Abstract
E2F6 is believed to repress E2F-responsive genes and therefore plays an important role in cell-cycle regulation. However, the role of E2F6 in the control of apoptosis remains unknown. We show here that the expression of E2F6 was downregulated with a concurrent increase in BRCA1 mRNA and cleaved protein during ultraviolet (UV)-induced apoptosis in human embryonic kidney 293 cells. Moreover, E2F6 overexpression distinctly inhibited UV-induced apoptosis as well as UV-induced increases in BRCA1 expression and cleavage, accompanied with increases of the full-length BRCA1 and BRCA1 nuclear foci. In contrast, knockdown of E2F6 by small interfering RNA had opposite effects. Furthermore, these effects of E2F6 on BRCA1 depended upon the association of E2F6 with BRCA1 via its C-terminus in a UV-triggered manner and upon the transcriptional repression by E2F6 on the BRCA1 promoter. These findings provide the first demonstration of the important role for E2F6 in the control of apoptosis via targeting of BRCA1.
Similar content being viewed by others
Main
Apoptotic cell death (programmed cell death) is a normal physiological process that plays a critical role in development, tissue homeostasis, toxic stress, and tumor suppression through active removal of damaged and/or excess cells.1 Recent studies have shown that the E2F transcription factor family has a pivotal role in the regulation of cell proliferation and antiproliferative processes, such as apoptosis and senescence.2, 3 Members of this family act by establishing the correct transcription state of the genes that direct the cell-cycle process and trigger cell death when they become deregulated by oncogenic mutations.4 Therefore, by understanding the specific roles of the individual E2Fs and the mechanisms underlying their function, we can gain important insights into the processes of cellular proliferation and apoptosis.
The human E2F family of transcription factors has eight known family members (E2F1–8)2, 5, 6, 7 and two dimeric partners (DRTF1-polypeptide-1 and DRTF1-polypeptide-2), the latter of which forms heterodimers to generate functional E2F complexes.8, 9 E2F members possess highly conserved DNA binding and heterodimerization domains, with the exception of E2F7 and E2F8.6, 7 The E2F family can be functionally categorized into either transcriptional ‘activators’ (E2F1–2, E2F3a) or ‘repressors’ (E2F3b, E2F4–8).2, 10 Activators E2F1, E2F2, and E2F3a are negatively regulated by interaction with pRb, a tumor suppressor protein. The Rb-E2F pathway regulates expression of many genes whose products are required for DNA synthesis, progression of the cell cycle and apoptosis.11 E2F4 and E2F5 are thought to be important in the repression of E2F-responsive genes for the cell cycle.2, 10 E2F7 and E2F8, the most recently identified ones, can function as repressors, independent of DP interaction, to negatively influence cellular proliferation.6, 7
E2F6 lacks the transactivation and pocket protein (pRb, p107, and p130) binding domains found in other E2Fs, and can repress the transcription of known E2F-responsive genes.5 It has also been shown that its carboxyl terminus has repression activity.12 Overexpression of E2F6 leads to accumulation of cells in the S-phase and delays re-entry into the cell cycle in quiescent cells.13 More recently, E2F6 has been suggested to repress transcription of BRCA1, CTIP, ART27, HP-1a, and RBAP48 genes (which encode functions involved in tumor suppression and maintenance of chromatin structure) via its repression of the target promoter.14 However, the biological function of E2F6 is not fully understood.
The tumor suppressor BRCA1 is a multifunctional protein implicated in DNA repair, cell-cycle regulation, and transcription.15 BRCA1 becomes hyperphosphorylated in response to various DNA-damaging agents, including γ-irradiation.16 It is implicated in the repair of double-strand breaks by homologous recombination in embryonic stem cells from BRCA1 knockout mice17 and in global genomic repair of ultraviolet (UV)-induced DNA damage.18 In addition, BRCA1 has been implicated in the regulation of apoptosis. Exogenous expression of BRCA1 sensitizes cells to apoptosis.19 BRCA1 also enhances apoptosis induced by stimuli such as growth factor withdrawal and substratum detachment in ovarian and prostate cancer cell lines.20 Although it is well-established that BRCA1 plays a crucial role in mediating apoptosis, little is known about a possible link between E2F6 and BRCA1 during apoptosis.
In this study, we tested the hypothesis that E2F6 may repress apoptotic response via targeting of BRCA1. Dose- and time-dependent relationships between expression of E2F6 and BRCA1 during UV-induced apoptosis were examined. Further, the functional link between them was identified by overexpression of full-length E2F6 and RNA interference (RNAi)-mediated knockdown of E2F6 expression, and by co-transfection of E2F6 and the C-terminus of BRCA1 during UV-induced apoptosis in human embryonic kidney 293 (HEK293) cells, combined with a protein–protein interaction approach. We demonstrate that the human E2F6 gene plays an important role in UV-induced apoptosis. We also provide evidence for the regulatory mechanisms of the functional link between E2F6 and apoptosis via its modulation of BRCA1.
Results
Dose- and time-dependent changes of E2F6 and BRCA1 levels during UV-induced apoptosis
To investigate the temporal effect of genotoxic agent-induced apoptosis on the transcript and protein levels of E2F6 and BRCA1, HEK293 cells were irradiated with various doses of UV ranging from 25 to 100 J/m2 or were observed for up to 24 h after irradiation at 100 J/m2. Induction of apoptosis was confirmed by the appearance of condensed chromatin or apoptotic bodies 24 h after irradiation in a dose- and a post-time-dependent manner (Figure 1a). In the same sets of experiments, the transcript and protein levels of endogenous E2F6 were reduced significantly in a UV dose- and a post-time-dependent manner, accompanied by UV dose- and post-time-dependent upregulation of BRCA1 transcript at higher doses (Figures 1b and c). The protein level of full-length BRCA1 decreased in dose- and post-time-dependent manners, whereas the cleaved 90 kDa C-terminal fragments of BRCA121 accumulated greatly with the increased UV doses and time points following UV irradiation. Therefore, the total BRCA1 protein levels increased after UV irradiation (Figure 1c). Taken together, these data imply that downregulation of E2F6 may be involved in UV-induced apoptosis via deregulation of BRCA1.
