Abstract
Upon DNA damage signaling, p73, a member of the p53 tumor suppressor family, accumulates to support transcription of downstream apoptotic genes. p73 interacts with Yes-associated protein 1 (Yap1) through its PPPY motif, and increases p73 transactivation of apoptotic genes. The ubiquitin E3 ligase Itch, like Yap1, interacts with p73. Given the fact that both Itch and Yap1 bind p73 via the PPPY motif, we hypothesized that Yap may also function to stabilize p73 by displacing Itch binding to p73. We show that the interaction of Yap1 and p73 was necessary for p73 stabilization. Yap1 competed with Itch for binding to p73, and prevented Itch-mediated ubiquitination of p73. Treatment of cells with cisplatin leads to an increase in p73 accumulation and induction of apoptosis, but both were dramatically reduced in the presence of Yap1 siRNA. Altogether, our findings attribute a central role to Yap1 in regulating p73 accumulation and function under DNA damage signaling.
Similar content being viewed by others
Main
p73 is a member of the p53 family of transcription factors. This family of proteins, which includes p53, p63 and p73, shares similar structure, sequence and function.1 In response to DNA damage and other cell stress, p53 activates target genes that induce cell cycle arrest, senescence, or apoptosis. Owing to their structural similarity with p53, p73 and p63 can also bind to p53-responsive elements, and activate target genes. Interestingly, it has been shown that in mouse fibroblasts p63 and p73 are required for the ability of p53 to induce apoptosis.2 In addition, p73 is induced by irradiation and DNA damaging agents and can induce apoptosis independently of p53.3, 4, 5 p73 is regulated at both the transcriptional and post-translational levels. At the level of transcription, p73 gene expression is regulated by E2F1. It was found that the p73 promoter region contains at least three E2F1-binding sites, and indeed a direct binding of E2F1 to the p73 promoter results in a significant stimulation of p73 transcription.6, 7
The post-translational regulation of p73 activity is complex, and is still incompletely understood. Several mechanisms are involved in the stabilization and modification of p73 activity in response to DNA damage (reviewed by Oberst et al.8). A key player is the c-Abl nonreceptor tyrosine kinase that upon DNA damage insults triggers phosphorylation of p73 and increases its protein level and its proapoptotic activity.4, 5, 9 c-Abl phosphorylation of p73 also causes it to become associated with the nuclear matrix.10 Activation of p73 by c-Abl has been found to involve the p38 MAP kinase pathway.11 In addition to phosphorylation, p73 can be modified through SUMOylation12 and acetylation by p300.13
Phosphorylated p73 interacts with the prolyl isomerase Pin1 that is capable of specifically recognizing phosphorylated serine or threonine residues followed by proline (pS/pT-P) and catalyzing conformational changes in its substrates,14, 15 leading to p73 acetylation by p300.16, 17 The p300-mediated acetylation of p73 protects it against ubiquitinylation and the promyelocytic leukemia (PML) protein regulates its stability by positively modulating its acetylation levels.18
The transcriptional co-activator Yes-associated protein 1 (Yap1)19 determines p73 gene targeting in response to DNA damage. When cells are exposed to cisplatin, a DNA damaging agent, Yap1 translocates to the nucleus in a p73-dependent manner. In the nucleus, Yap1 promotes p73-dependent apoptosis through the specific and selective co-activation of the p53AIP1 gene, an apoptotic p73 target gene. To do so Yap1 requires PML and nuclear body (NB) localization. On the promoter, Yap1 functions to recruit p300 that mediates p73 and histone acetylation and transcriptional activation. Interestingly, silencing of Yap1 markedly impairs p73 accumulation in response to DNA damage, suggesting that Yap1 might also be involved in p73 stabilization.20
The 65 kDa Yap1 is a modular adapter protein with multiple protein interaction domains that was originally isolated based on its ability to bind the SH3 domain of the Src family kinase c-Yes.21 Although initially Yap1 was thought to play a role in the anchoring or targeting of c-Yes to specific subcellular compartments, additional studies however suggest that Yap1 plays a role of a transcription co-activator in regulating gene expression through direct association with a broad array of transcription factors. 20, 22, 23, 24 In addition to its SH3-binding motif, Yap1 contains a proline-rich amino terminus, a WW domain, a coiled-coil and a PDZ-binding motif at the extreme COOH terminus.25, 26, 27 Yap1 binds to p73 through its WW domain and the PPPY motif of p73.28 This interaction is required for the ability of Yap1 to co-activate p73-responsive genes.28 Yap1 also contains a 14-3-3 interacting motif. Upon phosphorylation by Akt at serine 127, 14-3-3 is recruited and promotes Yap1 localization to the cytoplasm. As a result, it loses its main function as a co-activator in the nucleus.29 Inhibition of Akt activity potentiates the nuclear relocalization of Yap1 to induce apoptosis by p73.20, 29
Accumulating evidence suggests that p73 turnover is regulated through both ubiquitin-independent and ubiquitin-dependent degradation pathways. p73 is prone to ubiquitin-independent 20S proteasomal degradation and that this process is regulated by NAD(P)H quinone oxidoreductase 1 (NQO1) together with NADH.30 Recently, Rossi et al. have shown that Itch, a human ubiquitin-protein ligase, which belongs to the Nedd4-like E3 family containing a WW domain,31 binds and ubiquitinates p73 via its PPPY motif and determines its rapid proteosome-dependent degradation in a ubiquitin-dependent manner.32 In response to DNA damage the Itch protein level dramatically decreases with concomitant accumulation of p73.
