Abstract
Autophagy is in principle a nonselective, bulk degradation system within cells, with a contribution to intracellular protein degradation estimated to be as large as that of the ubiquitin–-proteasome system. The primary roles of autophagy are baseline turnover of intracellular proteins and organelles, production of amino acids in nutrient emergency, and regression of retired tissues. These functions guarantee rejuvenation and adaptation to adverse conditions, and even underlie dynamic processes such as development/metamorphosis. In addition, several other roles for autophagy have recently been discovered, such as presentation of endogenous antigens and degradation of invasive bacteria. This review will discuss the biological significance of autophagy from yeast to higher eukaryotes.
Similar content being viewed by others
Introduction
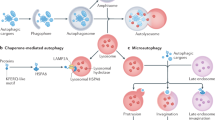
Proteins are dynamically synthesized and degraded. About 200 g of proteins are degraded in our bodies each day, an amount significantly larger than dietary protein intake (about 60–80 g), highlighting the physiological importance of this process. Intracellular protein degradation systems can be roughly classified into two groups: selective and nonselective. The former group includes the ubiquitin-proteasome system and the calpain system present in the cytoplasm, while the latter type occurs mainly in the lysosome. The lysosome is an organelle specialized for degradation, and it degrades extracellular materials, which enter the cell via endocytosis (phagocytosis and pinocytosis). In addition to outside materials, the lysosome can degrade cytoplasmic constituents by a process called autophagy.1, 2 There are at least three types of autophagy, (i) macroautophagy, (ii) chaperon-mediated autophagy (CMA), and (iii) microautophagy. CMA is mediated by chaperones that specifically recognize substrates. Thus, CMA is a type of selective, rather than a bulk or nonselective, degradation system. Microautophagy has been proposed to occur based on lysosome morphology, but its molecular mechanism remains unknown. Since it is believed that macroautophagy is the primary means for cytoplasm-to-lysosome delivery, it is most commonly referred to simply as autophagy. For the purposes of this review, the term ‘autophagy’ will be used to refer to the process of macroautophagy.
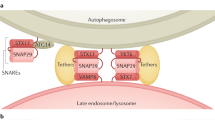
Upon induction of autophagy, a membrane sac called the isolation membrane elongates and encloses a portion of cytoplasm (Figure 1). The origin of this membrane has not yet been identified. It is derived from a small precursor structure, not from a large pre-existing cisterna such as endoplasmic reticulum (ER). However, it is still possible that small vesicles derived from ER and other organelles fuse with the isolation membrane to effect its elongation. Complete enclosure of the isolation membrane results in formation of a double membrane structure called the autophagosome, the diameter of which is usually 0.5–1.5 μm, depending on cell type and conditions. Often, even organelles such as mitochondria and ER fragments are taken up. At this stage, the enclosed cytoplasm is sequestered, but is not degraded until fusion with the lysosome, at which point the inner membrane of the autophagosome and the cytoplasm-derived materials are degraded by lysosomal hydrolases. Since sequestration is basically a nonselective process, autophagy is essentially a bulk degradation system. Yeast genetic studies have identified more than 20 genes (ATG genes) required for autophagy, most of which function in autophagosome formation.3, 4 With these remarkable findings, studies of autophagy have been rapidly progressed for the past decade. In particular, biological significance of autophagy is now being revealed in various organisms.
Autophagosome formation. (Upper) Electron microscopic analysis of nutrient-starved mouse F9 (teratocarcinoma) cells. An arrow indicates autophagosome and arrow heads indicate autolysosomes. (Provided by Dr. Akitsugu Yamamoto). (Lower) Model for autophagy in mammalian cells. A portion of cytoplasm is enclosed by the autophagic isolation membrane, which eventually results in the formation of a double membrane structure, called autophagosome. The outer membrane of the autophagosome then fuses with the lysosome where the cytoplasm-derived materials are degraded
Adaptation to Starvation by ‘Induced Autophagy’
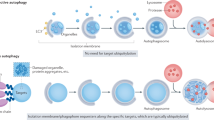
One of the most fundamental and evolutionarily conserved functions of autophagy is its role in the response to starvation. In budding yeast, autophagy is suppressed to undetectable levels under growing conditions. However, it is rapidly (within 30 min) induced during nitrogen starvation.5 This same type of response is also observed in whole animals. We have developed an autophagy indicator mouse model, in which autophagosomes are labeled with LC3 (mammalian Atg8) fused to green fluorescent protein (GFP) (Figure 2).6, 7 Through a systematic analysis of this mouse model, we found that autophagy is upregulated in almost all organs following initiation of starvation. During early starvation, glycogen is used for glucose homeostasis. However, in mammals, glycogen stores are consumed within 1 day. Subsequently, blood glucose is maintained by gluconeogenesis from pyruvate or tricarboxylic acid (TCA) cycle intermediates such oxaloacetate in the liver. Amino acids produced by autophagy in the liver and other tissues could be utilized for this aim. In addition, amino acids can be directly oxidized to produce energy in some tissues, such as muscle, or used for synthesis of proteins that are required for proper starvation adaptation. Autophagy could account for the majority of the so-called protein catabolism during starvation. However, our analyses also revealed some interesting exceptions. In the brain, autophagic activity is very low even under starvation conditions.6 This may be because the brain is not starved by food withdrawal within the time-frame examined, or because an unknown autophagy-inducing factor fails to trigger autophagy in the brain. In contrast, some cell types, such as thymic epithelial cells, show constitutively high autophagic activity.