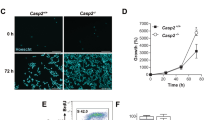
Dose- and time-dependent changes of E2F6 and BRCA1 mRNA and protein levels during UV-induced apoptosis in HEK293 cells. (a) Nuclear morphology (upper panel) and percentage (lower panel) of apoptotic cells showing condensed chromatin or apoptotic bodies at indicated UV doses and time points post-UV irradiation. Nuclei were stained with Hoechst 33342. (b) RT-PCR analyses of transcripts for E2F6, BRCA1, and housekeeping gene M28S during UV-induced apoptosis. (c) Protein levels of E2F6, full-length BRCA1, cleaved BRCA1, and actin were analyzed by Western blotting in whole-cell extracts with or without UV irradiation. Results are mean±S.E.M. from at least three individual experiments; >200 cells were scored per experiment in (a)
E2F6 represses UV-induced apoptosis
To confirm the role of E2F6 in the regulation of apoptosis, we examined UV-induced apoptosis after overexpression of wild-type E2F6 or knockdown of E2F6 by using small interfering RNA (siRNA). HEK293 cells were transfected with constructs that express control enhanced green fluorescent protein (EGFP), full-length wild-type E2F6-EGFP (E2F6wt), mutant E2F6 with C-terminal deletion (220–281)-EGFP (E2F6ΔC), or were transfected with nonspecific siRNA (NS) or E2F6-specific siRNA (siE2F6), respectively, (Figures 2a and b). Western blot analysis for HA showed that cells transfected with E2F6wt and E2F6ΔC expressed the proper proteins (Figure 2a). Treatment of siE2F6, but not of a nonspecific control, resulted in a significant reduction in E2F6 mRNA and protein expression (Figure 2b). Fewer E2F6wt-expressing cells demonstrated apoptotic morphology and nuclear changes than did control cells at 24 h after UV irradiation (100 J/m2), whereas more cells transfected with siE2F6 showed apoptotic changes than did cells transfected with NS (Figures 2c and d). Consistently, overexpression of E2F6wt significantly reduced the percentage of cells in the sub-G1 phase analyzed by flow cytometry (FACS), a condition corresponding to apoptotic cells (Figure 2e), and showed less DNA fragmentation (Figure 2f). In contrast, cells treated with siE2F6 were more numerous in the sub-G1 phase and had more DNA fragmentation (Figures 2e and f). These data demonstrate a functional role for E2F6 in the regulation of the apoptotic response to UV irradiation, that is, that upregulation of E2F6 expression can increase the resistance of cells to UV-induced apoptosis and that downregulation of endogenous E2F6 can sensitize cells to UV-induced apoptosis.
The effects of E2F6wt and siE2F6 on UV (100 J/m2, 24 h post)-induced apoptosis in HEK293 cells. (a) Schematic representation of control (EGFP), wild-type E2F6 (E2F6wt), and mutant E2F6 with repression domain deletion (E2F6ΔC, 220–281, left panel); expression of E2F6wt and E2F6ΔC detected by Western blotting using an anti-HA antibody 24 h post-transfection (right panel). (b) Expression of E2F6 mRNA and protein in HEK293 cells transfected with NS or siE2F6 by RT-PCR and Western blot analysis, respectively. M28S or actin was used as the control. (c) Cell morphology and percentage of apoptotic cells (showing round and distorted morphology) with or without UV irradiation after transfection of EGFP or E2F6wt (left panel), or NS or siE2F6 (right panel). (d) Nuclear morphology and percentage of apoptotic cells treated as (c). Nuclei were stained with Hoechst 33342 and apoptotic cells showed condensed chromatin or apoptotic bodies. (e) Flow cytometric analysis of propidium iodide-stained cells treated as (c). (f) DNA fragmentation. DNA of ∼4 × 106 cells was loaded, and DNA fragmentation analyzed by 2% agarose gel and visualized with ethidium bromide under UV light. Data are represented as mean±S.E.M.; n=4 each, and >200 cells were scored per experiment in (c) and (d). **P<0.01 compared to the UV-treated group expressing control EGFP or to the UV-treated group transfected with NS
E2F6 abrogates the increases of BRCA1 transcription and accumulation of cleaved BRCA1 C-terminus during UV-induced apoptosis
BRCA1 is cleaved by caspase-3 during UV irradiation, and the cleaved C-terminal 90 kDa fragment triggers the apoptotic response through activation of the BRCA1 downstream effectors, GADD45, and JNK.21 To determine whether E2F6 represses UV-induced apoptosis through modulation of BRCA1, we then examined the effect of overexpression or specific knockdown of E2F6 on BRCA1 transcription and accumulation of the cleaved BRCA1 C-terminus. HEK293 cells were transiently transfected with control EGFP, E2F6wt, NS or siE2F6, followed by exposure to various doses of UV irradiation. Cells expressing EGFP did not show significant changes in the transcript levels of BRCA1 compared to that of the control group without transfection (data not shown), whereas cells expressing E2F6wt had significantly lower transcript levels of BRCA1 compared to that of EGFP-transfected cells at doses of 25–100 J/m2 UV irradiation (Figure 3a). In contrast, the transcript of BRCA1 in cells treated with siE2F6 was significantly increased in a UV dose-dependent manner (Figure 3a). Consistently, EGFP expression did not affect the accumulation of cleaved BRCA1 fragments, whereas the accumulation was significantly repressed by expression of E2F6wt at ⩾50 J/m2 UV irradiation, and increased by siE2F6 treatment at 25–100 J/m2 UV irradiation (Figure 3b). In parallel, after exposure to different doses of UV, there were fewer apoptotic cells when expressing E2F6wt and more when treated with siE2F6 (Figure 3c). These results suggest that E2F6 may increase cell resistance to UV-induced apoptosis via its inhibition on the BRCA1 transcript and accumulation of cleaved BRCA1 C-terminus.
