Abstract
Myofibroblasts are critical cellular elements of wound healing generated at sites of injury by transdifferentiation of resident cells. A paradigm for this process is conversion of hepatic stellate cells (HSC) into hepatic myofibroblasts. Treatment of HSC with DNA methylation inhibitor 5-aza-2′-deoxycytidine (5-azadC) blocked transdifferentiation. 5-azadC also prevented loss of IκBα and PPARγ expression that occurs during transdifferentiation to allow acquisition of proinflammatory and profibrogenic characteristics. ChIP analysis revealed IκBα promoter is associated with transcriptionally repressed chromatin that converts to an active state with 5-azadC treatment. The methyl-CpG-binding protein MeCP2 which promotes repressed chromatin structure is selectively detected in myofibroblasts of diseased liver. siRNA knockdown of MeCP2 elevated IκBα promoter activity, mRNA and protein expression in myofibroblasts. MeCP2 interacts with IκBα promoter via a methyl-CpG-dependent mechanism and recruitment into a CBF1 corepression complex. We conclude that MeCP2 and DNA methylation exert epigenetic control over hepatic wound healing and fibrogenesis
Similar content being viewed by others
Main
Myofibroblasts are key cellular elements of wound healing responses, responsible for wound contraction, recruitment of inflammatory cells and remodelling of the extracellular matrix (ECM) to promote formation of a temporary scar. Severe or chronic injury is associated with persistence and proliferation of wound healing myofibroblasts and formation of crosslinked scars also known as fibrotic tissue.1 Fibrosis is a disease process common to major organs such as the kidney, liver, pancreas, heart and lungs in the context of reiterative injury or infection.2, 3, 4, 5 Recent evidence also suggests that the stroma reaction, in which tumours benefit from the antiapoptotic and growth promoting properties of their surrounding ECM, is driven by myofibroblast secretion of collagen-rich ECM into the extracellular space surrounding the growing tumour.6 There is therefore interest in determining the molecular events that regulate the production and behaviour of myofibroblasts during injury, infection and tumour formation.
Cell transdifferentiation is a generic process by which myofibroblasts are generated in injured tissues such as the liver, kidney, pancreas and lung.3, 4, 5, 6, 7 Cell transdifferentiation enables fully differentiated nonstem cells to dramatically alter their phenotype and function in a stable and long-term manner.8 In the context of wound repair, transdifferentiation enables rapid responses to damage and injury by providing a mechanism to generate wound healing myofibroblasts from resident cells located in the vicinity of the damaged tissue. Differentiated cells that undergo myofibroblast transdifferentiation (MTD) in response to tissue injury include hepatic stellate cells (HSC),7 kidney mesangial cells and epithelia,2 pancreatic stellate cells,4 vocal fold stellate cells9 and lung fibroblasts.3 It is therefore likely that MTD is a generic process in wound healing responses of the majority of soft tissues. Understanding the molecular regulation of MTD should therefore provide therapeutic solutions to ineffective and aberrant wound healing responses. In addition, the emerging role of the myofibroblast in tumour development and growth suggests that MTD may be a therapeutic target in cancer. However, despite the identification of numerous intracellular signalling molecules and transcription factors implicated as regulators of MTD,7, 8 the critical regulatory events that orchestrate the transformation of a committed differentiated cell into a myofibroblast remain poorly defined.
Transition of HSC to the hepatic myofibroblast (HM) represents a well characterised and widely accepted paradigm for MTD in the injured liver.7, 10 HSC-derived HM are transitionary components of the hepatic wound healing response, however, their persistence and proliferation in chronic liver disease promotes the progressive remodelling of the hepatic ECM from normal to fibrotic and eventually cirrhotic tissue.10, 11 The HSC exists as a quiescent retinoid-storing element of the liver that has a perisinusoidal location in the Space of Disse. The MTD or so-called ‘activation’ of HSC generates a highly proliferative, contractile, smooth muscle-alpha actin (α-SMA) positive cell that acquires a plethora of biochemical and functional phenotypic characteristics not observed with the quiescent HSC.7, 12, 13 Transition of HSC to the myofibroblastic phenotype is most readily studied by a widely accepted and reproducible in vitro model in which freshly isolated rodent or human HSC are cultured on plastic in the presence of serum-containing medium.7, 10, 11, 12, 13 This cell culture model recapitulates events occurring in the injured liver and enables the earliest events of MTD and the functional characteristics of the HSC-derived myofibroblast to be analysed in fine detail.7, 13 HSC-derived myofibroblasts are highly profibrogenic and secrete vast quantities of collagen I and III as well as the tissue-inhibitor of metalloproteinases-1 (TIMP-1) which combine to promote collagen deposition.10 In addition, the HSC-derived myofibroblast is proinflammatory producing an array of cytokines (e.g. IL-6), chemokines (e.g. MCP-1), mitogens (e.g. PDGF, TGFβ and leptin) and adhesion molecules (e.g. ICAM1).14, 15, 16, 17, 18 These dramatic functional changes are associated with a global reprogramming of the HSC transcriptome, including the up- and downregulation of several hundred different genes.12 Similar dramatic changes in phenotype are described for MTD occurring in the kidney, pancreas and lung indicating that a common transcriptional reprogramming process may operate to generate myofibroblasts in response to tissue injury.2, 3, 4 In support of this concept there are many reports describing alterations in expression and/or activity of key transcription factors such as PPARs, NF-κB and AP-1 common to MTD of distinct organ-specific myofibroblast-precursor cells.19, 20, 21, 22 However, no single transcription factor or combinations of transcription factors are so far described as responsible for exerting global control of MTD.
Improved understanding of the relationship between chromatin structure, DNA methylation and gene transcription provides new epigenetic paradigms with which to explain how the differentiated state of a mammalian cell can be regulated at a global level.23, 24 For example, methyl-CpG-binding proteins such as MeCP2 have the potential to exert regulation over the expression of multiple genes via their interaction with methylated DNA and their association with histone modifying enzymes inclusive of histone deacetylases (HDACs) and histone methyltransferase activity.24 In the present study, we demonstrate a role for DNA methylation and MeCP2 as orchestrators of MTD. Treatment of isolated HSC with the DNA methylation inhibitor 5-aza-2′-deoxycytidine (5-azadC) blocks MTD including prevention of the loss of expression of genes associated with the HSC phenotype such as PPARγ and IκBα. We also demonstrate that MeCP2 is selectively induced in HM of the injured liver, during culture-induced MTD of HSC, and provide evidence for a novel epigenetic mechanism by which MeCP2 promotes transcriptional repression of the IκBα gene. These data provide direct experimental evidence for epigenetic control of MTD and a physiolgical role for MeCP2 in wound healing and fibrogenesis.