Molecular mechanism of autophagy regulation and autophagosome formation. Insulin is thought to be one of the major suppressive factors for autophagy. Accordingly, PI3 kinase (class I), PDK1, Rheb and (m)TOR suppress autophagy and PTEN and TSC1/TSC2 induce autophagy. Activity of TOR may be influenced also by amino-acid levels and intracellular ATP. Other growth factors such as IL-3 can inhibit autophagy. Downstream of the TOR-suppressive signaling pathway, Atg proteins function to form the autophagosome, although a direct relationship between TOR and Atgs has not been confirmed. The PI3 kinase complex (class III) including Beclin-1/Atg6 is required for generation of preautophagosome structures. The Atg1 complex is thought to be involved in initiation or regulation of autophagic activity. Elongation of the isolation membrane requires the Atg12 and Atg8 (LC3) ubiquitin-like modification systems. A ubiquitin-like protein, Atg12, is conjugated with Atg5 and forms a large complex with Atg16(L). Only a small fraction of this complex targets to the isolation membrane. Atg8 (LC3 in mammals) is another ubiquitin-like protein that associates with both the isolation membrane and the completed autophagosome as a conjugate with phosphatidylethanolamine (PE)
The critical role played by autophagy in maintaining viability during starvation has been shown in Sacharomyces cerevisiae,8 Dictyostelium discoideum 9 and Drosophila melanogaster.10 In addition, we recently revealed that autophagy is also quite important to help mammals survive a unique period of starvation, the birthing process.11 It is known that neonates face severe starvation at birth due to sudden termination of the placental nutrient supply. Upon extending the GFP-LC3 mice study to the embryonic and perinatal stages, we found that autophagy is transiently upregulated in various tissues soon after birth. We then determined the significance of this neonatal autophagy by analyzing mice deficient in Atg5, a gene essential for elongation of the isolation membrane (Figure 2).12 Atg5 knockout (KO) mice appear almost normal at birth, but exhibit severe nutrient and energy insufficiency within 10 h after birth. In particular, the amino-acid concentrations in plasma and tissues are significantly lower in Atg5 KO mice. Atg5 KO mice display significantly shorter survival times than wild-type mice under starvation conditions, but death could be delayed by forced milk ingestion. Therefore, these findings suggest that neonates use amino acids degraded from tissue proteins for energy homeostasis.11
The importance of amino-acid production via autophagy was also shown in cultured cells. Lum et al.13 reported that IL-3-dependent cells maintain cell viability even after IL-3 withdrawal if the apoptotic pathway is blocked by knocking out both Bax and Bak, both of which are key initiators of apoptosis. Owing to the absence of IL-3 stimulation, these cells are unable to take up enough nutrients from the outside environment, but do still survive for a lengthy period of time. Lum et al. showed that nutrient levels in these cells are maintained by upregulation of autophagy, as RNAi-mediated inhibition of autophagy was quickly lethal to these cells. As a final confirmation, they demonstrated that cell death due to inhibition of autophagy could be avoided by addition of methylpyruvate, a cell-permeable TCA substrate. Taken together, these studies emphasize that increased intracellular generation of amino acids by autophagy is a physiologically important starvation response (Figure 3).