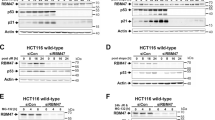
UV-induced increases of BRCA1 transcripts, accumulation of cleaved BRCA1 C-terminus, and apoptosis are suppressed by E2F6 overexpression, but enhanced by RNAi-mediated E2F6 downregulation in HEK293 cells. (a) RT-PCR analysis for BRCA1 and housekeeping gene M28S transcripts in cells with or without transfection of EGFP or E2F6wt (left panel), or NS or siE2F6 (right panel) exposed to increased doses of UV. Upper panel, representative gel images; lower panel, averaged data. (b) Western blot analysis for cleaved BRCA1 C-terminus and actin in cells treated as (a). (c) Percentage of apoptotic cells determined by nuclear morphology in parallel cells treated as (a). Nuclei were stained with Hoechst 33342. Results are mean±S.E.M.; n=4, and >200 cells were scored per experiment in (c). *P<0.05 and **P<0.01 compared to corresponding control values
It has been reported that E2F6 negatively regulates BRCA1 at the mRNA level via transcriptional repression.14 To confirm whether the increase of BRCA1 mRNA level by UV irradiation is due to the decreased repression of E2F6 on BRCA1 promoter, we performed the chromatin immunoprecipitation assays (ChIP) to examine the recruitment of E2F6 to the BRCA1 promoter. As shown in Figure 4, the BRCA1 promoter was robustly enriched by the E2F6 antibody without UV irradiation, whereas the binding of E2F6 to BRCA1 promoter was significantly decreased after UV irradiation and decreased further by siE2F6 treatment. However, more recruitment of E2F6 to BRCA1 promoter was detected in cells overexpressing E2F6wt. Taken together, these data indicate that E2F6 can bind to BRCA1 promoter, but the recruitment of E2F6 to the BRCA1 promoter is mitigated by UV irradiation.
E2F6 recruits to the BRCA1 promoter and this recruitment is decreased by UV irradiation and further by siE2F6 but increased by E2F6 overexpression. HEK293 cells transfected with or without EGFP, E2F6wt, NS or siE2F6 were subjected to ChIP analysis, using antibodies of E2F6 and normal rabbit IgG, 24 h post-UV irradiation (100 J/m2). A total of 0.2% of the total input of each sample was subjected to PCR to show that equal amounts of starting input chromatin were used
E2F6 represses UV-induced increases of the transcripts, cleavage of BRCA1, and apoptosis in a transfection dose-dependent manner
To provide more pertinent evidence for the possible regulation of E2F6 in UV-induced apoptosis via targeting of BRCA1, we then examined the effect of E2F6wt or siE2F6 at various doses of transfection on the transcript of BRCA1 and the accumulation of cleaved BRCA1 C-terminus during 50 J/m2 UV-induced apoptosis. With the increased amounts of transfected E2F6wt, BRCA1 transcription was correspondingly repressed during UV-induced apoptosis, whereas it was increased with increased amounts of siE2F6 transfection (Figure 5a). Similarly, the accumulation of the cleaved BRCA1 C-terminus was regulated by the transfection amount of E2F6wt or siE2F6 in a dose-dependent manner as seen above (Figure 5b). In parallel experiments, UV-induced apoptosis was repressed with an increased transfection amount of E2F6wt, but the apoptosis was increased along with increased amounts of transfected siE2F6 (Figure 5c). Taken together, these results provide further evidence that E2F6 inhibits UV-induced apoptosis by repressing the transcription of BRCA1 and the accumulation of the cleaved BRCA1 C-terminus.
UV-induced increases of the transcripts and cleavage of BRCA1, and apoptosis are suppressed by E2F6 overexpression, but enhanced by RNAi-mediated E2F6 downregulation in a transfection dose-dependent manner. (a) RT-PCR analysis for the transcripts of BRCA1 and housekeeping gene M28S in cells with the transfection of different doses of E2F6wt or siE2F6 24 h post-UV irradiation. The transfection amount was equal to that of control vectors. (b) Western blot analysis for cleaved BRCA1 and actin in cells treated as above. (c) Percentage of apoptotic cells determined by nuclear morphology in parallel cells treated as (a). Nuclei were stained with Hoechst 33342. Results are mean±S.E.M. from four individual experiments. >200 cells were scored per experiment in (c)
Co-expression of BRCA1 C-terminus (BRCA1-KE) with E2F6 partially abrogates antiapoptotic effects of E2F6
To further evaluate the contribution of decreased accumulation of cleaved BRCA1 C-terminus to the protection of cells from apoptosis by E2F6, cells were co-transfected with E2F6wt and BRCA1-KE and subjected to UV irradiation. The apoptosis seen in the cells co-expressing control EGFP and pcDNA4 was inhibited by co-overexpression of E2F6wt and pcDNA4, but was enhanced by co-transfection of BRCA1-KE and EGFP. Moreover, the apoptosis seen in the cells co-expressing BRCA1-KE and EGFP was inhibited in the cells co-expressing E2F6wt and BRCA1-KE (Figure 6a). Similar observations were obtained from the analysis of cells in sub-G1 phase using FACS (Figure 6b). These results confirmed that overexpressed BRCA1 C-terminus sensitizes cells to UV-induced apoptosis; therefore, the reduction of the cleaved BRCA1 C-terminus is involved in the antiapoptotic effects of E2F6. Furthermore, these results also suggest that E2F6 may inhibit apoptosis by other mechanisms in addition to its repression of the cleavage of BRCA1.
E2F6 inhibits apoptosis by reducing the accumulation of cleaved BRCA1 C-terminus. HEK293 cells were co-transfected with control EGFP and control pcDNA4, E2F6wt and pcDNA4, BRCA1 C-terminal fragment KE (BRCA1-KE) and EGFP, or E2F6wt and BRCA1-KE. After 24 h, the cells were exposed to UV irradiation and observed at 24 h post-UV. (a) Nuclear morphology of cells (left panel) and percentage of apoptotic cells (right panel). (b) Flow cytometric analysis of propidium iodide-stained cells. Results are mean±S.E.M. from four individual experiments. A total of >400 cells were scored per experiment in (a)
E2F6 increases UV-induced BRCA1 nuclear foci
Endogenous BRCA1 accumulates into discrete subnuclear foci termed ‘BRCA1 nuclear dots’ in the S-phase.22 Following UV-induced DNA damage, these dots become re-distributed into more dispersed nuclear foci and are involved in DNA repair.23, 24 To confirm whether antiapoptotic effects of E2F6 are related to the increase of the BRCA1 nuclear foci, we next examined the BRCA1 nuclear foci in cells transfected with E2F6wt or siE2F6 during UV-induced apoptosis. Without UV irradiation, only 25% of control cells showed <10 foci, whereas after UV irradiation, ∼41–48% of cells, with or without transfection of control EGFP or NS, displayed >50 foci (Figure 7a and b). The BRCA1 foci were increased to 71% in cells expressing E2F6wt, but decreased to 27% in cells treated with siE2F6 (Figure 7a and b). These results indicate that the increases of UV-induced BRCA1 nuclear foci may also contribute to the antiapoptotic effect of E2F6.
