Abstract
DNA topoisomerase I (Topo1) contributes to vital biological functions, but its regulation is not clearly understood. The BTBD1 protein was recently cloned on the basis of its interaction with the core domain of Topo1 and is expressed particularly in skeletal muscle. To determine BTBD1 functions in this tissue, the in vitro model used was the C2C12 mouse muscle cell line, which expresses BTBD1 mainly after myotube differentiation. We studied the effects of a stably overexpressed BTBD1 protein truncated of the 108 N-terminal amino-acid residues and harbouring a C-terminal FLAG tag (Δ-BTBD1). The proliferation speed of Δ-BTBD1 C2C12 cells was significantly decreased and no myogenic differentiation was observed, although these cells maintained their capacity to enter adipocyte differentiation. These alterations could be related to Topo1 deregulation. This hypothesis is further supported by the decrease in nuclear Topo1 content in Δ-BTBTD1 proliferative C2C12 cells and the switch from the main peripheral nuclear localization of Topo1 to a mainly nuclear diffuse localization in Δ-BTBTD1 C2C12 cells. Finally, this study demonstrated that BTBD1 is essential for myogenic differentiation.
Similar content being viewed by others
Introduction
DNA topoisomerase 1 (Topo1) belongs to the DNA topoisomerase multimember family, which is essential for DNA-topology modulation. Topo1 relaxes supercoiled duplex DNA by introducing a transient break in the phosphodiester bond of one DNA strand and passing the complementary strand through the cleaved one. Religation involves reformation of the phosphodiester bond and release of the enzyme. Topo1 belongs to the IB subfamily, which acts by DNA rotation, that is, the segments resulting from transient nick in one DNA strand can rotate relatively to each other.1 Several recent papers review the cellular roles of Topo1.1, 2, 3, 4 A role in chromosome condensation has been proposed for Topo1,5 and this enzyme also supports the elongation step of DNA replication, like other type IB enzymes. Concerning gene expression mechanisms, Topo1 is involved in solving topological problems associated with transcription, and it has been shown to be present in actively transcribed regions.2, 4 In addition, Topo1 interacts with transcription factors and participates in the construction of an active TFIID–TFIIA protein complex in a DNA strand breakage-independent manner.6, 7 Topo1 seems to enhance the response to transcription activators in the case of TATA boxes present in promoters and represses basal transcription in the absence of activators.4 Finally, Topo1 contributes to RNA splicing.8, 9 Regarding the Topo1 subcellular localization, a nuclear localization signal is present in its N-terminal region10 and a direct interaction with nucleolin, a major nucleolar protein, has been shown by Bharti et al.11 thus indicating a probable nuclear localization. It is therefore not surprising that repression or overexpression of the Topo1 gene can lead to cell death,12, 13, 14 indicating that preservation of a normal level of Topo1 is crucial for cell survival. Moreover, Topo1 is more abundant in acute lymphocytic leukaemia and several human tumours than in normal tissue,15, 16, 17, 18, 19, 20 possibly because of a defect in its degradation,21 and Topo1 inhibitors are used as antitumour agents. Although many proteins have been described as interacting with Topo1 to regulate its activity.7, 9, 22, 23, 24 in particular through its ubiquitination,21, 25, 26, 27 little is known with regard to Topo1 regulators and it is of major interest to determine the factors involved in Topo1 inhibition.
In the present work, we investigated the role of BTBD1, a protein with an unknown function, which binds to Topo1.28 This protein was detected in humans in 2001 by Carim-Todd et al.29 along with its homologue BTBD2, which shares 82% identity with the BTBD1 amino-acid residue sequence. Both proteins contain a BTB domain (for ‘broad-complex, Tramtrack and bric à brac’),30 which is a conserved protein–protein interaction motif; this domain is also named the POZ domain for ‘Pox virus and Zinc finger’.31 In 2002, Xu et al.28 demonstrated that BTBD1 and BTBD2 interact with Topo1 and that BTBD2 harbours slight Topo1 inhibition activity in vitro. The BTBD1 gene is localized on human chromosome 15q24 and is transcribed as a 3.2 kb mRNA.29 It encodes a 482-amino-acid protein with an N-terminal end containing a proline-rich region and a BTB domain, which are both involved in protein–protein interactions. The C-terminal region (317–482 positions) was described to be essential for binding Topo1.28 BTBD1 is likely expressed in the whole animal kingdom from Caenorhabditis elegans to humans, but not in plants. BTBD1 is expressed in many human tissues and this expression is enhanced in heart, testis and skeletal muscle.28, 29 These observations confirmed previous results obtained by Pietu et al.32 in 1999. These authors found a number of novel human skeletal muscle genes, including a partial cDNA named GENX-702 (GenBank accession numbers: F00425 and Z19393), preferentially expressed in heart, testis and skeletal muscle. This cDNA actually corresponds to BTBD1.