Results
MeCP2 is induced in wound healing myofibroblasts
The first suggestion that the MTD may be subject to epigenetic regulation was provided by the discovery that the methyl-CpG-binding protein MeCP2 is selectively induced in α-SMA positive myofibroblasts in the diseased liver (Figure 1). Normal rat liver lacked MeCP2 expression. However, livers subjected to chronic hepatotoxic injury to induce fibrosis displayed strong MeCP2 staining associated with cells found within fibrotic tissue. By contrast parenchymal and endothelial cells remained MeCP2 negative. To help identify the MeCP2 positive cells we performed dual staining with anti-α-SMA and anti-MeCP2. Double positive cells were localised to fibrotic tissue indicative of MeCP2 being expressed by wound healing smooth muscle myofibroblasts. Of note, smooth muscle cells surrounding vessels staining strong positive for α-SMA were uniformly negative for MeCP2 expression. MeCP2 is therefore induced selectively in wound healing myofibroblasts of the experimentally injured liver. Similarly, we demonstrated MeCP2 expression in diseased human liver (primary biliary cirrhosis) associated predominantly with myofibroblastic cells populating fibrotic tissue (Figure 1b). HSC can be isolated and purified from normal rodent liver (described in Materials and Methods) and upon culture on plastic in serum-containing media spontaneously transdifferentiate into a myofibroblastic cell in a manner that closely resembles MTD events occurring in the injured liver.7, 12, 13 By employing this well-established in vitro model of MTD, we showed that MeCP2 mRNA expression was increased during transdifferentiation of HSC (Figure 2a). Moreover, Western blot analysis of MeCP2 revealed that the protein was not expressed at detectable levels in quiescent HSC but was powerfully induced during culture-induced MTD of these cells (Figure 2a). The dramatic induction of MeCP2 protein expression compared with the relatively modest increase in MeCP2 transcript levels suggests that the major regulatory check point for controlling MeCP2 expression in HSC is likely to be at the level of translation of MeCP2 transcript to protein. MeCP2 mRNA and protein species were also expressed at high levels in human myofibroblasts generated by culture-induced MTD of isolated human HSC (Figure 2b). These data indicate a role for MeCP2 in the pathobiology of liver fibrosis and suggest that epigenetic mechanisms may regulate the phenotype and function of the myofibroblast.
MeCP2 immunostaining on liver sections. (a) MeCP2 immunostaining on sections of normal (olive oil sham injured) rat liver or livers from rats at week 5 of chronic injury with CCl4 to induce fibrosis. Top × 20 magnification images show absence of MeCP2 in normal liver (left-hand micrograph) and selective MeCP2 expression located in fibrotic lesions of diseased liver (brown staining, in right-hand micrograph with examples noted by red arrows). Central × 20 images show α-SMA staining (brown) localised selectively to smooth muscle cells lining vessels of normal liver (left) and to myofibroblasts in association with fibrotic lesions in diseased liver (right). High magnification (× 100) images show dual positive MeCP2 (black) and α-SMA (brown) in myofibroblasts associated with fibrotic lesions of diseased liver (examples noted by red arrows in micrograph on the right) which contrasts with the absence of MeCP2 staining in vascular smooth muscle cells lining hepatic vessels which stain single positive for α-SMA (brown staining cells in micrograph on the left). Photomicrographs are representative of four control or CCl4 injured rat livers. (b) MeCP2 immunostaining on normal human liver and primary biliary cirrhosis (PBC) liver sections. Original magnifications: × 20
MeCP2 expression in rat and human HSC. (a) Left panel-total RNA isolated from day 0, 3, 7 and 10 cultures of rat HSC (rHSC) was used to obtain first strand cDNA, which was then utilised as a template in RT-PCRs using protocols described under Materials and Methods. cDNA species encoding MeCP2 and β-actin were amplified over 28 and 25 PCR cycles, respectively. Right panel – whole-cell extracts were isolated from quiescent rHSCs (day 0) or culture transdifferentiated rHSC on day 7. Of total cell extract, 30 μg was used to immunoblot for MeCP2 and β-actin control. Gel was representative of three independent experiments. (b) Left panel – total RNA isolated from two independent preparations of human HSC-derived myofibroblasts (1+2) were used to obtain first strand cDNA, which was then used as a template in RT-PCRs using protocols described under Materials and Methods. cDNA species encoding MeCP2 and was amplified over 28 PCR cycles. Right panel – whole-cell extracts were isolated from sister cell cultures and 30 μg were used immunoblot for MeCP2
5-azadC treatment blocks MTD of HSC
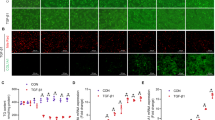
Epigenetic control of gene expression by MeCP2 is mediated at least in part through its association with hypermethylated CpG-rich islands found in gene promoters and enhancers. CpG methylation can be inhibited by the nucleotide analog 5-azadC. Treatment of freshly isolated rat HSC with 5-azadC had a profound inhibitory effect on culture-induced MTD (Figure 3). Seven days culture of HSC under control conditions was associated with the classic morphological transdifferentiation from a rounded weakly adherent quiescent cell into a highly adherent cell that spread out onto the plastic substrate (Figure 3a). HSC maintained in the presence of 5-azadC remained in their rounded phase-bright state indicating a blockade of morphological transdifferentiation. Of note we failed to observe significant signs of morphological change as long as the cells were maintained in 5-azadC-containing medium (up to 6 weeks). However, cells remained viable and upon removal of the drug would, after a lag period of 7–10 days, subsequently undergo the typical morphological changes associated with the MTD (Figure 3a). Hence, 5-azadC effectively stalls morphological features of the MTD. 5-azadC also inhibited proliferation of cultured HSC, a behavioural phenotype characteristically associated with MTD (Figure 3b). Induction of procollagen 1(I) and TIMP-1 gene expression are classic events associated with MTD, both were repressed in 5-azadC-treated cultures (Figure 3c). 5-azadC treatment also blocked the diminution in expression of PPARγ and IκBα that accompanies MTD, both genes are believed to maintain the quiescent phenotype of HSC (Figure 3d). Glial fibrillary acidic protein (GFAP), which is expressed at equivalent levels in quiescent and transdifferentiated HSC, was unaffected by 5-azadC (Figure 3e). Hence, the effects of 5-azadC are restricted to genes that undergo changes in expression characteristic of MTD.