Roles of induced autophagy and baseline autophagy. The level of autophagy is usually low but can be upregulated by starvation such as birth and fasting. The induced autophagy is important for intracellular generation of amino acids. On the other hand, the baseline autophagy is crucial for intracellular quality control
Intracellular Clearance by ‘Baseline Autophagy’: Antidegeneration and Antiaging
Autophagy occurs constitutively at low levels even under normal growth conditions. During embryogenesis, autophagic activity is generally quite low. However, Atg5 KO mice show a suckling defect at birth.11 The reason for this has not yet been determined, but is probably attributable to unknown central nervous system (CNS) dysfunction. Recently, it has been observed that the hepatocytes of liver-specific conditional Atg7 KO mice develop ubiquitin-positive aggregates.14 We also observed such aggregates in Atg5−/− neonates (unpublished data). These findings suggest that baseline autophagy is critical for intracellular clearance (Figure 3). The mechanism underlying the accumulation of ubiquitin-positive aggregates is unknown. Inhibition of continuous turnover of soluble proteins might lead to aggregate formation. Alternatively, aggregates may naturally be degraded by autophagy, although such specific engulfment has never been documented.
In contrast, autophagy-defective yeast cells,8 embryonic stem (ES) cells12 and embryonic fibroblasts11 are quite healthy and show no apparent abnormalities under growing conditions. In rapidly dividing cells, abnormal proteins may be quickly diluted even if they are not degraded. Therefore, intracellular clearance by autophagy should be much more important in postmitotic cells. In this vein, baseline autophagy is under intense scrutiny in the fields of neurodegeneration and aging.
Intracellular protein aggregates containing mutant proteins is a hallmark of many neurodegenerative diseases, such as Parkinson's disease, Alzheimer's disease, polyglutamine diseases (ex. Huntington's disease) and amyotropic lateral sclerosis. It is still controversial whether formation of these protein aggregates is or is not pathogenic. A recent study indicated that, rather than aggregates, high levels of mutant proteins diffusely present within cells may be the largest source of toxicity for cells.15 Thus, formation of aggregates may be a protective mechanism to reduce the levels of mutant proteins elsewhere. Either way, accumulation of misfolded proteins, irrespective of aggregation, are likely mediators of cell damage. Although the ubiquitin–proteasome system has been considered to be the primary means for degradation of these misfolded proteins, autophagy is beginning to receive a great deal of attention.16, 17 Inhibition of autophagy induces aggregate formation, whereas rapamycin-induced upregulation of autophagy reduces it.18, 19, 20, 21 Furthermore, amelioration of neurodegeneration and improvement of behavior were reported in a D. melanogaster Huntington model following treatment with a rapamycin analog, CCI-779.21 It has been speculated that aggregates can be degraded by autophagy (possibly in a specific manner), although this idea has not been proven.16 Alternatively, nonselective degradation (baseline turnover) of diffusely present misfolded proteins could account for prevention of aggregate formation.
Degenerating cells in these diseases often demonstrate vacuoles thought to be related to autophagy. How autophagy is induced under such conditions remains unknown. Ravikumar et al.17 found that mTOR is sequestered into protein aggregates, which impairs mTOR function. They proposed that this could lead to induction of autophagy as a protective mechanism in neurodegeneration. In contrast, it has been speculated that increased levels of autophagy itself might cause degeneration and cell death. However, this idea is still controversial because it has not been proved using genetic models.22
It is known that, in aged cells, protein turnover is slowed and various damaged proteins are accumulated. It has been suggested that autophagic activity might be downregulated with increased age. In addition, Bergamini et al.23 proposed that maintenance of autophagic activity could be at least partly responsible for the life-span extending property of caloric restriction in various species from yeast to mammals. Therefore, autophagic activity may be a critical determinant for life-span. There is possible genetic evidence. Melendez et al.24 demonstrated that beclin-1 (Atg6) is required for the life-span extension observed in a worm mutant lacking the insulin signaling gene daf2. Although the effects of knocking down other atg genes were not demonstrated, these data suggest that autophagy could be an important process regulating longevity.
Degradation of Invading Bacteria
It has been observed that some invasive bacteria may be found in autophagosome-like structures in host cells. 25 This phenomenon was interpreted to be either part of a degradation process or a pregnant phase to allow bacteria to multiply.26 Subsequently, several groups have proven that this is indeed a degradation process. The intracellular pathogen Mycobacterium tuberculosis resides in macrophage phagosomes by interfering with phagosome–lysosome fusion. However, Colombo and Deretic's group found that M. tuberculosis is transported into the lysosome and degraded upon induction of autophagy by starvation or rapamycin.27 Furthermore, they also reported that interferon-γ, which has an antimycobacterial effect, can induce autophagy in macrophages. Thus, they raised the possibility that upregulation of autophagy may partially account for the antibacterial effect of interferon-γ. However, it remains to be clarified whether the phagosome itself is engulfed during autophagy and whether this happens by chance or with specific recognition.