E2F6 overexpression increases nuclear BRCA1 foci, whereas siRNAs targeted against E2F6 decrease BRCA1 foci. (a) Representative confocal images showing nuclear foci of BRCA1 in cells with or without EGFP, E2F6wt, NS, or siE2F6 after UV irradiation. The images were merged by BRCA1 nuclear foci stained with anti-BRCA1 (D-20) and nuclei stained with Hoechst33342. (b) Quantification of foci. Values shown are the percentage of cells containing foci from one representative experiment. Similar results were obtained in at least three other experiments
Association of E2F6 with BRCA1 via its C-terminus is required for its repression of cleavage of BRCA1 and increase of BRCA1 nuclear foci
As demonstrated above, E2F6 was able to repress the cleavage of BRCA1 and increase BRCA1 nuclear foci during UV-induced apoptosis. To determine whether these effects of E2F6 are related to its association with BRCA1, we expressed E2F6wt or mutant E2F6ΔC in HEK293 cells, after which the cells were exposed to 100 J/m2 UV irradiation and allowed to recover for 24 h. Cell lysates were immunoprecipitated with control (normal rabbit IgG and normal goat IgG), anti-E2F6, or anti-BRCA1 antibodies. The resultant complexes were recovered and resolved by SDS-PAGE, and were immunoblotted with anti-E2F6 or anti-BRCA1 antibodies. With E2F6wt expression, BRCA1 could be detected in the complexes with anti-E2F6, and E2F6 was also observed in BRCA1 immunoprecipitates after UV irradiation (Figure 8a). However, without UV irradiation, neither BRCA1 in E2F6 immunoprecipitates nor E2F6 in BRCA1 complexes could be detected (Figure 8b). Next, we tested whether the C-terminal fragment of E2F6 is a necessary domain for the association of E2F6 and BRCA1. Because of undetected content of endogenous E2F6 in immunocomplexes with 100 J/m2 UV irradiation (Figure 8a), we could exclude the effect of endogenous E2F6 on E2F6ΔC precipitation. HEK293 cells expressing E2F6ΔC were treated as above. After UV irradiation, the association between E2F6ΔC and BRCA1 could not be detected (Figure 8a). Thus, we conclude that E2F6 and BRCA1 can associate with one another via the C-terminus of E2F6 in a UV-triggered manner.
Association of E2F6 with BRCA1 is necessary for its modulation of BRCA1. Lysates were immunoprecipitated with normal rabbit IgG, normal goat IgG, anti-E2F6, or anti-BRCA1 (D-20) followed by immunoblotting with indicated antibodies. Original lysates were used for positive immunoblotting analysis. (a) Association of E2F6 and BRCA1 in HEK293 cells expressing control EGFP, E2F6wt, or E2F6ΔC after UV irradiation. (b) Association of E2F6 and BRCA1 in cells expressing E2F6wt without UV irradiation. (c) Cleavage of BRCA1 in the cells without (control) or with transfection of EGFP, E2F6wt, or E2F6ΔC after UV irradiation. (d) Nuclear BRCA1 foci in cells treated as (a). Values shown in (d) are from one representative experiment. Similar results were obtained in at least three other experiments
To further determine whether the association between them is necessary for the modulation of E2F6 on BRCA1, we then compared the effects of E2F6wt overexpression with mutant E2F6ΔC (the latter could not interact with BRCA1 as shown in Figure 8a) on the cleavage of BRCA1 and BRCA1 nuclear foci. E2F6wt significantly increased the protein level of full-length BRCA1 and reduced the accumulation of cleaved BRCA1 C-terminus after UV-irradiation; but the level of full-length BRCA1 was significantly decreased in E2F6ΔC cells, and was associated with an increased cleavage of BRCA1 (Figure 8c). Moreover, the increased BRCA1 foci in cells expressing E2F6wt was reduced to a control level in cells expressing E2F6ΔC (Figure 8d). These results provide evidence that the regulation of E2F6 on BRCA1, and thereby on apoptosis, is related to the association between the two proteins.
Discussion
In this study, we have demonstrated first that E2F6 can negatively regulate UV-induced apoptotic cell death. The E2F family of transcription factors has been shown to play critical roles in controlling normal and tumor cell proliferation.2 A number of lines of evidence show that E2F1 can induce both proliferation and apoptosis.11 Recent studies demonstrate that expression of the E2F6 repressor is influenced at the transcriptional level by E2F1.25 Although these observations imply that E2F6 may also be implicated in the regulation of apoptosis, no study so far has directly shown functional relevance of E2F6 in apoptosis. Here, we show that UV-induced apoptotic cell death is associated with downregulated endogenous E2F6 in a dose- and time-dependent manner. E2F6 overexpression significantly represses UV-induced cell apoptosis, whereas RNAi-mediated E2F6 downregulation sensitizes cells to UV-induced apoptosis. These findings also suggest that the ability of E2F6 to sensitize cells to a death response to genotoxic stress by its downregulation is important in tumor suppression and in radiotherapy.
