proliferation
Cellular Oncology (2021)
-
Emerging roles and the regulation of aerobic glycolysis in hepatocellular carcinoma
Journal of Experimental & Clinical Cancer Research (2020)
-
Hypoxia Pretreatment Promotes Chondrocyte Differentiation of Human Adipose-Derived Stem Cells via Vascular Endothelial Growth Factor
Tissue Engineering and Regenerative Medicine (2020)
-
Novel Peptide Inhibitors for Lactate Dehydrogenase A (LDHA): A Survey to Inhibit LDHA Activity via Disruption of Protein-Protein Interaction
Scientific Reports (2019)