Some bacteria enter into nonphagocytic cells via the endocytic pathway and escape from the endosome into the cytosol. Nakagawa and Yoshimori's group showed that group A Streptococci (GAS), upon escaping from endosomes into the cytosol by a streptolysin O-mediated process, are immediately and specifically sequestered by autophagosome-like structures.28 These GAS-containing structures are eventually fused with lysosomes for degradation. The formation of these structures is Atg5-dependent, and in Atg5−/− embryonic fibroblasts and embryonic fibroblasts, degradation of GAS is significantly reduced, suggesting that this autophagy-like process is indeed bactericidal. A similar phenomenon is observed in cells infected with Shigella.29 However, Shigella may be more clever, because it is able to escape from autophagy. A Shigella mutant lacking the secretory protein IcsB is defective in multiplication within host cells. Ogawa et al.29 reported that the icsB mutant is sequestered by autophagosome-like structures. In contrast, wild-type Shigella evades autophagy by secreting IcsB. They further found that another bacterial protein, VirG, which is required for actin-based intracellular motility, interacts with Atg5. Since the VirG–Atg5 interaction is competitively inhibited by IcsB, these interactions are most likely central to this escape mechanism. The physiological significance of the VirG–Atg5 interaction is unknown. Since Atg5 protein almost exclusively localized to the outer side of the isolation membrane (Figure 2),12 this interaction might be important at an earlier step before sequestration.
Many questions remain regarding bacterial autophagy. Is the membrane used for bacterial trapping the same as in conventional autophagy? What is the induction mechanism? How is specific engulfment achieved? Finally, how important is the contribution of such a killing system during the natural course of bacterial infection? Given that the engulfed contents are not ‘self’, a new term ‘xenophagy’ has been proposed for this system of defense against intracellular microorganisms.30
Supplying Intracellular Antigens to the Major Histocompatibility Complex (MHC) Class II Pathway
T cells recognize antigens presented on two types of MHC on the surface of antigen-presenting cells. In general, cytoplasmic antigens such as viral proteins are processed by the proteasome and presented via MHC Class I to CD8+ (cytotoxic) T cells. On the other hand, exogenous antigens are taken up by antigen-presenting cells via the endocytic pathway and processed in the lysosome. There, antigen peptides become associated with Class II MHC molecules and are then delivered to the plasma membrane and presented to specific CD4+ T cells. Although this is the basic concept, many reports have demonstrated that endogenous proteins, particularly those that are long lived, are also presented on Class II MHC.31 The mechanism by which cytoplasmic proteins are loaded on Class II MHC is incompletely understood, but autophagy has been proposed to be a candidate for this pathway.32, 33 However, Münz laboratory have demonstrated that autophagy is indeed involved in physiological processing of a viral antigen.34 The nuclear antigen 1 of Epstein–Barr virus (EBNA1) is known to escape recognition by CD8+ cytotoxic T cells. This is due to a Gly-Ala repeat domain of EBNA1, which inhibits its processing by the proteasome. However, EBNA1 can be recognized by CD4+ T cells. Paludan et al.34 showed that EBNA1 can be processed by autophagy and then associated with Class II MHC in lysosomes. This ‘autophagic route’ may contribute to presentation not only of viral antigens, but also of self-antigens in autoimmune diseases and tumor antigens. We previously found that thymic epithelial cells show constitutively active autophagy.6 During positive and negative selection, thymic epithelial cells are believed to present self-antigens to lymphocytes. Since thymic epithelial cells are not thought to have phagocytic activity, it is reasonable to hypothesize that they provide self-antigens from their own cytoplasm. Therefore, autophagy might be also involved in central immune tolerance.
Cell Death with Autophagy
It is now apparent that there are several types of programmed cell death other than typical apoptosis. One type, the so-called ‘autophagic cell death’ or ‘type 2 cell death’, is characterized by the appearance of cytoplasmic vacuoles, which are thought to be related to autophagy.35, 36 It should be emphasized that ‘autophagy’ is a term for a degradation process and not a cell death process. Therefore, ‘cell death with autophagy’ would be the most appropriate nomenclature for this kind of cell death.