Given the fact that both Yap1 and Itch bind the very same motif of p73, it appeared possible that upon Yap1 binding to the PPPY motif, this motif is sequestered to prevent access to Itch. This mechanism, whereby Yap1 enables p73 to escape the Itch E3 ligase, provides an explanation for the effect of Yap1 on p73 stability. This hypothesis was tested and confirmed in this study. We show that Yap1 can lead to a substantial increase in the amount and half-life of transfected and endogenous p73 protein. Yap1 inhibits proteasomal degradation of p73 provided that Yap1–p73 physical interaction is not hampered. Furthermore, we demonstrate that knocking down Yap1 results in compromised p73 accumulation and apoptosis in response to DNA damage insults. Our findings attribute to Yap1 a key role in induction of apoptosis in response to DNA-damage stress.
Results
Yap1 upregulates the p73α protein level
p73α physically interacts with Yap1.28 To study whether Yap1 affects p73 accumulation, we performed co-transfection experiments to express p73α alone or in combination with Yap1 in various cell lines. Twenty-four hours after transfection cell extracts were subjected to immunoblot analysis. Co-expression of p73 with increasing amounts of Yap1 gave rise to an increase in the steady-state levels of p73α (Figure 1a). Similar results were obtained in HCT116 p53−/− (data not shown) and human large cell lung carcinoma H1299 (Figure 1b). H1299, like the HCT116 cell line, does not express p53, suggesting that the role of Yap1 in p73 accumulation is p53 independent. To knockdown the expression of Yap1, we generated the plasmid pSuper Yap1, which encodes a Yap1-specific siRNA. As shown in Figure 1c, pSuper Yap1 was effective in knocking down Yap1, whereas the empty pSuper plasmid, or the nonspecific plasmid pSuper X, encoding siRNA against HBV X protein, had no effect on Yap1 protein levels. When the pSuper Yap1 plasmid was co-transfected with Yap1, Yap1 levels were reduced, and this resulted in a marked decrease in the level of p73α protein up to 72 h after transfection (Figure 1d).
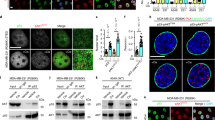
Ectopic Yap1 expression upregulates p73α protein level. (a) 293 cells were transfected with HA-p73α alone or together with increasing amounts of Flag-Yap1. Twenty-four hours after transfection, cell lysates were subjected to Western blotting analysis using anti-HA to detect HA- p73α protein level. The same blot was re-probed with anti-Flag and with anti-tubulin antibody to show equal loading (b) H1299 p53−/− cells were transfected with either HA-p73α alone or together with 250 ng of Flag-Yap1. Cells were then treated the same as in (a). (c) 293 cells were transfected with Flag-Yap1 in the presence of the indicated pSuper siRNA constructs. Cells were harvested after 48 h and were subjected to Western blot analysis. (d) 293 Cells were transiently transfected with HA-p73α alone or together with Yap1 without or with pSUPER Yap1 encoding Yap1-specific siRNA. At 24, 48 and 72 h after transfection cells were harvested and subjected to Western blot analysis as in (a)
To examine whether Yap1 overexpression increases the endogenous level of p73 as well, we overexpressed Yap1 in HEK293 cells and analyzed the p73 protein level by a specific p73 antibody. The obtained data revealed that Yap1 overexpression led to a significant accumulation of endogenous p73 (Figure 2a) (similar results were obtained with the H1299 cells (data not shown). To further confirm the importance of Yap1 in controlling endogenous p73 protein level, we knocked down the endogenous Yap1 level by pSUPER plasmid expressing Yap1-specific siRNA. Under this condition, a reduction in the endogenous level of Yap1 resulted in lower level of endogenous p73α (Figure 2b). These results suggest that Yap1 regulates the level of p73α.
Yap1 increases the steady-state levels of endogenous p73α (a) 293 cells were transfected with Flag-Yap1 and 24 h after transfection endogenous p73 protein level was detected with anti-p73 BL906 antibody. Yap1 was detected by anti-Flag, and anti-tubulin detection was used as a loading control. (b) 293 cells were transiently transfected without or with pSUPER Yap1 encoding Yap1-specific siRNA. Forty-eight hours after transfection endogenous p73 protein level was detected using the indicated antibodies
Yap1–p73 physical interaction is critical for p73 stabilization
p73α associates in vitro and in vivo with Yap1 via the WW domain of Yap1 and the PPPY motif of p73.28 To test the possibility that Yap1–p73 physical interaction is necessary for Yap1-dependent p73 accumulation, we used the Y487F p73α mutant. A mutation at this position was previously demonstrated to impair Yap1 binding to p73.28 Interestingly, this p73 mutant failed to accumulate in response to Yap1 overexpression (Figure 3a). Similar results were obtained in HCT116 p53−/− cells (Figure 3b, lanes 3 and 4). Taken together, our data suggest that the physical interaction between Yap1 and p73α is important for Yap1 to support p73 accumulation.
Yap1–p73 physical interaction is critical for p73 stabilization. (a) 293 cells were co-transfected with HA-p73α Y487F in the presence or absence of Flag-Yap1. Twenty-four-hour post-transfection cells were harvested and cell lysates were subjected to Western blot analysis. (b) HCT116 p53−/− cells were co-transfected with HA-p73α and HA-p73α Y487F in the presence or absence of Flag-Yap1. Cells were then treated the same as in (a)
Yap1 prolongs p73 half-life
The observed correlation between Yap1 expression and p73 accumulation suggests that Yap1 supports p73 accumulation by preventing its proteasomal degradation. To investigate this possibility, we measured p73 half-life in the presence or absence of overexpressed Yap1. HEK293 cells were transiently co-transfected with the expression plasmid for HA-p73 with or without the Flag-Yap1 expression plasmid. Twenty-four hours after transfection, cells were treated with cycloheximide (CHX) to inhibit protein synthesis. Whole-cell lysates were prepared at different time points and subjected to immunoblotting with the anti-HA antibody. This experiment revealed that overexpression of Yap1 significantly prolonged the half-life of the transfected p73 (Figure 4a). Next, we conducted experiments to determine the half-life of the endogenous p73 under Yap1 overexpression. The data revealed that Yap1 increased the half-life of the endogenous p73 as well (Figure 4b). Thus, these results indicate that Yap1 prolongs p73 half-life.
