Abstract
Investigations pursued during the last decade on neurodegenerative diseases have revealed a common mechanism underlying the development of such diseases: conformational disorder of certain proteins leads to the formation of misfolded protein oligomers, which subsequently develop into large protein aggregates. These aggregates entangle other denatured proteins and lipids to form disease-specific inclusion bodies. The failure of the ubiquitin-proteasome system to shred the protein aggregates has led investigators to focus their attention to autophagy, a bulk degradative system coupled with lysosomes, which is involved in non-selective shredding of large amounts of cytoplasmic components. Research in this field has demonstrated the accumulation of autophagic vacuoles and intracytoplasmic protein aggregates in patients with various neurodegenerative diseases. Although autophagy fails to degrade large protein aggregates once they are formed in the cytoplasm, drug-induced activation of autophagy is effective in preventing aggregate deposition, indicating that autophagy significantly contributes to the clearance of aggregate-prone proteins. The pivotal role of autophagy in the clearance of aggregate-prone proteins has been confirmed by a deductive approach using a brain-specific autophagy-ablated mouse model. In this review, we discuss the consequences of autophagy deficiency in neurons.
Similar content being viewed by others
Main
Cell proteins exist in a balance between continuous synthesis and degradation. In general, this flow of synthesis and degradation (i.e., turnover) contributes to exertion of cell-type-specific functions and maintenance of cell homeostasis. However, it is not uncommon that living cells are exposed to various environmental stresses, such as oxygen radicals and UV irradiation. Unfortunately, these stresses frequently cause various types of protein injuries that vitiate normal cellular functions or homeostasis and may eventually cause cell death. Prompt elimination of injured harmful proteins, which is particularly important in non-proliferative cells, such as neurons, is totally dependent on proper function of protein catabolic machineries, in which two major sophisticated apparatuses play principal roles. One is the proteasome, which is an elegantly organized multi-protease complex with catalytic activities inside its central proteinaceous chamber. It plays crucial roles in selective degradation of short-lived regulatory proteins as well as proteins with aberrant structures that should be eliminated from the cells.1 The other apparatus is the lysosome that contains many acidic hydrolases, which are separated from the cytosol by the limiting membrane. In this lysosomal pathway, degradation of plasma membrane proteins and extracellular proteins is mediated by endocytosis, whereas that of cytoplasmic components is achieved through distinct types of autophagic pathways; for example, macroautophagy, microautophagy, and chaperone-mediated autophagy.2, 3
Macroautophagy (hereafter referred to as autophagy), the major type of autophagy, is the bulk protein degradation pathway associated with marked membrane dynamics. In response to various stimuli, such as starvation (i.e., nutritional step-down) and humoral (trophic) factors (e.g., glucagon and cytokines), an isolation membrane appears promptly in the cytosol, where it gradually elongates to sequestrate cytoplasmic constituents. Subsequently, the edges of the membrane fuse together to form double-membrane structures termed autophagosomes. Autophagosomes rapidly fuse with lysosomes, and their contents engulfed together with the inner membrane are degraded by a variety of lysosomal digestive hydrolases (Figure 1).4 In addition to the importance of starvation-induced (i.e., adaptive) autophagy equipped as a fundamental survival strategy in all eukaryotic cells, growing lines of evidence point to the importance of basal autophagy that operates constitutively at low rate even under nutrient-rich environment and to its key role in global turnover of cellular components including organelles.
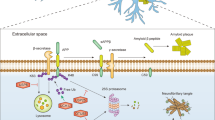
Schematic representation of the physiological functions of autophagy. Autophagy is induced in response to emergency states such as nutrient starvation or bacterial infection, which results in the degradation of cytoplasmic components for amino-acid supply (non-selective process) and the removal of bacteria that invade the cytoplasm (selective process). Autophagy also contributes to processing viral proteins such as EBNA1 and certain cytosolic proteins (e.g., tumor antigens) for presentation onto major histocompatibility complex (MHC) class II molecules. Autophagy also occurs constitutively even under a nutrient-rich state and contributes to global turnover of cellular components. It is an essential cellular process that maintains homeostasis in quiescent cells (e.g., hepatocytes and neurons)
Starvation-induced Autophagy
The most fundamental function of autophagy is cellular adaptation to nutritional stress. In yeast, autophagy is promptly induced upon nitrogen starvation.5 Transgenic mice overexpressing GFP (green fluorescent protein)-LC3 is an interesting animal model for monitoring autophagy.6 LC3 (microtubule-associated protein 1 light chain 3/MAP1LC3), originally identified as a small subunit of MAP-1A/MAP-1B, is processed by Atg4B protease to expose the carboxyl-terminal glycine whose residue serves as a donor site for conjugation of target molecules.7, 8, 9 The processed form (LC3-I) undergoes two consecutive ubiquitylation-like modification reactions catalyzed by Atg7 (E1, activating-like enzyme) and Atg3 (E2, conjugating-like enzyme), to be covalently coupled with phosphatidylethanolamine (PE).10, 11, 12 The PE-conjugated form, designated as LC3-II, is then recruited to autophagosomal membrane. Thus, LC3-II is a promising marker for autophagosomal membranes.7 Similar to endogenous LC3, GFP-LC3 responds to nutrient-starved conditions to form GFP-LC3-II, which is recruited to autophagosomes in GFP-LC3 transgenic mice.6 The autophagosomal GFP-LC3-II could be detected as dots in fluorescence microscopic analyses. Under fasting conditions, the numbers of fluorescent dots increase in the cytoplasm of the liver, heart, and skeletal muscles of GFP-LC3 transgenic mice.