The role of autophagy in this type of cell death has been an issue of great controversy. Is autophagy a cell death executor or a protective mechanism? Recently, a RNAi-mediated gene knockdown technique was applied in an attempt to answer this question. Yu et al.37 found that z-VAD, a common caspase inhibitor, induces nonapoptotic cell death in L929 and some other cell lines. They showed that knockdown of autophagy factors such as Atg7 and Beclin-1 prevents cell death. Shimizu et al.38 also showed that treatment of Bax−/−Bak−/− cells with apoptogenic reagents such as staurosporine and etoposide induces cell death associated with excessive autophagy. Again, this type of cell death is inhibited by RNAi against Beclin-1 and Atg5. These two studies suggest that autophagy can mediate cell death. However, there are no reports suggesting that autophagy has a causative role in cell death in other species than mammals.
In contrast, Lum et al.13 showed that autophagy acts to ensure survival in IL-3 dependent Bax−/−Bak−/− cells following IL-3 withdrawal. IL-3 withdrawal downregulates surface nutrient transporters, resulting in intracellular starvation. This is consistent with the conventional role of autophagy as a means to ensure cell survival during starvation. Similarly, inhibition of autophagy sensitizes mammalian cells to starvation-induced apoptosis.39 These reports, then, suggest that autophagy is a cytoprotective mechanism. Therefore, whether autophagy acts as a cell death executor or cell protector might depend on cellular environments such as nutrient conditions. In the presence of enough nutrients, prolonged autophagy would not play any protective roles, rather could induce cell death/degeneration.37, 38 However, it should be noted that these studies were performed using cells defective in apoptosis. The fact that Atg5 KO mice are born without an apparent increase or decrease in cell number suggests that autophagy is not critical for cell death, at least during mammalian embryogenesis.11 Also in Dictyostelium, treatment of cells with both starvation and differentiation-inducing factor (DIF) causes cell death with autophagy, but the same treatment can still induce nonvacuolar cell death in autophagy-deficient cells.40 Therefore, the physiological role of autophagy in cell death is still obscure. In addition, the concept of type 2 cell death might be too broad because some of them apparently depend on caspases activation.41
Tumor Suppression
It has been speculated that autophagy may both benefit and hinder tumor development and progression. On the one hand, tumor cells should require autophagy, as tumor tissues are often nutrient-poor environments. On the other, loss of autophagy may lead to accumulation of genotoxic substances in the cytoplasm that could promote mutation and hence tumorigenesis.2, 36, 42 Mouse genetic studies on Beclin-1 (Atg6/Vps30 homolog) indicated that autophagic activity could suppress tumor development.43, 44 Beclin-1 was originally identified as a Bcl2-interacting protein which turned out to be monoallelically deleted in 40–75% of sporadic human breast and ovarian cancers.45 The breast cancer cell line MCF7, which expresses very low levels of Beclin-1,45 as well as Beclin-1−/− ES cells44 are defective in autophagy, suggesting that Beclin-1 is required for autophagy. While Beclin-1−/− mice show early embryonic lethality,44 Yue et al.44 and Qu et al.43 reported that Beclin-1+/− mice develop spontaneous tumors such as lung cancer, hepatocellular carcinoma and lymphoma. Although these studies suggest that Beclin-1 is a tumor suppressor, the implications for autophagy remains to be clarified. Beclin-1/Atg6 is a component of the class III phosphatidylinositol 3 kinase (PI3 kinase) complex both in yeast46 and mammals,47 and Beclin-1 has ability to interact with Bcl-2.48 Finally, the phenotype of Beclin-1−/− mice is much more severe than that of Atg5−/− mice, which appear almost normal at birth.11, 44 Therefore, Beclin-1 likely has additional roles beyond its function in autophagy. It remains to be determined which phenotypes of Beclin−/− and Beclin+/− mice are caused by defects in autophagy.
Development
Thus far, a number of developmental defects have been reported in autophagy mutants in several species (Figure 4). In S. cerevisiae, autophagy mutants are defective in spore formation,8 while autophagy mutants of D. discoideum are defective in normal multicellular developmental processes such as aggregation formation and fruiting body formation.9 Premature death from the third larval to pupal stages was reported in D. melanogaster mutants.10, 49 Finally, dauer formation is abnormal in Caenorhabditis elegans autophagy mutants.24 Although these phenotypes seem to be diverse, all of them are closely related to nutrient status and starvation. For example, sporulation, fruiting body formation, and dauer formation are typical starvation responses. In Drosophila, the mutant third larvae leave food sources and starve as a result. During the pupal stage, larval tissues are degraded and used as nutrients to generate adult tissues. In dying larval tissues such as the salivary glands, massive autophagy is observed.35 Therefore, autophagy is extremely important for developmental steps during which generation of amino acids from endogenous sources is crucial. For mammals, birth is probably the first such experience. In contrast, plant autophagy mutants show only minimal abnormalities, that is, accelerated senescence and reduced seed number.50, 51, 52 This might be because plants have multiple backup systems for starvation and also develop a sophisticated root system to continuously absorb nutrients from the soil.