Treatment of quiescent rat HSC with 5-azadC prevents their activation in vitro. (a) Freshly isolated rHSC were plated out onto plastic in several separate dishes. Photomicrographs were taken of the quiescent HSC within 30 min of culture, while the remaining dishes were cultured in the presence or absence (control) of 1 μM 5-azadC for 7 days before image capture. Following the 7-day period, 5AzdC was removed and both control and previously treated cells were incubated in fresh media for further 2 weeks. All photomicrographs are shown at original × 20 magnification. (b) 1 × 105 quiescent rHSCs were seeded in triplicate on day 0 and incubated in 16% DMEM or same media supplemented with 1 μM 5-azadC for further 6 days. Proliferation was measured on day 7 by adding 0.5 μCi of [3H]thymidine per well for 16 h. Incorporated [3H]thymidine was measured as outlined in Materials and Methods and is expressed as counts per minute. All experiments shown represent n=3, where each separate experiment was carried out 15 times. Use of paired t-test revealed difference between control activated and 5-azadC treated rHSC to be highly significant with P<0.0001 as denoted by three stars. White bars represent control cells, whereas 5-azadC treated cells are shown as a black bar. (c) TaqMan analysis of procollagen I and TIMP1 mRNA was measured in 20 ng of cDNA from quiescent, 5-azadC treated (for 7 days from isolation) or activated (day 7) rHSCs. The relative level of transcriptional difference was calculated and expressed as an average±S.E.M. from three independent cell preparations. Average cycle numbers for 18sRNA were 17.5, 17.2 and 17.8 for quiescent, activated and 5-azadC treated rHSC, respectively. Average cycle numbers for TIMP1 and collagen1 are as shown below the graphs in Figure 3C. Use of paired t-test revealed difference between control activated and 5-azadC treated rHSC to be significant with P=0.0024 for TIMP1 (represented by two stars) and P=0.029 for collagen1 (represented by one star). (d and e) Whole-cell extracts were isolated from quiescent (Q) rHSCs (day 0), 5-azadC treated (AzadC) rHSC (for 7 days from day 0) or culture transdifferentiated/activated (A) rHSC on day 7. Of total cell extract, 30 μg was used in immunoblots for IκBα, PPARγ and GFAP. All gels were representative of at least two independent experiments
Epigenetic regulation of IκBα gene transcription
Diminution of IκBα expression during MTD is a requirement to reprogramme basal NF-κB activity, enable the myofibroblast to constitutively express proinflammatory molecules and avoid apoptosis.7, 11 5-azadC treatment of HSC-derived myofibroblasts stimulated elevated expression of IκBα mRNA and protein (Figure 4a) supporting the idea that an epigenetic mechanism operates to repress IκBα expression in myofibroblasts. Crosslinked ChIP (XChIP), which is capable of detecting indirect as well as direct protein: DNA interactions confirmed that MeCP2 interacts with the IκBα promoter and this interaction was lost in cells treated with 5-azadC (Figure 4b). However, native ChIP (NChIP), which favours the detection of stable and direct protein: DNA interactions, failed to detect MeCP2 interaction indicating that MeCP2 associates with the promoter in a relatively unstable manner or alternatively requires protein–protein interactions retained in the XChIP assay (data not shown). To determine a role for MeCP2 as a regulator of IκBα expression, the LX-2 human HSC-derived myofibroblast cell line,25 was transfected at high efficiency with an siRNA vector designed to inhibited MeCP2 expression (Figure 4c and d). Cells transfected with MeCP2 siRNA expressed higher levels of IκBα protein relative to cells transfected with a control siRNA (Figure 4d). In addition, MeCP2 siRNA stimulated a dose-dependent increase in IκBα promoter-luciferase reporter activity (Figure 4e). As MeCP2 recruits HDAC and HMT activities that promote transcriptionally repressed chromatin it was of interest to determine chromatin structure at the IκBα promoter in myofibroblasts. This was achieved by XChIP, which firstly confirmed that the IκBα promoter is located in the context of transcriptionally permissive euchromatin (enrichment for acetylated histone 4). However, enrichment for dimethyl-lysine 9 of histone 3 and lack of acetylated histone 3, acetylated K14 and dimethylated K4 of histone H3 (Figure 5) shows that the promoter is associated with transcriptionally repressed chromatin. 5-azadC-treatment promoted remodelling of these histone signatures resulting in loss of the repressive dimethyl-lysine 9, histone 3 modification and enrichment of the transcriptionally active acetylated histone 3 modification as well as the presence of acetylated K14 and dimethylated K4 of histone H3 (Figure 5). Similar results were obtained with native ChIP (data not shown). Furthermore, we could not detect association of tri-methylated K9 or K4 of histone H3 in either control or 5-azadC-treated cells suggesting that IκBα is neither found in fully active nor in fully repressed state (data not shown).
Transcriptional repressor MeCP2 binds to IκBα promoter in vivo; siRNA directed against MeCP2 is able to alleviate transcriptional repression. (a) Transdifferentiated rHSC were cultured in the presence of 1 μM 5-azadC for up to 48 h. Total RNA or whole-cell protein extracts were made from the cells at time points 24 and 48 h as well as the control untreated cells. Left panel – total RNA was used to obtain first strand cDNA, which was then used as a template in RT-PCRs using protocols described under Materials and Methods. cDNA encoding IκBα and β-actin was amplified over 28 and 27 PCR cycles, respectively. Of total cell protein extract, 50 μg was used to immunoblot for IκBα and β-actin (right panel). Both gels were representative of three independent experiments. (b) HSC (control and 5-azadC treated) were fixed and chromatin sheared by sonication. In total input sample the crosslinks between the DNA and proteins were reversed following the sonication, whereas in other samples 100 μg of chromatin was incubated with irrelevant or anti-MeCP2 antibody in an immunoprecipitation reaction and subsequently treated as outlined under Materials and Methods. Genomic DNA associated with precipitated MeCP2 was eluted off in 50 μl of TE buffer. Of DNA (contained in a volume of 3μl), 15 ng was used in PCRs with primers specific for rat IκBα promoter. The amplification was carried out over 35 cycles using PCR conditions as outlined under Materials and Methods. PCR products were separated by electrophoresis on a 1% agarose gel. Gel shown was representative of two independent experiments. (c) 5 × 105 LX2 cells were seeded onto 10 cm dishes and transfected with 1 μg psiRNA-MeCP2 which includes an expression cassette for green fluorescent protein (GFP) to enable visualization of transfected cells. At 48 h following the transfection, the cells were photographed at × 20 magnification either using light microscopy or a FITC filter to allow visualization of GFP. (d) LX2 cells transfected with psiRNA-MeCP2 or control psiRNA-Lux (as described in 4C) were harvested 48 h after transfection and whole-cell extracts made. Of total cell extract from each sample, 30 μg was used in immunoblots for MeCP2, IκBα and β-actin loading control. Gels shown were representative of two independent experiments. (e) LX2 cells were transfected with 100 ng of pRLTK, 0.5 μg IκBα promoter/luciferase reporter and 0.5 μg or 1 μg psiRNA-MeCP2 for 48 h. Luciferase activity was determined, normalized to pRLTK activity, and expressed as mean±S.E. of three independent triplicate transfections