Starvation-induced protein degradation has been best investigated in the liver. One unequivocal characteristic of hepatic protein degradation is its dependence on the nutrient conditions. Depending on the dietary cycle of the animal, the rate of protein degradation fluctuates between ∼1.5% (fed state) and ∼4.5% (fasted state) of total liver proteins per hour.13 Recently, we generated Atg7F/F:Mx1 mice in which autophagy could be successfully inactivated in the livers.14 Whereas the amount of total liver proteins decreased to about 66% in the control liver by 1-day fasting, fasting did not result in a significant decrease in the amount of total proteins in the autophagy-deficient liver, indicating that the decrease in total proteins upon fasting is indeed dependent on autophagy. Measurement of the activity of mitochondrial enzyme, succinate dehydrogenase (SDH), showed that fasting was also associated with a significant decrease in SDH activity in total extracts in the control livers, and such reduction was proportional with the decrease in the amount of total protein. On the other hand, fasting was not associated with any change in SDH activity in the autophagy-deficient livers. These results suggest that the mitochondria and cytoplasmic proteins are proportionally degraded upon fasting by autophagy. Thus, it is plausible that autophagosomes surround cytoplasmic components including mitochondria at random to adapt for starvation.
Yeast deficient in autophagy rapidly dies under nutrition-poor conditions,15 suggesting the important roles of autophagy in maintaining nutrient supply. Indeed, newborn mice deficient in Atg5 or Atg7, which are indispensable for autophagosome formation, show poor response to starvation with regard to production of amino acids, and die within the first day of life.14, 16 Furthermore, Lum et al.17 reported that in IL-3-dependent cells, which cannot undergo apoptosis due to knockout of both Bax and Bak, impairment of autophagy leads to rapid cell death by loss of IL-3, and such death is suppressed by addition of methylpyruvate, a TCA (tricarboxylic acid) substrate. These results suggest that one of the important roles of autophagy is the supply of amino acids under nutrient-poor environment (Figure 2, top panel).
Schematic presentation of induced autophagy and basal (constitutive) autophagy. Under nutrient-rich conditions, autophagic proteolysis proceeds in hepatocytes at a basal rate (top panel, pink zone), which is enhanced two- to threefold to an induced rate under nutrient-starvation conditions (blue zone). When animals are re-fed, the rate of autophagic proteolysis promptly returns to the basal level. In contrast to the liver, autophagy in the brain is thought to proceed at a basal rate, irrespective of nutrient conditions. However, this basal or constitutive autophagy plays a critical role in the quality control system of neurons. Rapamycin and its homologs, which upregulate autophagy to an induced level (bottom panel, broken line), are expected to prevent the accumulation of aggregate-prone proteins. It should be noted that autophagic activity declines with age, which may relate to the age-dependent onset of neurodegenerative diseases
Unique Features of Neuronal Autophagy
The brain appears to be a specially protected tissue where nutrients (e.g., amino acids, glucose, and ketone bodies) are compensated by constant supply from other organs even under starvation conditions and consequently autophagy does not operate in response to nutritional stress. Indeed, autophagosomes-related GFP-LC3 dots do not increase at all in the brain of GFP-LC3 transgenic mice, irrespective of fasting.6 A reasonable conclusion drawn from these observations is that autophagy in the brain proceeds at a basal rate but is not enhanced under fasting conditions (Figure 2, bottom panel). Another peculiar feature of autophagy in neurons is the free localization of autophagosomes in the cytoplasm but the restriction of lysosomes mainly to the juxtanuclear cytoplasm of the cell body in the neuron. This feature means that autophagosomes formed in dendrites and synaptic terminal regions must be transported to the lysosome in the cell body. A previous kinetic analysis of organelle movements in cultured neurons indicated that phase-dense vesicles, containing sequestered cytoplasmic proteins and materials taken up by endocytosis, move along microtubules in the axon to the lysosomes in the cell body.18 These data suggest that fusion of autophagosomes formed in the synaptic cytoplasm with lysosomes is strictly dependent on retrograde axonal transport. It has been reported that mutations of dynein and dynactin in mice and human cause motor neuron degeneration resembling amyotrophic lateral sclerosis.19, 20 Using cultured PC12 cells, flies and mice expressing different kinds of aggregate-prone proteins, such as expanded polyQ and mutant α-synuclein, Ravikumar et al.21 demonstrated that inhibition of dynein function retarded the clearance of aggregates by inhibiting autophagosome-lysosome fusion. It has been shown recently that aggregate-prone proteins formed in the cytoplasm of cultured HeLa cells and a cultured neural cell line, are degraded by autophagy, which is also dependent on intact microtubules.22