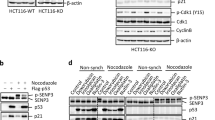
Yap1 prolongs p73α half-life. (a) 293 cells were transfected with plasmids expressing HA-p73α, along with empty vector or Flag-Yap1, as indicated. Twenty-four hours after transfection cells were treated with 20 μg/ml CHX for different time points. Equal amounts of total protein lysates were subjected to Western blotting with the indicated antibodies. (b) 293 cells were transfected with Flag-Yap1 or empty vector and were treated as in (a). Endogenous p73 level was detected using the anti-p73 BL906 antibody. (c) 293 cells were transfected with plasmids expressing HA-p73α Y487F along with Flag-Yap1 or empty vector and were treated as in (a). 35S pulse chase: 293 cells (d) and H1299 cells (e) were transfected with HA-p73α together with Flag-Yap1 (+Yap) or with an empty vector (−Yap). Twenty-four-hour post-transfection, cells were starved in a Met-free medium and then were labeled with 80 μCi/ml of 35S. Unlabeled Met (2%) were added and cells were collected at the indicated time points. IP was performed with anti-HA. Immunoprecipitates were washed and run on SDS-PAGE and detected by autoradiography. (f) 293 cells were treated with DMSO or 25 μM of MG132 for 2 h, respectively, and the cell extracts were analyzed by Western blotting for the indicated proteins. (g) 293 cells were transfected with plasmids expressing HA-p73α and Flag-Yap1, together or alone. After 24 h, cells were treated with 25 μM of the proteosome inhibitor MG132 or DMSO for 2 h. Cells lysates were collected and analyzed by Western blot with the indicated antibodies. (h) 293 cells were transfected with plasmids expressing HA-p73α Y487F and Flag-Yap1, together or alone. After 24 h, cells were treated with 25 μM of the proteasome inhibitor MG132 for different time points. Cells lysates were collected and analyzed by Western blot with the indicated antibodies
To determine whether Yap1–p73 interaction is the sole requirement for the increase in p73 half-life, we measured the half-life of the p73 Y487F mutant that is impaired in Yap1 binding. Consistent with previous studies,32 during the 8-h measurement period, this p73 mutant was found to be relatively stable. This is due to the fact that the Y487F mutation prevents binding of p73 to the E3 ligase Itch and leads to a significant decrease in ubiquitination. As expected, the stability of this p73 mutant was not affected by overexpressing Yap1 (Figure 4c). We further confirmed the role of Yap1 in p73 stabilization by pulse chase analysis in two different cell lines (Figure 4d and e).
To assess whether the effect of Yap1 is mediated through interference with the proteasomal degradation of p73, we first evaluated the effect of the proteasomal inhibitor MG132 on endogenous p73. Treatment of 293 cells with MG132 resulted in an increase in the level of endogenous p73 (Figure 4f), indicating that p73 is degraded through the proteasomal pathway, in agreement with a previous report.33 Notably, Yap1 protein level was also stabilized by MG132 treatment. Next, we evaluated the effect of MG132 on the transfected p73 cells in the presence or absence of Yap1 (Figure 4g). MG132 treatment resulted in an increased level of p73 expression in cells transfected with HA-p73α alone (Figure 4g, compare lanes 1 and 3). The accumulated level of p73 was comparable to that obtained in the presence of Yap1 in MG132-untreated cells. MG132, however, did not elicit a further increase in p73 above the levels obtained by Yap1 (Figure 4g, compare lanes 2 and 3). In contrast, MG132 did not affect the level of the p73α Y487F mutant, which is refractory to Itch and Yap1 (Figure 4h). This suggests that Yap1 and MG132 each mediate p73 accumulation through a similar mechanism, namely by inhibiting the proteasomal degradation of p73.
Yap1 competes with Itch for binding to p73
Itch, a human ubiquitin protein ligase bearing a WW domain, binds and ubiquitinates p73 and induces its rapid proteasomal degradation.32 The WW domain of Itch binds to p73α via the PPPY domain at position Y487, which is identical to the position where the WW domain of Yap1 binds.28 As both WW proteins bind to the same p73 PPPY box, we predicted that Yap1 stabilizes p73 by competing with Itch for the binding to p73α. To test this possibility, we treated 293 cells with MG-132 and then we co-immunoprecipitated p73 and Itch in the presence of an increasing amount of Yap1. In the absence of Yap1, Itch was co-immunoprecipitated with p73α, however, the level of p73α-associated Itch was decreased in the presence of Yap1, in a Yap1 dose-dependent manner (Figure 5a). In a reciprocal experiment, we co-immunoprecipitated Yap1 in the presence of an increasing amount of Itch. Yap1 was co-immunoprecipitated with endogenous p73, however, the p73–Yap1 interaction was dramatically reduced when Itch was added, in a dose-dependant manner (Figure 5b). To further examine the competition between Yap1 and Itch for p73, we immunoprecipitated endogenous p73. As expected, Itch was associated with p73. However, with the addition of Yap1, p73-Itch binding was reduced, in favor of p73-Yap1 binding (Figure 5c).
Yap1 competes with Itch for the binding to p73. (a) 293 cells were transfected with Flag-p73α, myc-Itch and increasing amounts of GFP-Yap1. Twenty-four hours after transfection cells were treated with 20 μg/ml MG132 and extracts were subjected to immunoprecipitation using anti-Flag antibody. Co-precipitating proteins were detected with the indicated antibodies. IP, immunoprecipitation; IB, immunoblot. (b) 293 cells were transfected with Flag-Yap1 and increasing amounts of myc-Itch. Cells were then treated as in (a). (c) 293 cells were transfectes with the indicated plasmids. Cell extracts were immunoprecipitaed with endogenous anti-p73 antibody and treated as in (a)
Taking together the data presented here, we conclude that the binding of Yap1 and Itch to p73α is mutually exclusive and accumulation of Yap1 displaces the binding of Itch and vice versa.