ChIP on crosslinked chromatin reveals ability of 5-azadC to modify histones associated with IκBα promoter in vivo. Crosslinked chromatin was prepared from control (denoted by a minus sign) or 48 h 5-AzdC treated (denoted by a plus sign) LX2 cells as described in Materials and methods. Of chromatin, 100 μg was incubated with 10 μg of either antidimethyl K4 of histone H3, antidimethyl K9 of histone H3, antiacetyl K14 of histone H3, antiacetyl H4 or antiacetyl H3 antibodies. The protein/DNA complexes were immunoprecipitated using sepharose/protein A+G beads (Santa Cruz Biotechnology, Inc.). DNA component of the immunoprecipitated complexes was separated from protein fraction by reversing crosslinks in 0.3 M NaCl at 65°C for 4 h followed by phenol/chloroform extraction and DNA isolation using PCR clean up columns. Obtained DNA was used as template in PCR reactions containing human IκBα promoter-specific primers
MeCP2 repression of IκBα transcription requires interaction with CBF1
MeCP2 classically regulates gene transcription via direct binding to methyl-CpG dinucleotides.23 However, despite the presence of an evolutionary conserved CpG-rich island spanning the promoter and transcription start site of the IκBα gene we failed to detect methylated CpGs in the promoter using methylation-sensitive restriction enzyme/polymerase chain reaction (PCR) mapping and bisulphite DNA sequencing (Figure 6). By contrast, methyl-CpG were detected in the exon 1/intron 1 regions of the IκBα gene, but of note, methylation status of this region of the gene was not altered during MTD. These observations coupled with our inability to detect direct interaction of MeCP2 with the IκBα promoter in the NChIP assay indicated that MeCP2 may be recruited to the promoter from a distal methylated site and via protein–protein interactions involving factors able to directly interact with the promoter. We previously described how CBF1 (RBP-Jκ) is induced with MTD of HSC (confirmed in Figure 7a) and represses IκBα transcription via direct sequence-specific binding to an overlapping CBF1/NF-κB site at the distal end of the IκBα promoter.26 However, the components of the CBF1 corepression complex assembling at the IκBα promoter were not characterised. We now show that MeCP2 interacts with CBF1 in myofibroblasts indicating that MeCP2 may be recruited to the IκBα promoter via interaction with CBF1 (Figure 7b). CBF1 can be converted from a transcriptional repressor to activator by the intracellular domain of Notch (NotchIC) through its direct interaction with CBF1 and removal of the corepressor complex.27 We therefore reasoned that overexpression of NotchIC should prevent the repressive effect of MeCP2 at the IκBα promoter. MeCP2 repression of IκBα transcription was blocked in the presence of NotchIC but was unaffected by mutant NotchIC proteins (CDN1 and CDNIT) lacking the RAM domain required for interaction with CBF128 (Figure 7c). From these data we propose that MeCP2 requires interaction with CBF1 in order to mediate repression of IκBα gene transcription and that MeCP2 is a component of the CBF1 corepression complex.
Methylation sensitive enzyme digests reveal presence of a methylated CpG island within the exonI/intronI of IκBα promoter in vivo. (a) Genomic DNA obtained from day 1, 3, 7 and 14 rHSC cultures was either fully digested with AciI (CpG methylation sensitive restriction enzyme) or left uncut. The cut or uncut gDNA was then used as template in PCR reaction with primer sets 1 (PS1) and 2 (PS2). PS1 primers were designed to amplify a region of the IκBα gene inclusive of the promoter, exon 1 and intron 1. PS2 primers were designed to amplify exon 1 and intron 1 independently of the promoter (see c). (b) Rat IκBα distal promoter and proximal gene genomic sequence were found to form a CpG island using a CpG island searcher (http://www.cpgislands.com/). Start of exon I was taken as +1 of the sequence, where around −120 to +800 of the gene form the CpG island denoted by thick blue line with individual CpG dinucleotides denoted by thin vertical lines. (c) Diagram of rat IκBα promoter/gene with illustrated binding sites for the primer sets 1 and 2. As determined using a combination of methylation sensitive restriction enzyme digest and bisulphite sequencing, the green arrows show position of nonmethylated CpGs, whereas the red arrows indicate methylated CpGs. *Above the arrows highlight CpGs confirmed as either methylated or nonmethylated using bisulphite sequencing. Position of AciI restriction enzyme cut sites are shown as yellow bars
MeCP2 associates with a transcriptional repressor CBF1 and is subject to control through NotchI signalling. (a) Whole-cell protein extracts were prepared from quiescent rHSCs (day 0) or transdifferentiated rHSC (day 7 of culture). Of protein extract, 30 μg was used in immunoblots for transcriptional repressor CBF1 and β-actin. Gel shown was representative of three independent experiments. (b) CBF1-Flag tagged expression construct was transfected into LX2 cells. The cells were harvested 48 h later and agarose/anti-FLAG beads were used to immunoprecipitate CBF1 as well as the proteins associated with CBF1. Immunoprecipitated fractions were separated on SDS-PAGE, transferred onto nitrocellulose membrane and blotted for CBF1 and MeCP2. (c) Seven-day cultured rHSC were transfected with 100 ng of pRLTK, 1 μg of wt pIκBα promoter/lux reporter and 2 μg of pBos (control vector) or pNotch1IC expression vector or vector encoding Notch1IC mutants (pBosCDN1 or pBosCDN1T) for 48 h. The cells were harvested, luciferase activity determined and normalized to pRLTK activity, and expressed as mean±S.E. of four independent triplicate transfections. Use of paired t-test revealed difference between control pBOS and pNotchIC transfected samples to be significant with P=0.0102 which is represented by one star
Discussion
Myofibroblasts can be both beneficial and harmful cellular elements of the wound healing response. On the one hand, the strong contractive forces and abundant secretion of collagen by the myofibroblast is critical for wound closure. On the other hand, the persistence, proliferation and migration of the myofibroblast in chronically injured tissues leads to fibrosis and provides a favourable environment for tumour cell growth and invasion.1, 2, 3, 4, 5, 6 Controlled transdifferentiation offers the prospect of the optimisation of wound healing by regulating the conversion of resident differentiated cells into myofibroblasts. In the present study, we provide evidence that global reprogramming of gene expression during the conversion of an HSC to a myofibroblast is subject to epigenetic control.
Perhaps our most convincing evidence for the epigenetic control of MTD was demonstration that 5-azadC, a drug used for epigenetic therapy of myeloproliferative disorders, was able to indefinitely suppress morphological, functional and biochemical features of MTD for cultured HSC. This state was also maintained over several days following removal of the drug before the eventual onset of the classic morphological changes associated with HSC transdifferentiation. Following incorporation into DNA, 5-azadC promotes DNA hypomethylation by inhibiting the activity of the maintenance DNA methyltransferase DNMT1.29 Recent evidence suggests that DNMT1 is rapidly degraded by the proteasomal pathway upon treatment of cells with 5-azadC.30 The requirement to replace 5-azadC-modified DNA and replenish DNMT1 in order to restore DNA methylation explains why removal of the drug from HSC cultures did not lead to a rapid onset of transdifferentiation but instead occurred after a lag-phase of several days. We propose that DNA methylation provides an epigenetic blueprint for the conversion of HSC to myofibroblasts. Perturbation of this blueprint by 5-azadC prevents transdifferentiation. We further suggest that methyl-CpG-binding proteins interpret this epigenetic blueprint to reprogram the HSC transcriptome to that of a myofibroblast. At present there are five known methyl-CpG-binding proteins of which four (MeCP2, MBD1, MBD2 and MBD4) bind through a conserved methyl-CpG-binding domain (MBD) while the fifth protein, Kaiso, binds through a zinc-finger motif.23 MBD1, MBD2, MeCP2 and Kaiso function as transcriptional repressors. Changes in the expression and/or activity of these four proteins may therefore provide mechanisms by which the myofibroblast phenotype is generated and maintained.