Autophagic Vacuoles in Neurodegenerative Diseases
Morphological analyses revealed that accumulation of abnormally large number of autophagic vacuoles (autophagosomes plus autolysosomes) or the appearance of irregularly shaped autophagic vacuoles is frequently observed as a common feature in many inherited neurodegenerative diseases.23, 24, 25, 26, 27 Inclusion bodies, composed of ubiquitin-positive cytoplasmic remnants, and lipofuscin deposits, together with dispersed autophagic vacuoles and lysosomes are the primary hallmarks of the late stages of these diseases. Increased density of autophagic vacuoles seems to reflect enhanced autophagosome formation, on one hand, but their accumulation with blurred structures of sequestered materials in their lumen may also imply impaired autolysosomal degradation, on the other. In fact, these morphological features are not present in normal neurons and resemble those of cultured HEK293 cells that have been placed under starvation conditions in the presence of lysosomal proteinase inhibitors.28 Under these conditions, autophagic response is markedly enhanced, but autophagic proteolysis is simultaneously inhibited. In addition, as observed in Danon disease, which is caused by mutation of the lysosome-associated membrane protein-2 (LAMP-2), and LAMP-2-deficient mice, impairment of autophagosome-lysosome fusion also leads to unequivocal increment of autophagosomes in cardiac muscles and hepatocytes.29, 30
Autophagy in Alzheimer's, Huntington's, and Parkinson's Diseases
Protein conformational disorders, such as Alzheimer's disease (AD), Huntington's disease (HD), and Parkinson's disease (PD), are characterized by abnormally high accumulation of misfolded and/or unfolded proteins in the surviving neurons as detected at postmortem examination. In this section, we will evaluate the role of autophagy in those hereditary neurodegenerative diseases.
It has become clear that autophagy is linked to the pathogenesis of HD. HD is an autosomal dominant disorder caused by mutations of huntingtin, a cytosolic protein that has a polyglutamine (polyQ) tract in its N-terminus. In HD, abnormal expansion of polyQ caused by codon (CAG) reiterations in exon 1 of the Huntingtin gene produces mutated huntingtin with an expanded polyQ repeat (more than 37 polyQs). Mutant huntingtin with a longer polyQ tract has a stronger tendency than the wild type to form aggregates, both accelerating the onset and worsening the severity of the disease, suggesting that the progressive formation of insoluble polyQ aggregates is a key event leading to manifestation of the disease. Indeed, model mouse with mutant polyQ is associated with formation of nuclear and cytoplasmic inclusions in their neurons.31 However, recent studies revealed that globular and protofibrillar intermediates form before the organization of mature huntingtin aggregates, and that these are toxic and could lead to disturbances of genetic transcription networks and mitochondrial dysfunctions.32, 33 Then, what are the mechanisms by which autophagy clears mutant huntingtin? Ultrastructural examination of huntingtin-transfected cells showed abundant accumulation of cathepsin D-positive autophagic vacuoles with or without sequestered cellular constituents, dense lysosomes, and multilamellar and tubuloveseicular structures.26 Ravikumar et al.34 investigated whether autophagy can degrade mutant huntingtin with expanded polyQ repeats. Degradation of 74 polyQ repeats fused to the amino terminus of GFP (polyQ74-GFP) transfected into COS7 or PC12 cells was inhibited by 3-methyladenine, a specific inhibitor of autophagy, and enhanced by rapamycin. Rapamycin acts by inhibiting the mammalian target of rapamycin (mTor) kinase, which forms the core of a nutrient- and growth factor-sensitive complex that control protein synthesis, and suppresses autophagy.35 Importantly, inhibitors of autophagy enhance cell death, whereas rapamycin prevents the effects. Furthermore, once the overexpressed polyQ74-GFP forms insoluble large aggregates, the insoluble aggregates become resistant to rapamycin-induced autophagy. The data clearly demonstrate that failure to degrade polyQ expansions by autophagy is associated with accelerated progression of HD and that stimulation of autophagy in the early stages of the disease by rapamycin treatment could prevent deposition of polyQ aggregates. Rapamycin enhances the autophagic clearance of different proteins with long polyQ and polyalanine (polyA)-expanded proteins, and reduces their neurotoxicity. Thus, rapamycin and its analogs can be potentially used therapeutically for neurodegenerative diseases caused by aggregate-prone proteins.36 It has been shown that mTor is sequestered in polyQ aggregates in transgenic mice expressing mutant huntingtin and patient brains of HD. Sequestration of mTor in polyQ aggregates inhibits nuclear-cytoplasm shuttling of mTor, leading to inactivation of mTor.37 The inactivation in turn induces autophagy. Hence, co-sequestration of aggregates with mTor leads to inhibition of mTor activity, which may provide a partial explanation for accumulation of autophagic vacuoles in neurodegenerative diseases. On the other hand, the activation of autophagy via an insulin signal pathway clears accumulated polyQ proteins independent of mTor, as reported by Yamamoto et al.38 They found that aggregates of mutant huntingtin activate insulin receptor substrate-2 involving the signaling cascades of insulin and insulin-like growth factor 1. Such activation turns on class III PI3K to induce autophagy, thus contributing to clearance of huntingtin aggregates.38