Concluding Remarks
In this discussion of the pleiotropic roles of autophagy, many of the works cited were published within the past year, highlighting the rapid growth of this field of research. However, many unsolved questions remain, particularly regarding the selectivity of macroautophagy. There is much suggestive data on this issue, particularly regarding the selectivity of organelle degradation, but none is entirely concrete. Most of the roles of autophagy are easily explained without substrate specificity. Probably, only the clear evidence for specificity in mammalian cells is autophagic degradation of bacteria (xenophagy). However, whether autophagy can degrade any self-contents specifically remains unknown. In yeast, Ald6 protein was shown to be rapidly degraded by autophagy compared to other cytosolic proteins.53 Although the mechanism behind this was not identified, there must be some specific recognition. In addition, two vacuolar proteins, aminopeptidase I and mannosidase, are selectively transported to the vacuole by an autophagy-like system called the cytoplasm-to-vacuole targeting (CVT) pathway.4 In this case, a common receptor for these proteins is present on the sequestration membrane. No pathway equivalent to the CVT pathway has been discovered in other species.
Aside from this question, it remains to be discovered how autophagy is regulated in vivo (Figure 2). Amino-acid concentrations are virtually unchanged or even elevated during starvation, suggesting that they are not a driving factor for autophagy under starvation conditions.54, 55 Insulin could be a suppressive factor, but almost no information is currently available regarding autophagy-inducing factors. Since artificial modification of autophagic activity could prove to be an important treatment avenue for degenerative diseases as well as an antiaging strategy, elucidation of the mechanism by which autophagy is regulated is a critical goal for future research.
Abbreviations
- ER:
-
endoplasmic reticulum
- CMA:
-
chaperon-mediated autophagy
- GFP:
-
green fluorescent protein
- TCA:
-
tricarboxylic acid
- ES:
-
embryonic stem
- PI3 kinase:
-
phosphatidylinositol 3 kinase
- TOR:
-
target of rapamycin
- GAS:
-
group A Streptococcus
- MHC:
-
major histocompatibility complex
- EBNA1:
-
Epstein-Barr virus nuclear antigen 1
- CVT:
-
cytoplasm-to-vacuole targeting
References
Levine B and Klionsky DJ (2004) Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev. Cell 6: 463–477
Cuervo AM (2004) Autophagy: in sickness and in health. Trends Cell Biol. 14: 70–77
Klionsky DJ, Cregg JM, Dunn Jr WA, Emr SD, Sakai Y, Sandoval IV, Sibirny A, Subramani S, Thumm M, Veenhuis M and Ohsumi Y (2003) A unified nomenclature for yeast autophagy-related genes. Dev. Cell 5: 539–545
Klionsky DJ (2005) The molecular machinery of autophagy: unanswered questions. J. Cell Sci. 118: 7–18
Takeshige K, Baba M, Tsuboi S, Noda T and Ohsumi Y (1992) Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J. Cell Biol. 119: 301–311
Mizushima N, Yamamoto A, Matsui M, Yoshimori T and Ohsumi Y (2004) In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15: 1101–1111
Mizushima N (2004) Methods for monitoring autophagy. Int. J. Biochem. Cell Biol. 36: 2491–2502
Tsukada M and Ohsumi Y (1993) Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 333: 169–174
Otto GP, Wu MY, Kazgan N, Anderson OR and Kessin RH (2003) Macroautophagy is required for multicellular development of the social amoeba Dictyostelium discoideum. J. Biol. Chem. 278: 17636–17645
Scott RC, Schuldiner O and Neufeld TP (2004) Role and regulation of starvation-induced autophagy in the Drosophila fat body. Dev. Cell 7: 167–178
Kuma A, Hatano M, Matsui M, Yamamoto A, Nakaya H, Yoshimori T, Ohsumi Y, Tokuhisa T and Mizushima N (2004) The role of autophagy during the early neonatal starvation period. Nature 432: 1032–1036
Mizushima N, Yamamoto A, Hatano M, Kobayashi Y, Kabeya Y, Suzuki K, Tokuhisa T, Ohsumi Y and Yoshimori T (2001) Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J. Cell Biol. 152: 657–667
Lum JJ, Bauer DE, Kong M, Harris MH, Li C, Lindsten T and Thompson CB (2005) Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell 120: 237–248