Yap1 prevents p73 ubiqitination by the Itch E3 ligase
The finding that Yap1 displaces Itch binding to p73 suggests that Yap1 can abrogate the activity of Itch in destabilizing p73. This prediction was tested by evaluating the level of p73 in the presence and absence of Yap1 and Itch. The data revealed that Itch overexpression significantly reduced the p73 level, but this was not the case in the presence of Yap1 (Figure 6a). The WW domain of Yap1 is important for the ability of Yap1 to interact with p73.28 In order to demonstrate the necessity of the Yap1 WW domain for the displacement of Itch, we used a mutant of Yap in which the second conserved tryptophan and the conserved proline of the WW domain were each substituted to alanine.25 The Yap1 WW mutant was inefficient in preventing Itch-mediated degradation of p73 (Figure 6b). Similar results were obtained when the half-life of p73 was determined. In that case, the short half-life of p73 in the presence of Itch was significantly prolonged when Yap1 was overexpressed (Figure 6c). To confirm that Yap1 reduces Itch–p73 interaction and p73 ubiquitination, HEK293 cells were co-transfected with HA-tagged Ub (HA ubiq.), myc-tagged Itch (myc-Itch), Flag-tagged p73α and increasing amounts of Yap1 in the presence of the proteasome inhibitor MG132. Cell extracts were subjected to IP with anti-Flag antibodies and detected with anti-HA. The data revealed that p73 was ubiquitinated in an Itch-dependent manner, but the level of ubiquitination was dramatically decreased by Yap1 in a Yap1 dose-dependent manner (Figure 6d). These data are consistent with the possibility that Yap1 competes with Itch for the binding to p73 and thus prevents p73 destabilization.
Yap1 prevents p73 ubiqitination by the E3 ligase Itch. (a) 293 cells were transfected with Ha p73α, Flag-Yap1 and myc-Itch as indicated. Twenty-four hours after transfection, cell lysates were subjected to Western blot analysis with the indicated antibodies. (b) 293 cells were trasnsfected with the indicated plasmids and treated as in (a). WW*-Yap1 mutant. (c) The same as in (a) except that 24 h after transfection cells were treated with 20 μg/ml CHX for different time points as indicated. (d) 293 cells were transfected with Flag-p73α, Flag-Yap1, myc-Itch and HA-Ub. Twenty-four hours after transfection, cells were treated with MG132 and then collected. Cells extracts were subjected to immunoprecipitation using anti-HA antibody. Co-precipitating proteins were detected with the indicated antibodies
p73 accumulation by cisplatin is mediated by Yap1
It has been demonstrated that p73 is stabilized following DNA damage induced by γ-irradiation and cisplatin (reviewed by Oberst et al.8). To evaluate the contribution of Yap1 to p73 stabilization following cisplatin treatment 293 cells were co-transfected with p73 and Yap1. Twenty-four hours after transfection, cells were treated with 25 μM cisplatin for an additional 24 h before Western blot analysis. Following cisplatin treatment, p73 accumulated to the level comparable to that achieved by overexpressing Yap1 (Figure 7a, lanes 2 and 3).
Yap1 mediates p73 stabilization after cisplatin treatment. (a) 293 cells were transfected with either HA-p73α alone or together with Flag-Yap1. After 24 h, cells were treated with 25 μM cisplatin for 24 h and then were harvested for immunoblotting. (b) 293 cells were transiently transfected with HA-p73 and without or with pSUPER Yap1 encoding Yap1-specific siRNA. Cells were harvested 48 h after transfection and subjected to Western blot analysis with the indicated antibodies. (c) H1299 cells were transfected as indicated with pSUPER Yap1 encoding Yap1-specific siRNA, together with an H2b-GFP (GFP) plasmid, and were treated with 25 μm cisplatin for 24 h. The number of H2b-GFP-positive cells undergoing apoptosis (sub-G1 population) were determined by flow cytometry 48 h after transfection. (d) Immunoblot showing endogenous Yap1 protein level in H1299 stably expressing pRetroSUPER Yap1 encoding Yap1-specific siRNA. (e) The same as C except the H1299 stable Yap1 siRNA cell line was used. Bars represent SD for at least three independent experiments
To further study, the role of Yap1 in p73 stabilization in response to cisplatin treatment, endogenous Yap1 was knocked down by pSUPER Yap1 encoding Yap1-specific siRNA. Cisplatin treatment led to stabilization of p73α, however, this was not the case in Yap1 knockdown cells (Figure 7b). On the basis of this result, reduction of endogenous Yap1 would be expected to hamper p73-mediated cell death. To challenge this prediction, H1299 cells were transfected with a plasmid encoding Yap1-specific siRNA and a marker pEGFP-H2B plasmid and treated with 25 μm cisplatin. Forty-eight hours after transfection, the amount of GFP-positive cells undergoing apoptosis (sub-G1) were determined by FACS (Figure 7c). A significant reduction in the level of apoptotic cells in response to cisplatin treatment was obtained in cells expressing Yap1 siRNA. Similar results were obtained using an H1299 cell line stably expressing Yap1 siRNA that was established by infecting with pRetroSuper Yap1 (Figure 7d and e). Our results indicate that in addition to its role as co-activator, Yap1 also has an important role in stabilization of p73 to induce apoptosis in response to cisplatin treatment.