Invariably, AD is the most prevalent form of neurodegenerative diseases with dementia and associates with extracellular deposition of beta-amyloid (Aβ). Presenilin-1 (PS1) is one of several proteins linked to early-onset familial AD, and together with PS2, plays a catalytic role in the γ-secretase complex necessary for intermediate proteolysis of the amyloid precursor protein (APP) followed by liberation of Aβ. Although it has been noticed that autophagic vacuoles accumulate in hippocampal and prefrontal cortical pyramidal neurons of Alzheimer-type dementia,23, 24 the mechanism remains unclear. Wilson et al.39 found the formation of enlarged late endosome-like structures, including α- and β-synuclein, in the perikarya of PS1−/− primary neurons and hippocampal tissue of patients with the Levy body variant of AD. Formation of such organelles is rescued by exogenous expression of not only wild-type PS but also dominant-negative PS1 lacking its activity, indicating that PS1 has another function besides γ-secretase. Esselens et al.40 demonstrated the accumulation of telencephalin, a neural specific intercellular adhesion molecule known to interact with PS1, in vacuoles positive for Atg12 and LC3, but not cathepsin D, in PS1−/− hippocampal neurons. Similar to the report of Wilson et al,39 the formation of such vacuoles was suppressed by not only wild-type but also mutant PS1. Furthermore, Esselens et al.40 used cathepsin D knockout mice to show the degradation of telencephalin in lysosomes. Collectively, these results suggest that PS1 might play important roles in autophagosome and lysosomal fusion step. Recently, Yu et al.41 reported the role of autophagy in Aβ production. Their exhaustive electron and immunoelectron microscopic analyses revealed accumulation of LC3-positive autophagic vacuoles in brains of AD patients and in AD model mice, neural cell lines, and in a non-neural APP-expressing cell line, and also the localization of PS1, Aβ40, Aβ42, and necastrin on internal and limiting membrane components of autophagic vacuoles. Further, they found that induction of autophagy evoked Aβ production, and inversely, inhibition of autophagy suppressed Aβ production. Finally, they observed the PS1-dependent γ-secretase activity in biochemical isolated autophagic vacuoles. Based on these findings, they proposed a novel mechanism for the generation of Aβ via autophagy that emphasized the prominent role of autophagy in AD pathogenesis.41
PD is a neurodegenerative disorder associated with progressive loss of dopaminergic neurons of the substantia nigra and locus coeruleus. The major clinical symptoms of PD are body rigidity, hypokinesia, and postural instability associated with trembling extremities.42 Pathological examination shows marked accumulation of cytoplasmic inclusions of proteinaceous material with lipids called Lewy bodies. Lewy bodies consist of lipids, ubiquitin, enzymes involved in ubiquitin-related pathways, neurofilament proteins, α-synuclein, synphilin-1, and other entangled proteins. Mutations in the gene encoding α-synuclein, which is localized in presynaptic terminals and is abundantly present in Lewy bodies, are identified in certain cases of familial PD.43, 44 α-Synuclein is a protein of unknown function and a major component of Lewy bodies. Among three point mutations in α-synuclein causing an autosomal dominant form of familial PD, two mutations of α-synuclein (A53T and A30P) have been studied extensively. These α-synuclein mutants have a stronger tendency to form fibrils than wild-type α-synuclein. Hence, similar to huntingtin with abnormal polyQ expansion, misfolded or aggregated α-synuclein is believed to cause cell toxicity and inhibits the ubiquitin-proteasome system. Lewy bodies may contribute to aggregation of α-synuclein into inclusions to moderate its toxicity.45, 46 It has been reported recently that autophagic-lysosomal dysfunction may be also involved in PD. Using stable PC12 transfectants expressing wild-type and A53T mutant α-synuclein, Stefanis et al.47 showed that marked accumulation of autophagic vacuoles and impairment of lysosomal and ubiquitin-proteasome functions are principal phenotypes in the cells. On the other hand, clearance of mutant α-synuclein is strongly dependent on both ubiquitin-proteasomes and macroautophagy,48 but not chaperone-mediated autophagy capable of degrading wild-type α-synuclein efficiently.49