MeCP2, the Rett syndrome factor, is the prototypic methyl-CpG-binding protein.31 The abundant neuronal expression of MeCP2, coupled with its association with bulk chromatin, provide hallmarks for a global repressor of neuronal gene expression. However, cDNA microarray studies failed to detect large scale changes in transcriptional derepression in the brains of Mecp2−/− mice.32 There is therefore an emerging concept for a more gene selective role for MeCP2 involving regulation of a smaller number of key regulatory genes such as brain-derived neurotrophic factor (BDNF), a bona fida MeCP2 target.33, 34 However, this concept does not exclude gene selective or global regulatory functions for MeCP2 in non-neuronal cell types of the fully developed adult. Absence of MeCP2 expression in normal adult human and rodent liver demonstrated in the present study helps to explain the lack of hepatic phenotype for Mecp2−/− mice. However, immunohistochemical detection of MeCP2 in diseased livers and localisation to HM implicate MeCP2 as a regulator of wound healing and fibrogenesis. Moreover, the powerful induction of MeCP2 during culture-induced transdifferentiation of HSC raises the possibility of a role as a repressor of key genes controlling the myofibroblast phenotype.
5-azadC treatment of HSC identified PPARγ and IκBα as transcripts that are repressed during transdifferentiation in a DNA methylation-dependent manner. PPARγ is a potential master transcriptional repressor of HSC transdifferentiation which undergoes loss of expression and activity during HSC culture.35 Replenishment of PPARγ expression by adenoviral delivery of PPARγ cDNA into transdifferentiated HSC was associated with reduction in many fibrogenic characteristics of the cell. Diminution of IκBα is also a requirement for transdifferentiation since this leads to reprogramming of basal NF-κB activity to elevated levels that enable the myofibroblast to express proinflammatory genes and to become resistant to apoptosis.11, 26 DNA methylation therefore exerts control over the key fibrogenic (PPARγ) and inflammatory (IκBα) transcriptional regulators of the myofibroblast phenotype. We recently reported a repression mechanism that operates at the IκB-α promoter during MTD of HSC involving the recruitment of the transcriptional repressor CBF1 (RBP-Jκ) to an overlapping NF-κB/CBF1 site.26 We were interested to determine if MeCP2 functions as a component of this mechanism. ChIP analysis at the IκBα promoter revealed a transcriptionally repressed state of chromatin in HSC-derived myofibroblasts that was converted to an active state in cultures treated with 5-azadC. siRNA studies confirmed that MeCP2 is a repressor of IκBα transcription while ChIP assays confirmed that MeCP2 associates with the IκBα promoter in a methylation-dependent manner (inhibited by 5-azadC). However, we consistently failed to detect methyl-CpG at the IκBα promoter and were unable to detect direct binding of MeCP2 to the promoter using native ChIP. We therefore propose a model in which MeCP2 and its associated HDAC and histone methyltransferase activites are recruited to the IκBα promoter indirectly as components of the CBF1 corepression complex (Figure 8). In support of this model, we showed that CBF1 and MeCP2 can be coimmunopreciptitated from myofibroblasts. Of relevance, MeCP2 directly interacts with SMRT, N-CoR1, mSin3A and Ski, all of which are core components of CBF1 corepressor complexes and are removed upon interaction of NotchIC with CBF-1.27, 36, 37 NotchIC prevented MeCP2 repression of IκBα transcription and that this effect was lost in mutant NotchIC proteins unable to bind CBF-1. We conclude that MeCP2 forms a key component of the CBF1 corepressor complex at the IκBα promoter.
Proposed model for the epigenetic regulation of IκBα gene transcription. CBF-1 directly binds at an overlapping NF-κB (κB2)/CBF-1 site located at nucleotides −319 to −310 from the start site of transcription.32 Interaction of CBF1 with MeCP2 located at methyl-CpG sites either 5′ or 3′ to the IκBα promoter (e.g. within exon 1/intron 1) and potentially located in distal regions of silent chromatin, will be facilitated by DNA/chromosome looping. Histone deacetylase (HDAC) and methyltranseferase (HMT) activities associated with MeCP2 will also be recruited to the IκBα promoter leading to histone modification events associated with transcriptional repression
Our model also proposes that MeCP2 recruitment would be facilitated by DNA or chromosomal looping enabling MeCP2 bound at methyl-CpG dinucleotides located at sites proximal (e.g. IκBα exon1/intron 1) or distal to the IκBα promoter to associate with the CBF1 corepressor complex. This is an important feature of the model since although 5-azadC treatment results in release of MeCP2 from the IκBα promoter, the promoter does not contain any detectable methyl-CpG dinucleotides. As such it is necessary to invoke recruitment of MeCP2 bound to methyl-CpG sites outside of the IκBα promoter. This idea is yet to be formally tested, however there is a precedence for MeCP2 mediating so-called ‘silent chromatin loops’ that serve to recruit genes into transcriptionally repressed chromatin domains. This type of mechanism has recently been described at the Dlx5-Dlx6 locus in which association of MeCP2 with its bona fida target gene Dlx5 generates a roughly 11-kb chromatin loop.38
Future studies will also be directed at determining if the epigenetic mechanisms we have identified are a conserved feature for other cell types able to transform into myofibroblasts. Of particular interest would be to determine events in HSC-like, quiescent, vitamin A storing cells in the pancreas, kidney and vocal folds that undergo transdifferentiation in response to injury and infection.2, 4, 9 The myofibroblast generated from these distinct cellular sources is very similar in terms of their key stimulatory growth factors (TGFβ and PDGF), expression of contractile proteins (α-SMA), secretion of fibrogenic factors (types I and III collagen and TIMP-1) and their expression/activity of transcriptional regulators PPARγ, NF-κB and AP-1.19, 20, 21, 22 It is therefore not unreasonable to speculate that the transdifferentiation process responsible for generating these shared phenotypic characteristics is underpinned by a common epigenetic blueprint. Finally, there are obvious implications of our findings for the clinic, especially as we have identified MeCP2 expression in human liver disease. With the emergence of epigenetic therapeutics such as 5-azadC and HDAC inhibitors, there is clearly potential for this class of drug to be applied in the treatment of chronic inflammatory and fibrotic disease.
Materials and Methods
CCl4 liver injury model
Liver fibrosis was generated by 5-week treatment of adult male Sprague–Dawley (200–225 g) rats with CCl4 (CCl4/olive oil, 1 : 1(vol/vol) per kg body weight by intraperitoneal injection twice weekly) as previously described.11 Vehicle control animals were treated intraperitoneally with 1 ml of olive oil/kg body weight at same time intervals. At 24 h after the final CCl4 administration, animals were culled by CO2 asphyxiation and liver samples prepared.