Discussion
p73 shares a significant structural and functional homology with the tumor suppressor p53, however, the mechanisms which regulate their stabilization are different.1 The major E3 ligase of p53 is Mdm2 that binds the N-terminal transcription activation domain of p53. A number of signaling pathways culminate in modification of the N-terminal domain of p53 to escape Mdm2 interaction. Interestingly, p73 binds Mdm2 but this interaction does not result in p73 modification and destabilization.33 By analogy with p53, phosphorylation of p73 at the Y99 position by c-Abl in response to DNA damage would be expected to interrupt binding of an E3 ligase to p73. In fact, c-Abl phosphorylation of p73 does lead to an increase in p73 levels.9 However, attempts to identify the putative E3 ligase that would be expected to interact with p73 at this position have so far been unsuccessful. Instead, a role for c-Abl phosphorylation of p73 on Y99 in p73 stabilization has been demonstrated to involve the p38/Pin1 pathway.16 Interestingly, the Itch E3 ligase that interacts with p73 and sensitizes p73 to proteasomal degradation binds at the C-terminal region of p73. We have shown that Yap1 competes with Itch for binding to p73 at the PPPY motif, and this prevents Itch ubiquitination, and subsequent degradation, of p73. The Yap1-mediated stabilization of p73 appears to be independent of the c-Abl phosphorylation of p73 on Y99, as a Y99F p73 mutant was stabilized by overexpression of Yap1 (data not shown). This is consistent with the findings that Yap1 association with p73 is independent of c-Abl binding and phosphorylation.28 Our data demonstrate that in addition to c-Abl-mediated p73 stabilization, Yap1-mediated p73 stabilization represents a robust element of the p73-stabilization network in response to DNA damage.
Our results further show that the stabilization of p73 that is mediated by Yap1 is important for the p73-mediated response to DNA damage. Strano, et al20, 28 have shown Yap1 increases p73 transcriptional activation of proapoptotic genes. We have demonstrated that knock-down of Yap1 by siRNA significantly reduces the amount of p73 accumulation and p73-induced apoptosis in response to DNA damage. Rossi, et al32 demonstrated that Itch overexpression reduces p73 levels and p73 transactivation of apoptotic genes, and that Itch is downregulated following DNA damage, resulting in higher TAp73 levels. Although DNA damage-induced downregulation of Itch does increase p73 stability, it is not sufficient to ensure the proapoptotic function of p73, as demonstrated by the knock-down of Yap1 in cisplatin-treated cells. Under this condition, where Itch was downregulated by the DNA damage, Yap1 was still required to ensure p73 stabilization and apoptotic activity. We conclude that Yap1 stabilizes p73 by competing with residual Itch for binding to p73, thereby preventing its ubiquitination and degradation. Taken together, these data demonstrate that Yap1 works on two fronts to increase p73-mediated apoptosis: co-activation of p73 transcription and stabilization of the p73 protein.
Yap1 stabilization of p73 may not be restricted to the ubiquitin-dependent pathway. It is possible that Yap1 may also regulate the degradation of p73α by the 20S proteasomes, which occurs in a ubiquitin-independent manner. A number of cellular proteins, including p53 and p73, have been shown to be subject to degradation of this type.30 The defining characteristic determining susceptibility appears to be the possession of an unstructured protein region, although there may be additional factors that determine susceptibility. p53 and p73 can be protected from degradation via this pathway by binding to NQO1.30 Stabilization of p53 from degradation by this pathway was required for p53 accumulation following DNA damage, and for p53-mediated apoptosis.34 It appears that, in general, binding of 20S substrate proteins to any other protein will protect them from 20S degradation, as was shown for ornithine decarboxylase.35 For this reason, it is likely that Yap1 can also protect p73α from 20S degradation, by virtue of the Yap1–p73α interaction. This possibility and its ramifications for p73-mediated apoptosis are currently under investigation.
Rossi et al.32 have shown that Itch protein level is downregulated in response to DNA damage. This would tend to shift the balance in favor of Yap1 binding to p73, which would then increase p73 stabilization. Another factor that might favor Yap1/p73 interaction at the expense of p73/Itch interaction is the relocalization of both Yap1 and p73 following DNA damage. Colocalization of both proteins would be expected to increase their interaction and the ability of Yap1 to stabilize p73. It was shown by others,18, 20 that both p73 and Yap1 colocalize in the PML-NBs. Interestingly, PML modulates p73 half-life by inhibiting its degradation in a p300 acetylation-dependent manner.18 In addition, when cells are exposed to apoptotic conditions, Yap1 recruits p300 to potentiate p73 acetylation in a PML-dependant manner.20 The interaction of Yap1 and p73 with the NBs after DNA damage may provide an additional mechanism to insure Yap1/p73 interaction in an Itch-intact nuclear compartment. The redundancy in p73 stabilization mechanisms serves as a safety net to assure p73 function under stress conditions.
As a co-activator protein, Yap1 was found to activate many transcription factors,20, 22, 23, 24 including p73.28 It is an open question whether the activation of these proteins by Yap1 recruits RNA polymerase II to interact with the general transcription apparatus36 or whether Yap1 stabilizes the transcription factors. We cannot rule out the possibility that in fact Yap1 has a dual function. It is important to note that the identification of Yap1 as a transcription co-activator is based on luciferase reporter assays that could not exclude either possibility.
The observation that a transcriptional co-activator can also affect the stability of its client transcription factor is not restricted to Yap1/p73, and may represent a more general mechanism for gene regulation. Such relationships may have either positive or negative effects on transcription and stability. In the case of Yap1/p73, Yap1 is a positive regulator of p73 transcription and stability. Similarly, the transcription co-activator CBP was found to increase both the transcriptional activity and the stability of the transcription factor complex HNF6/Foxa2.37 The opposite relationship exists with p53 and the Mdm2 protein. Mdm2 is a ubiquitin ligase, which leads to p53 degradation, and also functions to repress p53 transcriptional activity.38 It was found that the RING domain of Mdm2 mediates histone monoubiquitinylation and transcriptional repression.38 By analogy with p53, it is tempting to speculate that besides the ubiquitination function of Itch, it might also mediate transcription repression, contributing to the regulation of gene silencing. Therefore, by possessing the power to control both the transcriptional activity and the stability of transcription factors, transcription co-activators or repressors can quickly and efficiently influence transcriptional output and cellular responses.