Impairment of Autophagy in Neurons
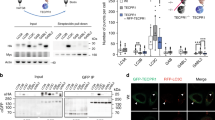
Recently, our group and Mizushima's group investigated the pathophysiological roles of basal or constitutive autophagy in the brain.50, 51 For this purpose, we generated neuron-specific autophagy-deficient mice (Atg7F/F:Nes mice) by crossing Atg7-conditional knockout mice (Atg7F/F) with transgenic mice expressing the Cre recombinase under the control of the neuron-specific Nestin (Nes) promoter, Nes-Cre. We found that mice lacking Atg7 (i.e., autophagy) in the central nervous system exhibited various behavioral deficits, such as abnormal limb-clasping reflexes and reduction of coordinated movement, and died within 28 weeks after birth. Histological analysis showed that Atg7-deficiency was associated with neuronal loss in the cerebral and cerebellar cortices. Intriguingly, Atg7-deficient neurons showed abundant accumulation of polyubiquitylated proteins, which appeared as inclusion bodies whose size and number increased with aging (Figure 3), but had functionally intact proteasomes, whose impairment is generally known to cause abnormal ubiquitin-mediated proteolysis.50 Hara et al.51 also reported that almost all these phenotypes, if not all, were observed in neural-specific mice deficient in Atg5, another autophagy-essential gene. Thus, many of the critical symptoms seen in neural-specific autophagy-deficient mice are similar to those of patients with neurodegenerative disorders.
Ubiquitin-positive inclusions in autophagy-deficient neurons. The presence of ubiquitin-positive dots was examined immunohistochemically in several regions of the brain including cerebral cortex, cerebellum (Purkinje cells), hippocampus, amygdala, hypothalamus, caudate putamen of Atg7F/+:Nes (left panels), and Atg7F/F:Nes (right panels) mice. Note the presence of numerous ubiquitin dots in the amygdala and hypothalamus of representative mutants. Bars, 100 μm in the panel of cerebral cortex, and 50 μm in others
Histological analyses of the brains of Atg7F/F:Nes mice revealed loss of specific neurons, such as pyramidal neurons in the cerebral cortex and hippocampus, and Purkinje cells in the cerebellum. Unexpectedly, immunohistological analysis using anti-ubiquitin antibody identified ubiquitin-positive proteinaceous aggregates throughout the brain, although the staining intensity varied from one region to another. Few ubiquitin-positive inclusions were recognized in brain regions with evident neuronal loss, whereas many ubiquitin inclusions were noted in areas with barely any neuronal loss such as the hypothalamus. Although we could not determine whether neuronal death is due to accumulation and subsequent inclusion formation of ubiquitylated proteins, neurons with large inclusions survived. Conversely, large pyramidal neurons and Purkinje cells seem vulnerable to ubiquitylated proteins and die before the formation of large inclusions. Whether the formation of inclusion bodies in neurons is protective or toxic is under debate, although emerging evidence emphasizes that protein aggregation can be a protective mechanism.32, 33
Although accumulation of ubiquitylated proteins and cell death were noted in autophagy-deficient hepatocytes14 and neurons,50, 51 such phenotypes were not observed in growing cells such as mouse embryonic fibroblasts (MEFs) and astroglial cells, irrespective of autophagy deficiency. Thus, it seems that autophagy is not required in rapidly dividing cells, at least with respect to multiplication of these cells. These results might also reflect the difference in autophagic activity among cell types. It is possible that the cell division cycle results in dilution of ubiquitylated proteins in autophagy-deficient MEFs, preventing their accumulation. Alternatively, other degradation pathways, such as chaperone-mediated autophagy, could contribute to degradation of long-lived proteins in growing MEFs. Considered together, it is clear that macroautophagy (the massive autophagy pathway discussed here) plays important roles in proteolysis in quiescent cells.
Autophagy deficiency is considered to result in delays of global turnover of cytoplasmic components, resulting in accumulation of misfolded and/or unfolded proteins followed by formation of inclusion bodies. However, a recent report showed that p62/SQSTM1 harboring a ubiquitin binding domain, interacted with LC3 and was degraded via autophagy.52 We also obtained similar results and found that p62 plays a critical role in the formation of ubiquitin-positive aggregates by impaired autophagy (unpublished data). These results imply that ubiquitylated unfavorable proteins might be selectively sequestered into autophagosomes in part via p62 (which may retain its shuttling ability of ubiquitylated proteins). In either case (non-selective or selective degradation of ubiquitylated proteins by autophagy), our results indicate that autophagy operates not only as a supplier of amino acids under nutrient-poor conditions but also as a house cleaner of damaged proteins under nutrient-rich conditions.