Komatsu M, Waguri S, Ueno T, Iwata J, Murata S, Tanida I, Ezaki J, Mizushima N, Ohsumi Y, Uchiyama Y, Kominami E, Tanaka K and Chiba T (2005) Impairment of starvation-induced and contitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169: 425–434
Arrasate M, Mitra S, Schweitzer ES, Segal MR and Finkbeiner S (2004) Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431: 805–810
Kopito RR (2000) Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol. 10: 524–530
Ravikumar B and Rubinsztein DC (2004) Can autophagy protect against neurodegeneration caused by aggregate-prone proteins? Neuroreport 15: 2443–2445
Teckman JH and Perlmutter DH (2000) Retention of mutant α1-antitrypsin Z in endoplasmic reticulum is associated with an autophagic response. Am. J. Physiol. Gastrointest. Liver Physiol. 279: G961–G974
Ravikumar B, Duden R and Rubinsztein DC (2002) Aggregate-prone proteins with polyglutamine and polyalanine expansions are degraded by autophagy. Hum. Mol. Genet. 11: 1107–1117
Fortun J, Dunn Jr WA, Joy S, Li J and Notterpek L (2003) Emerging role for autophagy in the removal of aggresomes in Schwann cells. J. Neurosci. 23: 10672–10680
Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG, Scaravilli F, Easton DF, Duden R, O’Kane CJ and Rubinsztein DC (2004) Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat. Genet. 36: 585–595
Yuan J, Lipinski M and Degterev A (2003) Diversity in the mechanisms of neuronal cell death. Neuron 40: 401–413
Bergamini E, Cavallini G, Donati A and Gori Z (2004) The role of macroautophagy in the ageing process, anti-ageing intervention and age-associated diseases. Int. J. Biochem. Cell Biol. 36: 2392–2404
Melendez A, Talloczy Z, Seaman M, Eskelinen EL, Hall DH and Levine B (2003) Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301: 1387–1391
Kirkegaard K, Taylor MP and Jackson WT (2004) Cellular autophagy: surrender, avoidance and subversion by microorganisms. Nat. Rev. Microbiol. 2: 301–314
Dorn BR, Dunn Jr WA and Progulske-Fox A (2002) Bacterial interactions with the autophagic pathway. Cell. Microbiol. 4: 1–10
Gutierrez MG, Master SS, Singh SB, Taylor GA, Colombo MI and Deretic V (2004) Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119: 753–766
Nakagawa I, Amano A, Mizushima N, Yamamoto A, Yamaguchi H, Kamimoto T, Nara A, Funao J, Nakata M, Tsuda K, Hamada S and Yoshimori T (2004) Autophagy defends cells against invading group A Streptococcus. Science 306: 1037–1040
Ogawa M, Yoshimori T, Suzuki T, Sagara H, Mizushima N and Sasakawa C (2005) Escape of intracellular Shigella from autophagy. Science 307: 727–731
Levine B (2005) Eating oneself and uninvited guests: autophagy-related pathways in cellular defense. Cell 120: 159–162
Lechler R, Aichinger G and Lightstone L (1996) The endogenous pathway of MHC class II antigen presentation. Immunol. Rev. 151: 51–79
Brazil MI, Weiss S and Stockinger B (1997) Excessive degradation of intracellular protein in macrophages prevents presentation in the context of major histocompatibility complex class II molecules. Eur. J. Immunol. 27: 1506–1514
Nimmerjahn F, Milosevic S, Behrends U, Jaffee EM, Pardoll DM, Bornkamm GW and Mautner J (2003) Major histocompatibility complex class II-restricted presentation of a cytosolic antigen by autophagy. Eur. J. Immunol. 33: 1250–1259
Paludan C, Schmid D, Landthaler M, Vockerodt M, Kube D, Tuschl T and Munz C (2004) Endogenous MHC class II processing of a viral nuclear antigen after autophagy. Science 307: 593–596
Baehrecke EH (2003) Autophagic programmed cell death in Drosophila. Cell Death Differ. 10: 940–945
Gozuacik D and Kimchi A (2004) Autophagy as a cell death and tumor suppressor mechanism. Oncogene 23: 2891–2906
Yu L, Alva A, Su H, Dutt P, Freundt E, Welsh S, Baehrecke EH and Lenardo MJ (2004) Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 304: 1500–1502
Shimizu S, Kanaseki T, Mizushima N, Mizuta T, Arakawa-Kobayashi S, Thompson CB and Tsujimoto Y (2004) Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat. Cell Biol. 6: 1221–1228