Another novel finding in the present study is the establishment of BRCA1 as a target for E2F6-regulated apoptotic cell death. In addition to cell-cycle arrest, BRCA1 is reported to be implicated in the regulation of apoptosis.26 Overexpression of wild-type BRCA1 increases susceptibility to apoptosis induced by apoptotic stimuli, including ionizing radiation and treatment with anticancer agents in breast and ovarian cancer cell lines.20, 21 Here, we show that the negative regulation of E2F6 in the development of apoptosis may be related to its repression of the transcription of BRCA1 that is upregulated by UV irradiation. This is possibly mediated by the transcriptional repression by E2F6 of the BRCA1 promoter.14 The BRCA1 mRNA upregulated by high doses of UV appears to be related to the reduced transcriptional repression of E2F6 as the recruitment of E2F6 to the BRCA1 promoter is mitigated by UV irradiation. The antiapoptotic role of E2F6 is also related to the preservation of full-length BRCA1. The full-length BRCA1 is downregulated during UV-induced apoptosis, accompanied by an accumulation of the cleaved BRCA1 C-terminus,21, 27 consistent with our observations. Expression of a non-cleavable mutant BRCA1 raises the cellular resistance to UV-induced cell death, whereas caspase-3-mediated cleavage of BRCA1 triggers the apoptotic response through activation of BRCA1 downstream effectors GADD5 and JNK.21 We demonstrate that E2F6wt overexpression significantly increases full-length BRCA1 and decreases the cleavage of BRCA1, whereas E2F6 siRNA or E2F6ΔC reverses these effects. Moreover, the antiapoptotic effect of E2F6 is inhibited by co-overexpression of BRCA1 C-terminus, further confirming that the antiapoptotic effect of E2F6 is related to its reduction of the accumulation of cleaved BRCA1 C-terminus. Importantly, we find that E2F6 can associate with BRCA1 after UV irradiation and that this association disappears after deletion of the C-terminal repression domain of E2F6. However, the detailed molecular mechanisms for the association between these two proteins still require elucidation. These consistent observations indicate that E2F6 may block the cleavage of BRCA1 via the association with full-length BRCA1 through its C-terminal domain (220–281). Therefore, downregulation of E2F6 may allow increases in BRCA1 transcripts and the accumulation of the cleaved BRCA1 C-terminus, which may consequently promote the apoptotic response to genotoxic stress.
The finding that E2F6 increases BRCA1 foci after UV-induced DNA damage reveals an important role of E2F6 in the process of DNA repair. Substantial evidence shows that BRCA1 plays an important role in mediating cell survival in response to DNA damage,28 in particular as regards double-strand DNA breaks17 and global genomic repair of UV-induced cyclobutane pyrimidine dimmers.29 The known functions of BRCA1 occur mostly in the nucleus.24 The molecule becomes localized as discrete nuclear dots during cellular DNA replication (S-phase) and, after DNA damage, nuclear BRCA1 becomes re-distributed into more numerous, smaller foci that co-stain with many proteins associated with DNA repair and replication, for example, Rad51,22 Rad50/hMre11/p95,30 and proliferating cell nuclear antigen.23 With transient overexpression of E2F6, we demonstrated that E2F6 increases UV-induced BRCA1 nuclear foci and that this effect is related to their association through the C-terminal domain of E2F6. Because full-length, but not cleaved, BRCA1 has an ability to form nuclear foci dots that are associated with DNA repair,31 it may be that the increased BRCA1 nuclear foci, when wild-type E2F6 is overexpressed, may not be due to the effect of E2F6 on nuclear foci formation, but rather to the higher level of the full-length BRCA1. Therefore, E2F6 may also be involved in DNA repair, at least, via its regulation of BRCA1.
Previous studies showed that BRCA1 mRNA level is downregulated by UV irradiation at doses of ⩽20 J/m2 in human breast cancer cells, MCF-7 cells, and SK-OV-3 cells.32 This downregulation at low doses of UV appears not to be directly related to the induction of apoptotic DNA fragmentation or cell death, because the prominent downregulation of BRCA1 is observed at a UV dose of 5 J/m2 for which cell survival was ⩾90%.32 Our data collected at low doses (⩽25 J/m2) of UV (<25 J/m2 data not shown) are consistent with the above observation. At higher doses of UV irradiation, following the decrease of E2F6 mRNA and protein levels, both mRNA and cleaved fragments of BRCA1 are upregulated with appearance of an increased percentage of apoptotic cells. This regulation may allow cells to cope with the increased DNA damage by inducing apoptotic cell death via activation of BRCA1 downstream effectors19, 20 and repair of DNA damage to maintain genomic integrity.28
In conclusion, we demonstrate here that E2F6 negatively regulates UV-induced apoptosis via modulation of BRCA1. At least two mechanisms are involved in this process: (i) E2F6 represses the expression of BRCA1 at the transcriptional level; (ii) E2F6 binds to full-length BRCA1 protein via the C-terminus of E2F6 and mitigates the accumulation of BRCA1 cleavage (Figure 9). The potential impact of E2F6 upon cancer mutation via its regulation on BRCA1 remains to be examined.
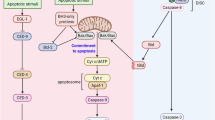
Model for the negative regulation of E2F6 on UV-induced apoptosis via modulation of BRCA1. E2F6 inhibits UV-induced apoptosis by repressing the expression of BRCA1 mRNA at the transcriptional level and cleavage of BRCA1. The effect is related to the association of E2F6 with the full-length BRCA1 via the C-terminal repression domain of E2F6 after UV irradiation, which accumulates full-length BRCA protein by mitigating the BRCA1 cleavage and may thereby increase nuclear BRCA1 foci
Materials and Methods
Cell culture
A human embryonic kidney cell line HEK293 was cultivated in Dulbecco's modified Eagle's medium supplemented with 10% fetal calf serum, 2 mM glutamine, 100 U/ml penicillin, and 100 μg/ml streptomycin at 37°C in 5% CO2.33 All cultivation media and other substances for cell culture were purchased from Gibco BRL
Plasmid construction and transfection
The plasmids, pcDNA3-HA-E2F6 (E2F6 cDNA constructs) and pcDNA3-HA-E2F6ΔC (Δ220–281, lacking the repression domain of E2F6), were provided by Professor David M Livingston (Harvard Medical School, Boston).12 The BRCA1 C-terminal fragments KE in pcDNA4 (BRCA1-KE) were obtained from Dr. QM Zhan.21 To generate the EGFP-containing E2F6wt and E2F6ΔC expression vectors, an EcoR I-Xab I CMV-EGFP-containing fragment from pAdtrack-CMV (Qbiogene) was subcloned into pcDNA3-HA-E2F6 and pcDNA3-HA-E2F6ΔC, respectively. At 24 h before transfection, cells were seeded onto six-well culture plates and transfected at a 70–80 % confluence with 1 μg or 0.25–4 μg of plasmid DNA in those experiments using the Ca-phosphate procedure, as described previously.34 Transfection efficiency was evaluated on the basis of the percentage of green fluorescent protein-positive cells.