Concluding Remarks
Considering the role of autophagy in neurodegenerative diseases, it is possible to align time-dependent enhancement and inactivation of autophagy. First, misfolded and/or unfolded protein aggregates formed in neurons cause sequestration of mTor along with the aggregates, leading to significant inactivation of mTor, which stimulates autophagy (autophagosome formation). Second, if the size of aggregates is small enough and the degree of aggregation is moderate, the aggregates are engulfed into autophagosomal lumen and subsequently degraded via autophagy. Unless the amounts of aggregates surpass the clearance capacity of autophagosomes, maximal activation of autophagy by pharmaceutical agents such as rapamycin could be effective in preventing the progression of the disease. It is noteworthy that the expression of aggregation-prone protein(s) observed frequently in familial neurodegenerative disorders is not required for the formation of inclusions associated with impaired autophagy, suggesting the involvement of autophagy even in sporadic neurodegenerative diseases. On the other hand, autophagic activity of rat liver decreases with aging and this decrease conversely correlates with an increase in the accumulation of oxidized proteins.36 Thus, age-dependent onset of neurodegenerative diseases most likely correlates with the age-dependent decline of autophagic activity. It is generally accepted that deficits of the proteasome-ubiquitin system are linked to various neurodegenerative disorders. Intriguingly, however, no obvious inhibition of the ubiquitin-proteasome system occurs in the mouse brain lacking autophagy, providing compelling evidence that constitutive autophagy plays a prominent role in neuronal survival, independent of proteasome function (Figure 4). In this context, we stress that proteasome inhibition induces augmented autophagy in order to eliminate unnecessary accumulated proteins, probably compensating the loss of proteasome functions.22 Thus, the two major proteolytic pathways naturally differ in the cooperative responses in cells. We emphasize the importance of autophagy as a process with adaptive and flexible responses. Finally, a better understanding of basal or constitutive autophagy, which maintains basal activity of macroautophagy in neurons, may help in the design of new strategies to prevent neurodegenerative diseases.
A schematic diagram of protein destruction pathways mediated by the proteasome and autophagy. The majority of cellular proteins, if not all, are polyubiquitylated before hydrolysis by the proteasome (an ATP-dependent proteolytic complex). Similarly, unfolded/misfolded proteins generated by environmental stresses or genetic mutations are discarded after polyubiquitylation by the same proteolytic system (a). Aging-related decline in autophagic activity causes accumulation of highly ubiquitylated proteins, which are recovered as both soluble and insoluble forms (b). As autophagy could feed both ubiquitylated and unubiquitylated protein(s), it is not clear at present whether polyubiquitylated aggregates/inclusions in autophagy-deficient neurons are formed consequent to impairment of degradation of unubiquitylated proteins or aggressively polyubiquitylated proteins. In addition, further work is needed to determine whether the two proteolytic systems (autophagy and proteasomes) work independently or cooperatively, and whether autophagy and the proteasome feed a similar set of normal and/or misfolded/unfolded proteins in general. Red text: protein dynamics associated with autophagy deficiency, Ub: ubiquitin
Abbreviations
- AD:
-
Alzheimer's disease
- APP:
-
amyloid precursor protein
- Aβ:
-
beta-amyloid
- GFP:
-
green fluorescent protein
- HD:
-
Huntington's disease
- LC3:
-
microtubule-associated protein 1 light chain 3/MAP1LC3
- mTor:
-
mammalian target of rapamycin
- PD:
-
Parkinson's disease
- PS1:
-
presenilin-1
- SDH:
-
succinate dehydrogenase
- TCA:
-
tricarboxylic acid
References
Goldberg AL . Protein degradation and protection against misfolded or damaged proteins. Nature 2003; 426: 895–899.
Levine B, Klionsky DJ . Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004; 6: 463–477.
Cuervo AM . Autophagy: in sickness and in health. Trends Cell Biol 2004; 14: 70–77.
Mizushima N, Ohsumi Y, Yoshimori T . Autophagosome formation in mammalian cells. Cell Struct Funct 2002; 27: 421–429.
Takeshige K, Baba M, Tsuboi S, Noda T, Ohsumi Y . Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J Cell Biol 1992; 119: 301–311.
Mizushima N, Yamamoto A, Matsui M, Yoshimori T, Ohsumi Y . In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell 2004; 15: 1101–1111.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–5728.
Kabeya Y, Mizushima N, Yamamoto A, Oshitani-Okamoto S, Ohsumi Y, Yoshimori T . LC3, GABARAP and GATE16 localize to autophagosomal membrane depending on form-II formation. J Cell Sci 2004; 117: 2805–2812.
Tanida I, Sou YS, Ezaki J, Minematsu-Ikeguchi N, Ueno T, Kominami E . HsAtg4B/HsApg4B/autophagin-1 cleaves the carboxyl termini of three human Atg8 homologues and delipidates microtubule-associated protein light chain 3- and GABAA receptor-associated protein-phospholipid conjugates. J Biol Chem 2004; 279: 36268–36276.