The data presented here support the concept that under DNA damage p73 should be upregulated and activated in order to induce apoptosis, whereas its negative regulator should be suppressed. On the one hand, DNA damage downregulates Itch protein level and on the other hand, Yap1 prolongs p73 half-life at the same physiological condition supporting the important role of p73 in the induction of apoptosis. Interestingly, endogenous Yap1 protein is elevated upon cisplatin treatment (Figure 7b), however, the molecular mechanism for it is not understood yet. Elevation of Yap1 levels in response to DNA damage demonstrates another level of regulation of this pathway, which suggests that p73 activation must be tightly controlled to insure a quick and efficient activation of p73 target genes in response to stress conditions. As there are several p73 isoforms with proapoptotic and antiapoptotic functions,1, 39, 40 it remains to be determined whether Yap1 can have a modular role in regulation of the different p73 splice variants.
Materials and Methods
Cells and cell culture
The cell lines used were 293 human embryonic kidney cells, human large cell lung carcinoma H1299 p53−/− and HCT116 p53−/− human colon carcinoma cells. 293 and HCT116 were grown in Dulbecco's modified Eagle's minimal essential medium (DMEM; GIBCO) supplemented with 8% fetal bovine serum (GIBCO), 100 U/ml penicillin, 100 μg/ml streptomycin and cultured at 37°C in a humidified incubator with 5.6% CO2. H1299 cells were cultured under the same conditions with Roswell Park Memorial Institute (RPMI) medium. The H1299 cell line expressing stable pRetroSuper Yap1 siRNA was generated as described.41
Plasmids and transfection
Overexpression of p73 was achieved by transfection of pSG5-HA-p73α and pIRES-Flag-p73α. Flag CMV Yap1 was a gift from M Sudol. pEGFP-Yap1 and pCDNA3-HA-p73α Y487F were kindly provided by G Blandino. The pRK5 myc-Itch plasmid was a gift from G Melino.32 Empty pSG5 and PCGN vectors were used to keep the amount of the transfected DNA constant among samples. All transfections were carried out by the calcium phosphate method. The precipitates were left for 8 h, after which the cells were washed with PBS and medium was changed. For transfection of H1299 cells, the culture medium was changed to DMEM before transfection, and RPMI medium was added after the PBS wash. The cells were harvested 24–48 h post-transfection. To generate a Yap1 siRNA plasmid, the pSUPER vector was ligated with annealed oligonucleotides, containing a 19-nucleotide (nt) sequence derived from human Yap1 transcript (CCAGAGAATCAGTCAGAGA), a 9 nts spacer and the reverse complement of the same 19 nts sequence as described.42 Empty pSuper plasmid, or the nonspecific plasmid pSuper X,43 encoding siRNA against HBV X protein were used for control. Cells were collected 48 h after transfection and cell lysates were subjected to immunoblot analysis as described above.
Chemicals
MG132 (Calbiochem) was dissolved in DMSO at a concentration of 50 mM and used at a final concentration of 25 μM. Protease inhibitor cocktail was purchased from Sigma and used at 1 : 100. Propidium iodide purchased from Sigma and cisplatin was from ABIC Israel.
Immunoblot analysis
Cell extracts were prepared by lysis of PBS-washed cells in RIPA lysis buffer (50 mM Tris-HCl, pH 8, 150 mM NaCl, 1% Nonidet P-40 (v/v), 0.5% deoxycholate (v/v), 0.1% SDS (v/v), 1 mM dithiothreitol and Sigma protease inhibitor cocktail (P 8340) diluted 1 : 100). The insoluble pellet was discarded, and the protein concentration was determined by using Bradford reagent (Bio-Rad). Equal amounts of protein were mixed with Laemmli sample buffer (4% SDS, 20% glycerol, 10% 2-mercaptoethanol and 0.125 M Tris-HCl pH 6.8), heated at 95°C for 5 min and loaded onto a 10% polyacrylamide-SDS gel. After electrophoresis, proteins were transferred to 0.45-μm cellulose nitrate membranes (Schleicher & Schuell). Loading equivalence and transfer efficiency were monitored by Ponceau S staining, and the membranes were then incubated with appropriate antibodies to proteins of interest followed by horseradish peroxidase-conjugated anti-IgG antibodies. Signals were developed by using Super Signal (Pierce) at 20°C for 5 min, and the membranes were then exposed to X-ray film (Fuji) for an appropriate time and developed. Membranes were stripped with 50 mM citric acid before a different primary antibody was used followed by PBST (PBS+0.1% Tween) washes. The antibodies used were anti-HA, monoclonal anti-α-tubulin and anti-Flag (Sigma); monoclonal anti-GFP (Babco); polyclonal anti-human p73 BL906 (Bethyl); and polyclonal anti-Yap1 H125 (Santa Cruz Biotechnology).
Half-life determination
293 cells were transfected with the indicated plasmids. Twenty-four hours after transfection cells were treated with 20 μg/ml CHX (Sigma) for different time points. Cells were harvested, and cell extracts were immunoblotted with the indicated antibodies.
35S pulse chase: 293 and H1299 cells were transfected with HA-p73α alone or together with Flag-Yap1. For 30 min, 24 h post-transfection, cells were starved in DMEM with dialyzed serum (Biological industries) and then labeled with 80 μCi/ml of 35S-Met for 45 min. Unlabeled Met (2%) was added and cells were collected in RIPA buffer at the indicated times. Samples were subjected to IP with anti-HA antibody. Immunoprecipitates were washed six times in RIPA buffer, run on SDS-PAGE and detected by autoradiography.
Ubiquitination assays
293 cells were transiently transfected with mammalian expression plasmids for HA-Ub, together with the indicated plasmid. At 24 h after transfection, cells were treated for 4 h with 100 μM MG132 before harvesting. Cell extracts were then subjected to IP using anti-Flag agarose conjugate beads (Sigma). Ub conjugates were detected using an anti-HA antibody (Biotest).