Boya P, Gonzalez-Polo RA, Casares N, Perfettini JL, Dessen P, Larochette N, Metivier D, Meley D, Souquere S, Yoshimori T, Pierron G, Codogno P and Kroemer G (2005) Inhibition of macroautophagy triggers apoptosis. Mol. Cell Biol. 25: 1025–1040
Kosta A, Roisin-Bouffay C, Luciani MF, Otto GP, Kessin RH and Golstein P (2004) Autophagy gene disruption reveals a non-vacuolar cell death pathway in Dictyostelium. J. Biol. Chem. 279: 48404–48409
Martin DN and Baehrecke EH (2004) Caspases function in autophagic programmed cell death in Drosophila. Development 131: 275–284
Edinger AL and Thompson CB (2003) Defective autophagy leads to cancer. Cancer Cell 4: 422–424
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A, Rosen J, Eskelinen EL, Mizushima N, Ohsumi Y, Cattoretti G and Levine B (2003) Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J. Clin. Invest. 112: 1809–1820
Yue Z, Jin S, Yang C, Levine AJ and Heintz N (2003) Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc. Natl. Acad. Sci. USA 100: 15077–15082
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H and Levine B (1999) Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 402: 672–676
Kihara A, Noda T, Ishihara N and Ohsumi Y (2001) Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J. Cell Biol. 152: 519–530
Kihara A, Kabeya Y, Ohsumi Y and Yoshimori T (2001) Beclin-phosphatidylinositol 3-kinase complex functions at the trans-Golgi network. EMBO Rep. 2: 330–335
Liang XH, Kleeman L, Jaing HH, Gordon G, Goldman JE, Berry G, Herman B and Levine B (1998) Protection against fatal Sindbis virus encephalitis by Beclin, a novel Bcl-2-interacting protein. J. Virol. 72: 8586–8596
Juhasz G, Csikos G, Sinka R, Erdelyi M and Sass M (2003) The Drosophila homolog of Aut1 is essential for autophagy and development. FEBS Lett. 543: 154–158
Hanaoka H, Noda T, Shirano Y, Kato T, Hayashi H, Shibata D, Tabata S and Ohsumi Y (2002) Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of an Arabidopsis autophagy gene. Plant Physiol. 129: 1181–1193
Doelling JH, Walker JM, Friedman EM, Thompson AR and Veirstra RD (2002) The APG8/12-activating enzyme APG7 is required for proper nutrient recycling and senescence in Arabidopsis thaliana. J. Biol. Chem. 277: 33105–33114
Yoshimoto K, Hanaoka H, Sato S, Kato T, Tabata S, Noda T and Ohsumi Y (2004) Processing of ATG8s, ubiquitin-like proteins, and their deconjugation by ATG4s are essential for plant autophagy. Plant Cell 16: 2967–2983
Onodera J and Ohsumi Y (2004) Ald6p is a preferred target for autophagy in yeast, Saccharomyces cerevisiae. J. Biol. Chem. 279: 16071–16076
Shenoy ST and Rogers QR (1977) Effect of starvation on the charging levels of transfer ribonucleic acid and total acceptor capacity in rat liver. Biochim. Biophys. Acta 476: 218–227
Chikenji T, Elwyn DH and Kinney JM (1983) Protein synthesis rates in rat muscle and skin based on Lysyl-tRNA radioactivity. J. Surg. Res. 34: 68–82
Acknowledgements
I thank Dr. Akitsugu Yamamoto (Nagahama Institute of Bio-Science and Technology) for providing the electron microscopic picture.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by M Piacentini
Rights and permissions
About this article
Cite this article
Mizushima, N. The pleiotropic role of autophagy: from protein metabolism to bactericide. Cell Death Differ 12 (Suppl 2), 1535–1541 (2005). https://doi.org/10.1038/sj.cdd.4401728
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401728
Keywords
This article is cited by
-
Hsp70-Bim interaction facilitates mitophagy by recruiting parkin and TOMM20 into a complex
Cellular & Molecular Biology Letters (2023)
-
The E3 ubiquitin ligase TRIM31 attenuates NLRP3 inflammasome activation in Helicobacter pylori-associated gastritis by regulating ROS and autophagy
Cell Communication and Signaling (2023)
-
Autophagy, a critical element in the aging male reproductive disorders and prostate cancer: a therapeutic point of view
Reproductive Biology and Endocrinology (2023)
-
BMP-induced non-canonical signaling is upregulated during autophagy-mediated regeneration in inflamed mesothelial cells
Scientific Reports (2023)
-
Pyrazole-sulfonamide scaffold featuring dual-tail strategy as apoptosis inducers in colon cancer
Scientific Reports (2023)