siRNA transfection
At 24 h before transfection, HEK293 cells were plated onto six-well culture plates and transfected with siRNAs at 70–80% confluence. Double-stranded siRNAs were purchased from GenePharma and included siE2F6 (AAGGAUUGUGCUCAGCAGCUG-custom order), or scrambled siE2F6 (GGUUGUGUGUACCAACCGGAA-custom order) as nonspecific control siRNA (NS). Transfections were performed with RNAiFect reagent (Qiagen) according to manufacturers’ instructions. RT-PCR and Western analysis were performed as described below to verify the efficiency of the siRNA.
Cell irradiation
HEK293 cells or transfected HEK293 cells were irradiated with 25–100 J/m2 UVC (254 nm) using a UV lamp HL-2000 Hybrilinker (UVP, California, USA) at 25°C, as described previously.34 The culture medium was removed before irradiation and fresh medium added to the plates immediately after irradiation. Cells were then incubated at 37°C for 24 h before further experiments or for 1–24 h for the time course study.
RT-PCR analysis
Total cell RNA was extracted from monolayer cells using TriZol reagent (Invitrogen), according to the manufacturer's instructions. RNA (0.5 μg) was converted to cDNA by using Superscript II reverse transcriptase (Invitrogen) in a final volume of 20 μl, and 0.4 μl of this was used for each polymerase chain reaction (PCR). Semiquantitative PCR was performed using Taq DNA polymerase (Promega) in a Mastercycler gradient (Eppendorf, Hamburg, Germany) under the following conditions: 5 min at 95°C followed by 35 cycles. Each cycle consisted of denaturation at 95°C for 45 s, annealing at 55°C for 40 s, and extension at 72°C for 45 s. After completion of the last cycle, there was an autoextension for 5 min at 72°C. PCR products were visualized on a 1% agarose gel containing ethidium bromide. Gel images were captured with a Gel Doc 2000 system (Bio-Rad, California, USA) and signals were quantified using Quantity One software (Bio-Rad) and normalized against M28S signal. The primers of BRCA1 (forward: 5′-CAGAACAAAGCACATCAG-3′, reverse: 5′-AATCCAGATTCCAGGTAA-3′), E2F6 (forward: 5′-AGATGCTTTGGATGAGTTA-3′, reverse: 5′-GTTGGTGCTCCTTATGTG-3′), and house keeping gene M28S (forward: 5′-AGCAGCCGACTTAGAACTGG-3′, reverse: 5′-TAGGGACAGTGGGAATCTCG-3′).
DNA fragmentation assay
DNA fragmentation was determined according to previous methods,35 with slight modifications. In brief, floating and attached cells (3–4 × 106) were pelleted by centrifugation at 500 × g for 10 min and lysed in 500 μl TTE buffer (containing 10 mM Tris-HCl, pH 7.5, 1 mM EDTA, and 0.2% Triton X-100). Lysates were centrifuged at 13 000 × g for 15 min and the supernatants containing low molecular weight fragmented DNA were collected. RNA was removed by addition of RNase A (0.25 μg/μl) and incubated at 37°C for 1 h. The DNA was de-proteinized by one extraction in phenol : chloroform : isoamylalcohol (25 : 24 : 1) and two extractions in chloroform/isoamylalcohol (24 : 1), followed by precipitation at −20°C in 50% isopropanol and 130 mM NaCl. DNA was visualized on a 2% agarose gel containing ethidium bromide.
FACS analysis
Samples were prepared for FACS as described previously.36 Briefly, 1 × 106 treated cells were fixed in cold 70% ethanol for 30 min, treated with 100 μg/ml DNase-free RNase A (Sigma), and labeled with 50 μg/ml propidium iodine (Sigma). Cells were then analyzed by FACS (FACStar Plus Flow Cytometer, Becton-Dickson, San Jose, USA).
Immunoprecipitation assay
Cells were lysed in lysis buffer containing 25 mM Tris (pH 7.6), 150 mM NaCl, 1 mM NaF, 1% Triton X-100, 1 mM PMSF, 1 mM DTT (Sigma), 1 mM sodium orthovanadate, and 1 μg/ml proteinase inhibitor mixture (Sigma), as described previously.21 Immunoprecipitations were performed exactly as described,37 using 1 μg of goat anti-E2F6, rabbit anti-BRCA1 (D-20) (Santa Cruz), normal rabbit IgG, or normal goat IgG antibodies (Santa Cruz), respectively, with end-over-end mixing at 4°C for 6 h. Protein G-agarose beads (20 μl, Santa Cruz) were then added and the reaction mixtures mixed further at 4°C for 1 h. Immunoprecipitates were separated from the supernatant by centrifugation and washed with PBS containing 0.05% Nonidet P-40. Proteins eluted from the agarose beads were boiled in 1 × SDS gel-loading buffer and resolved on 8% SDS-PAGE.
Western blot analysis
Cells were lysed as described previously.23 Cell lysates or immunoprecipitates were separated on 8% (for BRCA1) or 12% (for E2F6) SDS-PAGE, and transferred to polyvinylidene difluoride membranes (MILLIPORE). Membranes were blocked with 5% non-fat milk in Tris-buffered saline-Tween 20 for 1 h at room temperature and then incubated overnight at 4°C with antibodies: rabbit anti-BRCA1 (C-20, 1 : 400, Santa Cruz), goat anti-E2F6 (1 : 1000, Santa Cruz), or mouse anti-actin (1 : 1000, Sigma). The membranes were then incubated with horseradish peroxidase-linked secondary anti-rabbit (1 : 4000, Santa Cruz), anti-goat (1 : 2000, Santa Cruz), or anti-mouse (1 : 8000, Sigma). The immunoreaction was visualized with a chemiluminescent detection kit (Western LightingTM), exposed to X-ray film and quantified with a video documentation system (Gel Doc 2000, Bio-Rad, California, USA).