Cell isolation and model of HSC to MTD
Rat HSC were isolated in a quiescent state from normal livers of 350-g Sprague–Dawley rats by sequential perfusion with collagenase and Pronase, followed by discontinuous density centrifugation in 11.5%. Optiprep (Invitrogen Ltd., UK), which enables purification of HSC based on their high buoyancy (due to high vitamin A content). Subsequent culture of quiescent HSC on plastic in serum-containing media results in their spontaneous transdifferentiation to an activated myofibroblastic cell. This process occurs over a period of roughly 7 days and provides a widely used and robust model in which to study the molecular events that regulate MTD. Cultured HSC loose their vitamin A, enter the cell cycle and display de novo expression of a wide variety of profibrogenic and proinflammatory molecules that are characteristic of HM and which are described in detail elsewehere.7, 12, 13 Rat HSC and the LX2 human stellate cell line25 were cultured on plastic in Dulbecco's modified Eagle's medium (DMEM), supplemented with 100 U/ml penicillin, 100 μg/ml streptomycin, 2 mM L-glutamine, and 16% or 10% fetal calf serum, respectively. Cell cultures were maintained at 37°C at an atmosphere of 5% CO2. Human HSCs were isolated from liver resections from two patients (with consent and following approved ethical guidelines) as previously described39 and were subsequently cultured as described for rat HSC.
Crosslinked chromatin immunoprecipitation (XChIP) assay
ChIP assay was carried out using 100 μg crosslinked chromatin prepared from activated rHSC using protocol outlined in Upstate Biotechnology immunoprecipitation (ChIP) assay kit. Antibodies used for immunoprecipitation were all rabbit polyclonal directed against MeCP2, acetyl H3, dimethyl K9 and dimethyl K4 of histone H3 (all from Abcam) and rabbit antiacetylated histone H4 and antiacetyl K14 of histone H3 (Upstate Biotechnology Inc., UK). Ten microgram of each antibody or irrelevant control were used in each ChIP reaction. PCR amplification of the rat IκBα promoter was carried out using specific oligonucleotide primers 5′-cgc taa gag gaa cag cct ag-3′, and 5′- cag ctg gtc gaa aca tgg c-3′.
Cell proliferation assay ([3H]thymidine incorporation assay)
Quiescent rat HSC were seeded into individual wells of a 96-well plate following isolation. Cultures were either left untreated or their media was supplemented with 1 μM 5-azadC for 6 days. Proliferation was measured by adding 0.5 μCi of [3H]thymidine per well during the last 16 h of culture. Incorporated [3H]thymidine was measured by harvesting cells onto UniFilter-96, GF/C plates using a 96-well plate harvester and the plates analysed for [3H]thymidine incorporation using a Packard Topcount Microplate Scintillation counter. Incorporated radioactivity was expressed as counts per minute. Significance of differences was assessed using paired one tailed t-test.
Genomic DNA preparation, bisulphite sequencing and methylation-sensitive restriction enzyme mapping
Genomic DNA was isolated from rat HSCs cultures using DNeasy tissue kit (Qiagen) as per the manufacturer's instructions. Genomic DNA was bisulphite treated using EZ DNA methylation kit (Zymo Research, USA) as per the manufacturers instructions. For methylation sensitive digest, genomic DNA was either left untreated or was cut with AciI CpG methylation sensitive restriction enzyme. Uncut or cut DNA was then used as template in PCR reaction with primers designed to recognize sequence encompassing distal promoter/exon I/proximal intron I (primer set 1), and distal exon I/intron I (primer set 2). Primers used in methylation-sensitive restriction enzyme mapping were- primer set 1 5′-cca ctc agg gct cat caa-3′ (sense) and 5′-cat ctg ctc gta atc ctc-3′ (antisense) and primer set 2 was 5′-tgc cga gcc cac aac agt-3′ (sense) and 5′-gga acc tgc ggg atg aca-3′ (antisense) which were based on the rat IκBα promoter/gene sequence obtained from Ensemble Genome Browser (http://www.ensembl.org/Rattus_norvegicus) numbered ENSRNOG00000007390. PCR reactions were set up, amplified and detected as already described in the reverse transcription (RT)-PCR section.
Immunohistochemistry
Slides were dewaxed in xylene and dehydrated in alcohol, antigen retrieval was achieved by microwaving in citric saline for 15 min. Endogenous peroxidase activity was blocked by hydrogen peroxide pretreatment for 15 min then further blocked using the Vector Laboratories Ltd., UK, avidin/biotin blocking kit, 3 drops/section for 20 min, with Tris-buffered saline (TBS) washes between each stage. Slides were incubated with complete culture medium for 20 min then rabbit polyclonal anti-MeCP2 (Abcam) was diluted 1 : 300 and 1 : 500 for human and rat sections, respectively, in TBS and applied to the slides, then incubated overnight at 4°C. Slides were washed in TBS and then the secondary and the anti-IgG HRP conjugated tertiary antibodies were incubated for 20 min with TBS washes between antibody incubation, (Vector Laboratories, UK). MeCP2 positive cells by were visualised by 3,3′-diaminobenzidine tetrahydrochloride (DAB) staining. Slides were counterstained with Mayers Haematoxylin for 30 s, dehydrated, cleared in xylene and mounted in DPX. For MeCP2/α-SMA dual staining the methods are as described above for an MeCP2 antibody and the rat α-SMA stains with the exception of an additional serum-blocking step between the DAB visualization of α-SMA (brown) and the incubation of rabbit polyclonal anti-MeCP2 antibody. To distinguish between α-SMA and MeCP2 DAB plus nickel (black) was used to visualize MeCP2 in dual stained sections.
Plasmid DNA
All plasmid DNA was prepared using a commercial DNA extraction and isolation kit (Maxiprep, Qiagen). IκBα promoter function was studied using the luciferase reporter vector pIκBα wt-Luc vector containing nucleotides –332 to +35 of the human gene promoter (provided by Professor Ron Hay (St Andrews, UK)). The control Renilla luciferase reporter vector driven by a thymidine kinase promoter (pRLTK) was purchased from Promega (Southampton, UK). CBF1 expression vector pJH282 (provided by Dr Diane Hayward, Baltimore, MD, USA) has been described previously.26 Expression vectors pBos-ZEDN1 (Notch1 IC), pBos-CDN1 (Notch1 IC containing the ankyrin repeat and pest domain), pBos-CDCN1T (Notch1 IC containing the ankyrin repeat domain only) have been described elsewhere.26 SiRNA expressing plasmid psiRNA-MeCP2 expresses siRNA directed against human MeCP2. psiRNA-MeCP2 was custom made (see http://www.invivogen.com/family.php?ID=65&ID_cat=3&ID_sscat=18#groupe132, psiRNA-hMeCP2) based on the backbone of a commercially available vector (InVivoGen) with siRNA target sequence 5′-gga gtc ttc tat ccg atc tgt-3′ which forms a part of hairpin loop 5′-gga gtc ttc tat ccg atc tgt tca aga gac aga tcg gat aga aga ctc c-3′.