Ichimura Y, Kirisako T, Takao T, Satomi Y, Shimonishi Y, Ishihara N et al. A ubiquitin-like system mediates protein lipidation. Nature 2000; 408: 488–492.
Tanida I, Tanida-Miyake E, Ueno T, Kominami E . The human homolog of Saccharomyces cerevisiae Apg7p is a Protein-activating enzyme for multiple substrates including human Apg12p, GATE-16, GABARAP, and MAP-LC3. J Biol Chem 2001; 276: 1701–1706.
Tanida I, Tanida-Miyake E, Komatsu M, Ueno T, Kominami E . Human Apg3p/Aut1p homologue is an authentic E2 enzyme for multiple substrates, GATE-16, GABARAP, and MAP-LC3, and facilitates the conjugation of hApg12p to hApg5p. J Biol Chem 2002; 277: 13739–13744.
Mortimore GE, Hutson NJ, Surmacz CA . Quantitative correlation between proteolysis and macro- and microautophagy in mouse hepatocytes during starvation and refeeding. Proc Natl Acad Sci USA 1993; 80: 2179–2183.
Komatsu M, Waguri S, Ueno T, Iwata J, Murata S, Tanida I et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol 2005; 169: 425–434.
Tsukada M, Ohsumi Y . Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett 1993; 333: 169–174.
Kuma A, Hatano M, Matsui M, Yamamoto A, Nakaya H, Yoshimori T et al. The role of autophagy during the early neonatal starvation period. Nature 2004; 432: 1032–1036.
Lum JJ, Bauer DE, Kong M, Harris MH, Li C, Lindsten T et al. Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell 2005; 120: 237–248.
Hollenbeck PJ . Products of endocytosis and autophagy are retrieved from axons by regulated retrograde organelle transport. J Cell Biol 1993; 121: 305–315.
Puls I, Jonnakuty C, LaMonte BH, Holzbaur EL, Tokito M, Mann E et al. Mutant dynactin in motor neuron disease. Nat Genet 2003; 33: 455–456.
Hafezparast M, Klocke R, Ruhrberg C, Marquardt A, Ahmad-Annuar A, Bowen S et al. Mutations in dynein link motor neuron degeneration to defects in retrograde transport. Science 2003; 300: 808–812.
Ravikumar B, Acevedo-Arozena A, Imarisio S, Berger Z, Vacher C, O'Kane CJ et al. Dynein mutations impair autophagic clearance of aggregate-prone proteins. Nat Genet 2005; 37: 771–776.
Iwata A, Riley BE, Johnston JA, Kopito RR . HDAC6 and microtubules are required for autophagic degradation of aggregated huntingtin. J Biol Chem 2005; 280: 40282–40292.
Okamoto K, Hirai S, Iizuka T, Yanagisawa T, Watanabe M . Reexamination of granulovacuolar degeneration. Acta Neuropathol (Berlin) 1991; 82: 340–345.
Cataldo AM, Hamilton DJ, Barnett JL, Paskevich PA, Nixon RA . Properties of the endosomal-lysosomal system in the human central nervous system: disturbances mark most neurons in populations at risk to degenerate in Alzheimer's disease. J Neurosci 1996; 16: 186–199.
Anglade P, Vyas S, Javoy-Agid F, Herrero MT, Michel PP, Marquez J et al. Apoptosis and autophagy in nigral neurons of patients with Parkinson's disease. Histol Histopathol 1997; 12: 25–31.
Kegel KB, Kim M, Sapp E, McIntyre C, Castano JG, Aronin N et al. Huntingtin expression stimulates endosomal-lysosomal activity, endosome tubulation, and autophagy. J Neurosci 2000; 20: 7268–7278.
Petersen A, Larsen KE, Behr GG, Romero N, Przedborski S, Brundin P et al. Expanded CAG repeats in exon 1 of the Huntington's disease gene stimulate dopamine-mediated striatal neuron autophagy and degeneration. Hum Mol Genet 2001; 10: 1243–1254.
Tanida I, Minematsu-Ikeguchi N, Ueno T, Kominami E . Lysosomal turnover, but not a cellular level, of endogenous LC3 is a marker for autophagy. Autophagy 2005; 1: 84–91.
Nishino I, Fu J, Tanji K, Yamada T, Shimojo S, Koori T et al. Primary LAMP-2 deficiency causes X-linked vacuolar cardiomyopathy and myopathy (Danon disease). Nature 2000; 406: 906–910.
Tanaka Y, Guhde G, Suter A, Eskelinen EL, Hartmann D, Lullmann-Rauch R et al. Accumulation of autophagic vacuoles and cardiomyopathy in LAMP-2-deficient mice. Nature 2000; 406: 902–906.