Flow cytometry
Cells were seeded at a density of 2.5 × 106/10 cm dish. Twenty-four hours after transfection, some plates were treated with 25 μM cisplatin for an additional 24 h. Floating and attached cells were collected and combined together, washed twice with PBS and fixed in 70% ethanol. After fixation, the cells were washed with PBS and resuspended in 50 μg/ml RNase A and 25 μg/ml propidium iodide in PBS. In each assay, 100 000 cells were collected by FACScan and analyzed with the Cellquest program (BD Biosciences).
Abbreviations
- CHX:
-
cycloheximide
- DMSO:
-
dimethyl sulfoxide
- FACS:
-
fluorescence-activated cell sorting
- GFP:
-
green fluorescent protein
- HA:
-
hemagglutinin
- IB:
-
immunoblot
- IP:
-
immunoprecipitation
- NB:
-
nuclear body
- NQO1:
-
NAD(P)H quinone oxidoreductase 1
- PBS:
-
phosphate-buffered saline
- PML:
-
promyelocytic leukemia
- SH2, SH3:
-
src homology (domain) 2,3
- Ub:
-
ubiquitin
- Yap1:
-
Yes-associated protein 1
References
Irwin MS, Kaelin WG . p53 family update: p73 and p63 develop their own identities. Cell Growth Differ 2001; 12: 337–349.
Flores ER, Tsai KY, Crowley D, Sengupta S, Yang A, McKeon F et al. p63 and p73 are required for p53-dependent apoptosis in response to DNA damage. Nature 2002; 416: 560–564.
Yuan ZM, Shioya H, Ishiko T, Sun X, Gu J, Huang YY et al. p73 is regulated by tyrosine kinase c-Abl in the apoptotic response to DNA damage. Nature 1999; 399: 814–817.
Agami R, Blandino G, Oren M, Shaul Y . Interaction of c-Abl and p73alpha and their collaboration to induce apoptosis. Nature 1999; 399: 809–813.
Gong JG, Costanzo A, Yang HQ, Melino G, Kaelin Jr WG, Levrero M et al. The tyrosine kinase c-Abl regulates p73 in apoptotic response to cisplatin-induced DNA damage. Nature 1999; 399: 806–809.
Pediconi N, Ianari A, Costanzo A, Belloni L, Gallo R, Cimino L et al. Differential regulation of E2F1 apoptotic target genes in response to DNA damage. Nat Cell Biol 2003; 5: 552–558.
Stiewe T, Putzer BM . Role of the p53-homologue p73 in E2F1-induced apoptosis. Nat Genet 2000; 26: 464–469.
Oberst A, Rossi M, Salomoni P, Pandolfi PP, Oren M, Melino G et al. Regulation of the p73 protein stability and degradation. Biochem Biophys Res Commun 2005; 331: 707–712.
Tsai KK, Yuan ZM . c-Abl stabilizes p73 by a phosphorylation-augmented interaction. Cancer Res 2003; 63: 3418–3424.
Ben-Yehoyada M, Ben-Dor I, Shaul Y . c-Abl tyrosine kinase selectively regulates p73 nuclear matrix association. J Biol Chem 2003; 278: 34475–34482.
Sanchez-Prieto R, Sanchez-Arevalo VJ, Servitja JM, Gutkind JS . Regulation of p73 by c-Abl through the p38 MAP kinase pathway. Oncogene 2002; 21: 974–979.
Minty A, Dumont X, Kaghad M, Caput D . Covalent modification of p73alpha by SUMO-1. Two-hybrid screening with p73 identifies novel SUMO-1-interacting proteins and a SUMO-1 interaction motif. J Biol Chem 2000; 275: 36316–36323.
Costanzo A, Merlo P, Pediconi N, Fulco M, Sartorelli V, Cole PA et al. DNA damage-dependent acetylation of p73 dictates the selective activation of apoptotic target genes. Mol Cell 2002; 9: 175–186.
Stukenberg PT, Kirschner MW . Pin1 acts catalytically to promote a conformational change in Cdc25. Mol Cell 2001; 7: 1071–1083.
Lu KP, Hanes SD, Hunter T . A human peptidyl-prolyl isomerase essential for regulation of mitosis. Nature 1996; 380: 544–547.
Mantovani F, Piazza S, Gostissa M, Strano S, Zacchi P, Mantovani R et al. Pin1 links the activities of c-Abl and p300 in regulating p73 function. Mol Cell 2004; 14: 625–636.
Mantovani F, Gostissa M, Collavin L, Del Sal G . KeePin’ the p53 family in good shape. Cell Cycle 2004; 3: 905–911.
Bernassola F, Salomoni P, Oberst A, Di Como CJ, Pagano M, Melino G et al. Ubiquitin-dependent degradation of p73 is inhibited by PML. J Exp Med 2004; 199: 1545–1557.
Sudol M, Hunter T . NeW wrinkles for an old domain. Cell 2000; 103: 1001–1004.
Strano S, Monti O, Pediconi N, Baccarini A, Fontemaggi G, Lapi E et al. The transcriptional coactivator Yes-associated protein drives p73 gene-target specificity in response to DNA Damage. Mol Cell 2005; 18: 447–459.
Sudol M . Yes-associated protein (YAP65) is a proline-rich phosphoprotein that binds to the SH3 domain of the Yes proto-oncogene product. Oncogene 1994; 9: 2145–2152.
Espanel X, Sudol M . Yes-associated protein and p53-binding protein-2 interact through their WW and SH3 domains. J Biol Chem 2001; 276: 14514–14523.
Howell M, Borchers C, Milgram SL . Heterogeneous nuclear ribonuclear protein U associates with YAP and regulates its co-activation of Bax transcription. J Biol Chem 2004; 279: 26300–26306.