ChIP
The ChIP assay was performed as described previously38 on HEK293 cells with the following exceptions. The chromatin was sheared to an average size of 500–2000 bp. After cross-linking reversal and proteinase K digestion, each individual IP was purified to isolate DNA using a QIA quick PCR purification kit (Qiagen) according to the manufacturer's instructions. The ChIP-enriched DNA was amplified by PCR with specific primers for BRCA1 promoter (forward, 5′-TGGCCAATCCAGAGCCCCGAGAG-3′ and reverse, 5′-TGTCCCCCGTCCAGGAAGTCTCA-3′). E2F6 antibody used in the ChIP assays was purchased from Santa Cruz.
Immunofluorescence staining
Immunostaining was carried out as described previously.39 Cells were seeded on glass coverslips 2 days before UV irradiation. After 24 h post-irradiation incubation, cells were fixed in 3.7% formaldehyde in PBS for 15 min, washed with PBS, permeabilized with 0.2% Triton X-100 in PBS for 10 min, and again washed with PBS. Following a blocking step with 10% goat serum in PBS for 45 min, cells were incubated with a primary antibody, rabbit anti-BRCA1 (D-20) (1 : 100, Santa Cruz) for 1 h at 37°C. After washing with PBS, cells were incubated with Texas-Red-conjugated anti-rabbit secondary antibody (1 : 1000, Sigma) at 37°C for 1 h and then were rinsed three times with PBS. To visualize the nucleus, cells were then incubated with Hoechst 33342 (Sigma) at room temperature for 15 min. The subcellular localization of BRCA1 protein was observed with a confocal scanning laser microscope (Zeiss, LSM510, Carl Zeiss, Oberkoken, Germany).
Scoring of BRCA1 intranuclear foci
Cells were prepared for immunostaining of BRCA1 as described above. Cells with or without UV exposure (100 J/m2) were cultured for 24 h before fixation. Cells were then assessed by confocal scanning laser microscopy and scored as containing either no nuclear foci, <10, or >50 BRCA1 nuclear foci. At least 90 transfected cells were scored in each well.
Statistical analysis
All data are expressed as mean±S.E.M. Data were analyzed using one-way analysis of variance and Fisher's protected least significant difference test for multiple comparisons (SPSS 10.0, SPSS Inc, USA). P<0.05 was considered to be statistically significant.
Abbreviations
- UV:
-
ultraviolet
- EGFP:
-
enhanced green fluorescent protein
- FACS:
-
flow cytometry
- HEK293:
-
human embryonic kidney 293
- siRNA:
-
small interfering RNA
References
Hengartner MO . The biochemistry of apoptosis. Nature 2000; 407: 770–776.
Trimarchi JM, Lees JA . Sibling rivalry in the E2F family. Nat Rev Mol Cell Biol 2002; 3: 11–20.
Kowalik TF, DeGregori J, Schwarz JK, Nevins JR . E2F1 overexpression in quiescent fibroblasts leads to induction of cellular DNA synthesis and apoptosis. J Virol 1995; 69: 2491–2500.
Harbour JW, Dean DC . Rb function in cell-cycle regulation and apoptosis. Nat Cell Biol 2000; 2: E65–E67.
Trimarchi JM, Fairchild B, Verona R, Moberg K, Andon N, Lees JA . E2F-6, a member of the E2F family that can behave as a transcriptional repressor. Proc Natl Acad Sci USA 1998; 95: 2850–2855.
de Bruin A, Maiti B, Jakoi L, Timmers C, Buerki R, Leone G . Identification and characterization of E2F7, a novel mammalian E2F family member capable of blocking cellular proliferation. J Biol Chem 2003; 278: 42041–42049.
Maiti B, Li J, de Bruin A, Gordon F, Timmers C, Opavsky R et al. Cloning and characterization of mouse E2F8, a novel mammalian E2F family member capable of blocking cellular proliferation. J Biol Chem 2005; 280: 18211–18220.
Helin K, Wu CL, Fattaey AR, Lees JA, Dynlacht BD, Ngwu C et al. Heterodimerization of the transcription factors E2F-1 and DP-1 leads to cooperative trans-activation. Genes Dev 1993; 7: 1850–1861.
Zhang Y, Chellappan SP . Cloning and characterization of human DP2, a novel dimerization partner of E2F. Oncogene 1995; 10: 2085–2093.
Takahashi Y, Rayman JB, Dynlacht BD . Analysis of promoter binding by the E2F and pRB families in vivo: distinct E2F proteins mediate activation and repression. Genes Dev 2000; 14: 804–816.
DeGregori J, Leone G, Miron A, Jakoi L, Nevins JR . Distinct roles for E2F proteins in cell growth control and apoptosis. Proc Natl Acad Sci USA 1997; 94: 7245–7250.
Gaubatz S, Wood JG, Livingston DM . Unusual proliferation arrest and transcriptional control properties of a newly discovered E2F family member, E2F-6. Proc Natl Acad Sci USA 1998; 95: 9190–9195.
Ogawa H, Ishiguro K, Gaubatz S, Livingston DM, Nakatani Y . A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 2002; 296: 1132–1136.
Oberley MJ, Inman DR, Farnham PJ . E2F6 negatively regulates BRCA1 in human cancer cells without methylation of histone H3 on lysine 9. J Biol Chem 2003; 278: 42466–42476.
Chen Y, Farmer AA, Chen CF, Jones DC, Chen PL, Lee WH . BRCA1 is a 220-kDa nuclear phosphoprotein that is expressed and phosphorylated in a cell cycle-dependent manner. Cancer Res 1996; 56: 3168–3172.
Cortez D, Wang Y, Qin J, Elledge SJ . Requirement of ATM-dependent phosphorylation of brca1 in the DNA damage response to double-strand breaks. Science 1999; 286: 1162–1166.
Moynahan ME, Chiu JW, Koller BH, Jasin M . Brca1 controls homology-directed DNA repair. Mol Cell 1999; 4: 511–518.