Quantitative PCR (Taqman)
Taqman primers and probes for 18S RNA, TIMP1 and procollagen I were as previously described.11 Taqman quantitative RT-PCR reactions were composed of template cDNA, 0.3 μmol/l of forward, reverse and probe primers as well as 12.5 μl of Taqman master mix (Applied Biosystems, Europe) in a final volume of 25 μl. Reaction conditions were 50°C for 2 min and 95°C for 10 min, followed by denaturing for 15 s at 95°C and annealing and extension temperature at 60°C for 1 min and 40 cycles. The relative level of transcriptional difference between quiescent, activated and 5-azadC treated rat HSCs was calculated with the equation (1/(2A)) × 100, where A indicates the cycle threshold (ct) for the gene of interest in activated rHSC minus the ct of the quiescent rat HSC of the same gene after the 18S RNA ct has been deducted for the target gene for each experiment. Significance of differences was assessed using paired one tailed t-test.
SDS-PAGE and immunoblotting
Whole-cell extracts were prepared, and protein concentration of samples determined using a Bradford DC assay kit (Bio-Rad, UK). Whole-cell extracts (30 or 50 μg) from samples of interest were then fractionated by electrophoresis through a 9% sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE). Gels were run at a 100 V for 1.5 h before transfer onto nitrocellulose. After blockade of nonspecific protein binding, nitrocellulose blots were incubated for 1 h with primary antibodies diluted in TBS/Tween 20 (0.075%) containing 3% Marvel. Rabbit polyclonal antibody recognizing MeCP2 (ab-2828, Abcam PLc) was used at 1 μg/ml, as were anti-PPARγ and anti-glial fibrillary acidic protein (GFAP)(Sigma, Poole, UK). Mouse monoclonal antibody directed against IκBα (BioSource, Camerino, USA) was used at 1 : 1000 as was anti-β-actin (Sigma). Following incubation with primary antibodies, blots were washed three times in TBS/Tween-20 before incubation for 1 h in goat anti-mouse or mouse anti-rabbit horseradish peroxidase conjugate antibody at 1 : 4000 dilution in TBS/Tween-20 (0.075%) containing 3% Marvel. After extensive washing in TBS/Tween-20, the blots were processed with distilled water for detection of antigen using the enhanced chemiluminescence system (Amersham Biosciences, USA).
Semiquantitative reverse transcriptase-polymerase chain reaction
Total RNA was purified from isolated cells using the total RNA purification kit (Qiagen, UK) following the manufacturer's instructions and was used to generate first strand cDNA using a random hexamer primer [p(dN)6] and MMLV reverse transcriptase. Primers for mouse MeCP2 were 5′-gag cgg cac tgg gag acc-3′ (sense) and 5′-ctg gat ggt ggt gat gat-3′ (antisense); for mouse β-actin, 5′-tgg aat cct gtg gca tcc at-3′ (sense) and 5′-taa aac gca gct cag taa ca-3′ (antisense); for human MeCP2 5′-agg gag cgg cac cac gag acc-3′ (sense) and 5′-ctt cag tcc ttt ccc gct ctt-3′ (antisense) and for human IκBα 5′-atc acc aac cag cca gaa at-3′ (sense) and 5′-gca ccc aag gac acc aaa ag-3′ (antisense). PCR reactions comprised of 1 μl of cDNA template, 100 ng each of sense and antisense oligonucleotide primers, 2.5 μl of optimized TaqPCR buffer (Promega, UK), 0.4 mmol/l dNTP mixture, and 2 U of Taq polymerase in a total reaction volume of 25 μl. After an initial 5-min incubation at 94°C, PCRs were performed using a 1-min annealing followed by a 2-min elongation step at 72°C and a 45-s denaturation step at 94°C. Varied number of PCR cycles were performed for amplification of all cDNAs (as specified in Figure legend), followed by a final elongation for 10 min at 72°C. PCR products were separated by electrophoresis through a 1% agarose gel and detected by ethidium bromide staining. Expected sizes of specific PCR products were verified by reference to a 1-kb DNA ladder.
Transfections and reporter gene assays
LX2 cells were transfected by the nonliposomal Effectene protocol (Qiagen) according to the manufacturer's instructions. Luciferase assays were performed using a dual luciferase kit (Promega) according to the manufacturer's instructions. pIκBα promoter-driven expression of firefly luciferase was normalized for differences in transfection efficiency to the activity of a cotransfected Renilla vector pRLTK (Promega). Significance of differences was assessed using paired one tailed t-test.
Accession codes
Abbreviations
- ECM:
-
the extracellular matrix
- MTD:
-
myofibroblast transdifferentiation
- HSC:
-
hepatic stellate cells
- HM:
-
hepatic myofibroblast
- α-SMA:
-
smooth muscle-alpha actin
- HDACs:
-
histone deacetylases
References
Desmouliere A, Chaponnier C, Gabbiani G . Tissue repair, contraction and the myofibroblast. Wound Repair Regen 2005; 13: 7–12.
Desmouliere A, Darby IA, Gabbiani G . Normal and pathological soft tissue remodeling: role of the myofibroblast, with specific emphasis on liver and kidney fibrosis. Lab Invest 2003; 83: 1689–1707.
Thannickal VJ, Toews GB, White ES, Lynch III JP, Martinez FJ . Mechanisms of pulmonary fibrosis. Annu Rev Med 2004; 55: 395–417.
Apte MV, Wilson JS . Mechanisms of pancreatic fibrosis. Dig Dis 2004; 22: 272–279.
Wheeler MT, McNally EM . The interaction of coronary tone and cardiac fibrosis. Curr Atheroscler Rep 2005; 7: 219–226.
Desmouliere A, Guyot C, Gabbiani G . The stroma reaction myofibroblast: a key player in the control of tumor cell behaviour. Int J Dev Biol 2004; 48: 509–517.
Mann DA, Smart DE . Transcriptional regulation of hepatic stellate cell activation. Gut 2002; 50: 891–896.
Burke ZD, Tosh D . Therapeutic potential of transdifferentiated cells. Clin Sci (London) 2005; 108: 309–321.
Fuja TJ, Probst-Fuja MN, Titze IR . Transdifferentiation of vocal-fold stellate cells and all-trans retinol-induced deactivation. Cell Tissue Res 2005; 27: 1–8.
Benyon RC, Arthur MJ . Extracellular matrix degadation and the role of hepatic stellate cells. Semin Liver Dis 2001; 21: 373–384.
Oakley F, Meso M, Iredale JP, Green K, Marek CJ, Zhou X et al. Inhibition of inhibitor of kappaB kinases stimulates hepatic stellate cell apoptosis and accelerated recovery from rat liver fibrosis. Gastroenterology 2005; 128: 108–120.
Sancho-Bru P, Bataller R, Gasull X, Colmenero J, Khurdayan V, Gual A et al. Genomic and functional characterization of stellate cells isolated from human cirrhotic livers. J Hepatol 2005; 43: 272–282.
Kristensen DB, Kawada N, Imamura K, Miyamoto Y, Tateno C, Seki S et al. Proteome analysis of rat hepatic stellate cells. Hepatology 2000; 32: 268–277.