Ikeda H, Yamaguchi M, Sugai S, Aze Y, Narumiya S, Kakizuka A . Expanded polyglutamine in the Machado–Joseph disease protein induces cell death in vitro and in vivo. Nat Genet 1996; 13: 196–202.
Sanchez I, Mahlke C, Yuan J . Pivotal role of oligomerization in expanded polyglutamine neurodegenerative disorders. Nature 2003; 421: 373–379.
Arrasate M, Mitra S, Schweitzer ES, Segal MR, Finkbeiner S . Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 2004; 431: 805–810.
Ravikumar B, Duden R, Rubinsztein DC . Aggregate-prone proteins with polyglutamine and polyalanine expansions are degraded by autophagy. Hum Mol Genet 2002; 11: 1107–1117.
Inoki K, Corradetti MN, Guan KL . Dysregulation of the TSC-mTOR pathway in human disease. Nat Genet 2005; 37: 19–24.
Bergamini E . Autophagy: a cell repair mechanism that retards ageing and age-associated diseases and can be intensified pharmacologically. Mol Aspects Med 2006; 27: 403–410.
Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat Genet 2004; 36: 585–595.
Yamamoto A, Cremona ML, Rothman JE . Autophagy-mediated clearance of huntingtin aggregates triggered by the insulin-signaling pathway. J Cell Biol 2006; 172: 719–731.
Wilson CA, Murphy DD, Giasson BI, Zhang B, Trojanowski JQ, Lee VM . Degradative organelles containing mislocalized alpha-and beta-synuclein proliferate in presenilin-1 null neurons. J Cell Biol 2004; 165: 335–346.
Esselens C, Oorschot V, Baert V, Raemaekers T, Spittaels K, Serneels L et al. Presenilin 1 mediates the turnover of telencephalin in hippocampal neurons via an autophagic degradative pathway. J Cell Biol 2004; 166: 1041–1054.
Yu WH, Cuervo AM, Kumar A, Peterhoff CM, Schmidt SD, Lee JH et al. Macroautophagy – a novel Beta-amyloid peptide-generating pathway activated in Alzheimer's disease. J Cell Biol 2005; 171: 87–98.
Jenner P, Olanow CW . Understanding cell death in Parkinson's disease. Ann Neurol 1998; 44: S72–S84.
Kruger R, Kuhn W, Muller T, Woitalla D, Graeber M, Kosel S et al. Ala30Pro mutation in the gene encoding alpha-synuclein in Parkinson's disease. Nat Genet 1998; 18: 106–108.
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A et al. Mutation in the alpha-synuclein gene identified in families with Parkinson's disease. Science 1997; 276: 2045–2047.
Chung KK, Dawson VL, Dawson TM . The role of the ubiquitin-proteasomal pathway in Parkinson's disease and other neurodegenerative disorders. Trends Neurosci 2001; 24: S7–S14.
Mouradian MM . Recent advances in the genetics and pathogenesis of Parkinson disease. Neurology 2002; 58: 179–185.
Stefanis L, Larsen KE, Rideout HJ, Sulzer D, Greene LA . Expression of A53T mutant but not wild-type alpha-synuclein in PC12 cells induces alterations of the ubiquitin-dependent degradation system, loss of dopamine release, and autophagic cell death. J Neurosci 2001; 21: 9549–9560.
Webb JL, Ravikumar B, Atkins J, Skepper JN, Rubinsztein DC . Alpha-Synuclein is degraded by both autophagy and the proteasome. J Biol Chem 2003; 278: 25009–25013.
Cuervo AM, Stefanis L, Fredenburg R, Lansbury PT, Sulzer D . Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 2004; 305: 1292–1295.
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 2006; 441: 880–884.
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 2006; 441: 885–889.
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 2005; 171: 603–614.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by E Baehrecke
Rights and permissions
About this article
Cite this article
Komatsu, M., Ueno, T., Waguri, S. et al. Constitutive autophagy: vital role in clearance of unfavorable proteins in neurons. Cell Death Differ 14, 887–894 (2007). https://doi.org/10.1038/sj.cdd.4402120
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402120
Keywords
This article is cited by
-
Epigenetic regulation of autophagy by histone-modifying enzymes under nutrient stress
Cell Death & Differentiation (2023)
-
The dual roles of autophagy and the GPCRs-mediating autophagy signaling pathway after cerebral ischemic stroke
Molecular Brain (2022)
-
Estrogen and estrogen receptors chauffeur the sex-biased autophagic action in liver
Cell Death & Differentiation (2020)
-
Characterization of starvation-induced autophagy in cerebellar Purkinje cells of pHluorin-mKate2-human LC3B transgenic mice
Scientific Reports (2020)
-
Network approach identifies Pacer as an autophagy protein involved in ALS pathogenesis
Molecular Neurodegeneration (2019)