Hartman AR, Ford JM . BRCA1 induces DNA damage recognition factors and enhances nucleotide excision repair. Nat Genet 2002; 32: 180–184.
Shao N, Chai YL, Shyam E, Reddy P, Rao VN . Induction of apoptosis by the tumor suppressor protein BRCA1. Oncogene 1996; 13: 1–7.
Thangaraju M, Kaufmann SH, Couch FJ . BRCA1 facilitates stress-induced apoptosis in breast and ovarian cancer cell lines. J Biol Chem 2000; 275: 33487–33496.
Zhan Q, Jin S, Ng B, Plisket J, Shangary S, Rathi A et al. Caspase-3 mediated cleavage of BRCA1 during UV-induced apoptosis. Oncogene 2002; 21: 5335–5345.
Scully R, Chen J, Plug A, Xiao Y, Weaver D, Feunteun J et al. Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 1997; 88: 265–275.
Scully R, Chen J, Ochs RL, Keegan K, Hoekstra M, Feunteun J et al. Dynamic changes of BRCA1 subnuclear location and phosphorylation state are initiated by DNA damage. Cell 1997; 90: 425–435.
Scully R, Livingston DM . In search of the tumour-suppressor functions of BRCA1 and BRCA2. Nature 2000; 408: 429–432.
Lyons TE, Salih M, Tuana BS . The Activating E2Fs Mediate the Transcriptional Regulation of the Human E2F6 Repressor. Am J Physiol Cell Physiol 2006; 290: c189–c199.
Harkin DP, Bean JM, Miklos D, Song YH, Truong VB, Englert C et al. Induction of GADD45 and JNK/SAPK-dependent apoptosis following inducible expression of BRCA1. Cell 1999; 97: 575–586.
Fan S, Twu NF, Wang JA, Yuan RQ, Andres J, Goldberg ID et al. Down-regulation of BRCA1 and BRCA2 in human ovarian cancer cells exposed to adriamycin and ultraviolet radiation. Int J Cancer 1998; 77: 600–609.
Gowen LC, Avrutskaya AV, Latour AM, Koller BH, Leadon SA . BRCA1 required for transcription-coupled repair of oxidative DNA damage. Science 1998; 281: 1009–1012.
Hartman AR, Ford JM . BRCA1 and p53: compensatory roles in DNA repair. J Mol Med 2003; 81: 700–707.
Zhong Q, Chen CF, Li S, Chen Y, Wang CC, Xiao J et al. Association of BRCA1 with the hRad50-hMre11-p95 complex and the DNA damage response. Science 1999; 285: 747–750.
Au WW, Henderson BR . The BRCA1 RING and BRCT domains cooperate in targeting BRCA1 to ionizing radiation-induced nuclear foci. J Biol Chem 2005; 280: 6993–7001.
Andres JL, Fan S, Turkel GJ, Wang JA, Twu NF, Yuan RQ et al. Regulation of BRCA1 and BRCA2 expression in human breast cancer cells by DNA-damaging agents. Oncogene 1998; 16: 2229–2241.
Fan Y, Wu D, Jin L, Yin Z . Human glutamylcysteine synthetase protects HEK293 cells against UV-induced cell death through inhibition of c-Jun NH2-terminal kinase. Cell Biol Int 2005; 29: 695–702.
Wu S, Loke HN, Rehemtulla A . Ultraviolet radiation-induced apoptosis is mediated by Daxx. Neoplasia 2002; 4: 486–492.
Wyllie AH, Morris RG . Hormone-induced cell death. Purification ad properties of thymocytes undergoing apoptosis after glucocorticoid treatment. Am J Pathol 1982; 109: 78–87.
Zhang FB, Li L, Fang B, Zhu DL, Yang HT, Gao PJ . Passage-restricted differentiation potential of mesenchymal stem cells into cardiomyocyte-like cells. Biochem Biophys Res Commun 2005; 336: 784–792.
Wu X, Yang Z, Liu Y, Zou Y . Preferential localization of hyperphosphorylated replication protein A to double strand break repair and checkpoint complexes upon DNA damage. Biochem J 2005; 391: 473–480.
Oberley MJ, Tsao J, Yau P, Farnham PJ . High-throughput screening of chromatin immunoprecipitates using CpG-island microarrays. Methods Enzymol 2004; 376: 315–334.
Rodriguez JA, Henderson BR . Identification of a functional nuclear export sequence in BRCA1. J Biol Chem 2000; 275: 38589–38596.
Acknowledgements
This study was supported by Grants from Major and General Programs of the National Natural Sciences Foundation of China (30393133, 30370536) and Science & Technology Committee of Shanghai Municipality (02JC14038). We are grateful to Professor David M Livingston for providing us plasmids pCDNA3-HA-E2F6 and pCDNA3-HA-E2F6ΔC and to Professor Xin-Yuan Liu for offering us the plasmid pAdtrack-CMV. We also thank Zhi-Hu Qu for the assistance of Western blot analysis. We also thank Dr. Bao-Xue Ge for the constructive discussion and Dr. Sheri M Skinner for revising the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by B Dynlacht
Rights and permissions
About this article
Cite this article
Yang, WW., Wang, ZH., Zhu, Y. et al. E2F6 negatively regulates ultraviolet-induced apoptosis via modulation of BRCA1. Cell Death Differ 14, 807–817 (2007). https://doi.org/10.1038/sj.cdd.4402062
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402062
Keywords
This article is cited by
-
ClE2F1 Overexpression Enhances Plant Growth in Chrysanthemum lavandulifolium (Fisch. ex Trautv.) Makino
Plant Molecular Biology Reporter (2018)
-
Class I HDAC inhibitor mocetinostat induces apoptosis by activation of miR-31 expression and suppression of E2F6
Cell Death Discovery (2016)
-
Polycomb protein EZH2 suppresses apoptosis by silencing the proapoptotic miR-31
Cell Death & Disease (2014)
-
Type 3 inositol 1,4,5-trisphosphate receptor negatively regulates apoptosis during mouse embryonic stem cell differentiation
Cell Death & Differentiation (2010)
-
Downregulation of miR-205 and miR-31 confers resistance to chemotherapy-induced apoptosis in prostate cancer cells
Cell Death & Disease (2010)