Marra F, Valante AJ, Pinzani M, Abboud HE . Cultured human liver fat-storing cells produce monocyte chemotactic protein-1: Regulation by pro-inflammatory cytokines. J Clin Invest 1993; 92: 1674–1680.
Sprenger H, Kaufmann A, Garn H, Lahme B, Gemsa D, Gressner AM . Induction of neutrophil-attracting chemokines in transforming rat hepatic stellate cells. Gastroenterology 1997; 113: 277–285.
Oakley F, Mann J, Nailard S, Smart DE, Mungalsingh N, Constandinou C et al. Nuclear factor-kappaB1 (p50) limits the inflammatory and fibrogenic responses to chronic injury. Am J Pathol 2005; 166: 695–708.
Pinzani M, Marra F . Cytokine receptors and signalling in hepatic stellate cells. Semin Liver Dis 2001; 21: 397–416.
Ding X, Saxena NK, Lin S, Xu A, Srinivasan S, Anania F . The roles of leptin and adiponectin: a novel paradigm in adipocytokine regulation of liver fibrosis and stellate cell biology. Am J Path 2005; 166: 1655–1669.
Massy ZA, Guijarro C, O’Donnell MP, Kim Y, Kashtan CE, Egido J et al. The central role of nuclear factor-kappa B in mesangial cell activation. Kidney Int Suppl 1999; 71: S76–S79.
Masamune A, Kikuta K, Satoh M, Sakai Y, Satoh A, Shimosegawa T . Ligands of peroxisome proliferator-activated receptor-gamma block activation of pancreatic stellate cells. J Biol Chem 2002; 277: 141–147.
Burgess HA, Daugherty LE, Thatcher TH, Lakatos HF, Ray DM, Redonnet M et al. PPARgamma agonists inhibit TGF-beta induced pulmonary myofibroblast differentiation and collagen production: implications for therapy of lung fibrosis. Am J Physiol Lung Cell Mol Physiol 2005; 288: L1146–L1153.
Fitzner B, Sparmann G, Emmrich J, Liebe S, Jaster R . Involvement of AP-1 proteins in pancreatic stellate cell activation in vitro. Int J Colorectal Dis 2004; 19: 414–420.
Jaenisch R, Bird A . Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet Suppl 2003; 33: 245–254.
Margueron R, Trojer P, Reinberg D . The key to development: interpreting the histone code? Curr Opin Genet Dev 2005; 15: 163–176.
Xu L, Hui AY, Albanis E, Arthur MJ, O’Byrne SM, Blaner WS et al. Human hepatic stellate cell lines, LX-1 and LX-2: new tools for analysis of hepatic fibrosis. Gut 2005; 54: 142–151.
Oakley F, Mann J, Ruddell RG, Pickford J, Weinmaster G, Mann DA . Basal expression of IkappaBalpha is controlled by the mammalian transcriptional repressor RBP-J (CBF-1) and its activator Notch1. J Biol Chem 2003; 278: 24359–24370.
Kadesch T . Notch signaling: the demise of elegant simplicity. Curr Opin Genet Dev 2004; 14: 506–512.
Nofziger D, Miyamoto A, Lyons KM, Weinmaster G . Notch signaling imposes two distinct blocks in the differentiation of C2C12 myoblasts. Development 1999; 126: 1689–1702.
Robert MF, Morin S, Beaulieu N, Gauthier F, Chute IC, Barsalou A et al. DNMT1 is required to maintain CpG methylation and aberrant gene silencing in human cancer cells. Nat Genet 2003; 33: 61–65.
Ghoshal K, Datta J, Majumder S, Bai S, Kutay H, Motiwala T et al. 5-Aza-deoxycytidine induces selective degradation of DNA methyltransferase 1 by a proteasomal pathway that requires the KEN box, bromo-adjacent homology domain, and nuclear localization signal. Mol Cell Biol 2005; 25: 4727–4741.
Amir RE, Van den Veyver IB, Wan M, Tran CQ, Francke U, Zoghbi HY . Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat Genet 1999; 23: 185–188.
Tudor M, Akbarian S, Chen RZ, Jaenisch R . Transcriptional profiling of a mouse model for Rett syndrome reveals subtle transcriptional changes in the brain. Proc Natl Acad Sci USA 2002; 26: 15536–15541.
Chen WG, Chang Q, Lin Y, Meissner A, West AE, Griffith EC et al. Derepression of BDNF transcription involves calcium-dependent phosphorylation of MeCP2. Science 2003; 302: 885–889.
Martinowich K, Hattori D, Wu H, Fouse S, He F, Hu Y et al. DNA methylation-related chromatin remodelling in activity-dependent BDNF gene regulation. Science 2003; 302: 890–893.
She H, Xiong S, Hazra S, Tsukamoto H . Adipogenic transcriptional regulation of hepatic stellate cells. J Biol Chem 2005; 280: 4959–4967.
Kokura K, Kaul SC, Wadhwa R, Nomura T, Khan MM, Shinagawa T et al. The Ski protein family is required for MeCP2-mediated transcriptional repression. J Biol Chem 2001; 276: 34115–34121.
Stancheva I, Collins AL, Van den Veyver IB, Zoghbi H, Meehan RR . A mutant form of MeCP2 protein associated with human Rett syndrome cannot be displaced from methylated DNA by notch in Xenopus embryos. Mol Cell 2003; 12: 425–435.
Horike S, Cai S, Miyano M, Cheng JF, Kohwi-Shigematsu T . Loss of silent-chromatin looping and impaired imprinting of DLX5 in Rett syndrome. Nat Genet 2005; 37: 31–40.
Iredale JP, Goddard S, Murphy G, Benyon RC, Arthur MJ . Tissue inhibitor of metalloproteinases-1 and interstitial collagenase expression in autoimmune chronic active hepatitis and activated human hepatic lipocytes. Clin Sci (London) 1995; 89: 75–81.
Acknowledgements
This work was supported by grants from the Wellcome Trust (050443/Z and 068524/Z/02/Z).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Cossu
Rights and permissions
About this article
Cite this article
Mann, J., Oakley, F., Akiboye, F. et al. Regulation of myofibroblast transdifferentiation by DNA methylation and MeCP2: implications for wound healing and fibrogenesis. Cell Death Differ 14, 275–285 (2007). https://doi.org/10.1038/sj.cdd.4401979
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401979
Keywords
This article is cited by
-
Inhibition of DNMT3B expression in activated hepatic stellate cells overcomes chemoresistance in the tumor microenvironment of hepatocellular carcinoma
Scientific Reports (2024)
-
DNA 5mC and RNA m6A modification successively facilitates the initiation and perpetuation stages of HSC activation in liver fibrosis progression
Cell Death & Differentiation (2023)
-
Sarcomatoid renal cell carcinoma: biology, natural history and management
Nature Reviews Urology (2020)
-
A central role for MeCP2 in the epigenetic repression of miR-200c during epithelial-to-mesenchymal transition of glioma
Journal of Experimental & Clinical Cancer Research (2019)
-
Iso-alpha acids from hops (Humulus lupulus) inhibit hepatic steatosis, inflammation, and fibrosis
Laboratory Investigation (2018)