Vassilev A, Kaneko KJ, Shu H, Zhao Y, DePamphilis ML . TEAD/TEF transcription factors utilize the activation domain of YAP65, a Src/Yes-associated protein localized in the cytoplasm. Genes Dev 2001; 15: 1229–1241.
Komuro A, Nagai M, Navin NE, Sudol M . WW domain-containing protein YAP associates with ErbB-4 and acts as a co-transcriptional activator for the carboxyl-terminal fragment of ErbB-4 that translocates to the nucleus. J Biol Chem 2003; 278: 33334–33341.
Sudol M, Bork P, Einbond A, Kastury K, Druck T, Negrini M et al. Characterization of the mammalian YAP (Yes-associated protein) gene and its role in defining a novel protein module, the WW domain. J Biol Chem 1995; 270: 14733–14741.
Mohler PJ, Kreda SM, Boucher RC, Sudol M, Stutts MJ, Milgram SL . Yes-associated protein 65 localizes p62c-Yes to the apical compartment of airway epithelia by association with EBP50. J Cell Biol 1999; 147: 879–890.
Strano S, Munarriz E, Rossi M, Castagnoli L, Shaul Y, Sacchi A et al. Physical interaction with Yes-associated protein enhances p73 transcriptional activity. J Biol Chem 2001; 276: 15164–15173.
Basu S, Totty NF, Irwin MS, Sudol M, Downward J . Akt phosphorylates the Yes-associated protein, YAP, to induce interaction with 14-3-3 and attenuation of p73-mediated apoptosis. Mol Cell 2003; 11: 11–23.
Asher G, Tsvetkov P, Kahana C, Shaul Y . A mechanism of ubiquitin-independent proteasomal degradation of the tumor suppressors p53 and p73. Genes Dev 2005; 19: 316–321.
Perry WL, Hustad CM, Swing DA, O'Sullivan TN, Jenkins NA, Copeland NG . The itchy locus encodes a novel ubiquitin protein ligase that is disrupted in a18H mice. Nat Genet 1998; 18: 143–146.
Rossi M, De Laurenzi V, Munarriz E, Green DR, Liu YC, Vousden KH et al. The ubiquitin-protein ligase Itch regulates p73 stability. EMBO J 2005; 24: 836–848.
Balint E, Bates S, Vousden KH . Mdm2 binds p73 alpha without targeting degradation. Oncogene 1999; 18: 3923–3929.
Levanon D, Glusman G, Bangsow T, Ben-Asher E, Male DA, Avidan N et al. Architecture and anatomy of the genomic locus encoding the human leukemia-associated transcription factor RUNX1/AML1. Gene 2001; 262: 23–33.
Asher G, Bercovich Z, Tsvetkov P, Shaul Y, Kahana C . 20S proteasomal degradation of ornithine decarboxylase is regulated by NQO1. Mol Cell 2005; 17: 645–655.
Spiegelman BM, Heinrich R . Biological control through regulated transcriptional coactivators. Cell 2004; 119: 157–167.
Rausa III FM, Hughes DE, Costa RH . Stability of the hepatocyte nuclear factor 6 transcription factor requires acetylation by the CREB-binding protein coactivator. J Biol Chem 2004; 279: 43070–43076.
Minsky N, Oren M . The RING domain of Mdm2 mediates histone ubiquitinylation and transcriptional repression. Mol Cell 2004; 16: 631–639.
Melino G, Lu X, Gasco M, Crook T, Knight RA . Functional regulation of p73 and p63: development and cancer. Trends Biochem Sci 2003; 28: 663–670.
Melino G, De Laurenzi V, Vousden KH . p73: friend or foe in tumorigenesis. Nat Rev Cancer 2002; 2: 605–615.
Brummelkamp TR, Bernards R, Agami R . Stable suppression of tumorigenicity by virus-mediated RNA interference. Cancer Cell 2002; 2: 243–247.
Brummelkamp TR, Bernards R, Agami R . A system for stable expression of short interfering RNAs in mammalian cells. Science 2002; 296: 550–553.
Shlomai A, Shaul Y . Inhibition of hepatitis B virus expression and replication by RNA interference. Hepatology 2003; 37: 764–770.
Acknowledgements
We thank Drs Y Yarden for the HA-Ub plasmid, R Agami for the pSUPER, G Melino pRK5 Myc-Itch, G Blandino for GFP-Yap1 and pCDNA3-HA-p73α Y487F mutant, M Sudol for CMV Yap1 wt and WW* mutant plasmid. We also thank S Budilovsky for her technical assistance. This work was supported by grants from the Samuel Waxman Cancer Research Foundation, from Israel Cancer Research Fund and from the Minerva Foundation with funding from the Federal German Ministry for Education and Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by DR Green
Rights and permissions
About this article
Cite this article
Levy, D., Adamovich, Y., Reuven, N. et al. The Yes-associated protein 1 stabilizes p73 by preventing Itch-mediated ubiquitination of p73. Cell Death Differ 14, 743–751 (2007). https://doi.org/10.1038/sj.cdd.4402063
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402063
Keywords
This article is cited by
-
Imatinib inhibits SARS-CoV-2 infection by an off-target-mechanism
Scientific Reports (2022)
-
Tax1 banding protein 1 exacerbates heart failure in mice by activating ITCH-P73-BNIP3-mediated cardiomyocyte apoptosis
Acta Pharmacologica Sinica (2022)
-
P4HA2-induced prolyl hydroxylation suppresses YAP1-mediated prostate cancer cell migration, invasion, and metastasis
Oncogene (2021)
-
Riluzole-induced apoptosis in osteosarcoma is mediated through Yes-associated protein upon phosphorylation by c-Abl Kinase
Scientific Reports (2021)
-
NQO1 potentiates apoptosis evasion and upregulates XIAP via inhibiting proteasome-mediated degradation SIRT6 in hepatocellular carcinoma
Cell Communication and Signaling (2019)