In this study, we assessed the function of BTBD1 in skeletal muscle using the in vitro C2C12 mouse muscle cell line model. The analysis of stable overexpression in C2C12 cells of a BTBD1 protein lacking the 108 N-terminal residues led us to propose that BTBD1 activity is important for muscle growth and essential for muscle differentiation. Moreover, several characteristics of these modified cells may be related to Topo1 deregulation, and BTBD1 may function via a contribution to Topo1 activity regulation.
Results
BTBD1 is expressed in skeletal muscle fibre
BTBD1 expression was first assessed in mouse to validate the relevance of this species as a further experimental model. A wide tissue distribution of mRNA was found as in humans (not shown). Northern blot analysis in heart and skeletal muscle (Figure 1a), which strongly express BTBD1, revealed a 3.2 kb BTBD1 mRNA in mouse.
Muscle BTBD1 expression. (a) Detection of the 3.2 kb BTBD1 mRNA in mouse striated muscles by Northern blot. (b) BTBD1 mRNA expression during mouse C2C12 cell proliferation in GM and differentiation in DM. BTBD1 mRNA expression in C2C12 myoblasts and myotubes was analysed by semiquantitative RT-PCR. (c) BTBD1 expression in human skeletal muscle cells. RNAs were extracted from myocytes or myotubes prepared from single human satellite cells and analysed by RT-PCR. Myogenin was used as control of muscle differentiation. 18S rRNA was used as loading control. Scale bar=20 μm
Skeletal muscle is mainly composed of muscle fibres, fibroblastic structures (epimysium, endomysium, perimysium and tendon), nerves and endothelium. To assess more accurately BTBD1 expression within muscle fibres, we used muscle cells in culture, which are therefore not contaminated by any other components. First, BTBD1 expression was investigated in C2C12 cells, that is, a murine skeletal muscle cell line that is able to differentiate in vitro. In growth medium (GM), C2C12 cells are mononucleated and undifferentiated (myoblasts) (Figure 1b). At confluence, if these cells are switched to differentiation medium (DM), they differentiate into myocytes and after 2–3 days, these mononucleated cells fuse to form syncitial structures (myotubes). BTBD1 transcripts were quantified by semiquantitative RT-PCR using RNAs from proliferative and 4-day-old differentiated C2C12 cells. BTBD1 mRNA was found to be abundant in myotubes, whereas it was barely detectable in myoblasts, which may suggest a link between BTBD1 expression and C2C12 cell myogenic differentiation. Second, we amplified BTBD1 ORF by RT-PCR from mRNAs prepared with myocytes and myotubes obtained from a human clone derived from a single muscle satellite cell33 (Figure 1c). Skeletal muscle cell differentiation was assessed by myogenin cDNA amplification and BTBD1 expression was clearly detected in myocytes and myotubes. Together, these in vitro analyses strongly suggest that BTBD1 is expressed within skeletal muscle fibres.
Expression and subcellular localization of N-terminal truncated BTBD1 in C2C12 myoblasts
C2C12 cells were used to further investigate BTBD1 function, and we established stable C2C12 cell lines overexpressing a truncated BTBD1 lacking the first 108 amino-acid residues of the N-terminal region. This mutated protein will be designated as Δ-BTBD1. This truncated protein should be able to link Topo1 via its intact C-terminal domain.28 The missing N-terminal region includes the proline-rich domain and part of the N-terminal BTB domain, that is, two key domains for potential interactions with undetermined proteins. The Δ-BTBD1 protein was tagged with a FLAG peptide to allow detection of the fusion protein with a specific anti-FLAG antibody.
Western blot screening of stable lines using a monoclonal anti-FLAG antibody revealed a single 40 kDa band signal, while no signal was detected in control cells (Figure 2a). This is comparable to the theoretical 42 kDa size of the Δ-BTBD1-FLAG protein. Anti-FLAG immunofluorescence experiments on Δ-BTBD1 C2C12 cells showed specific cytoplasmic and possibly nuclear Δ-BTBD1 protein localization (Figure 2b and c). In addition, Δ-BTBD1 C2C12 cells were characterized by slower proliferation than the control C2C12 cells, as confirmed by a cell count over a 96 h period. In this experiment, we measured a Δ-BTBD1 C2C12 cell doubling time of 27 versus 19 h for control C2C12 cells (Figure 3). The values are means of three independent Δ-BTBD1 C2C12 clones and three independent empty vector C2C12 clones. Three independent myodulin-FLAG C2C12 clones, which express a nonrelated FLAG fusion protein, were used as negative control and had no significant difference in the doubling time with respect to the empty vector C2C12 clones.34
Expression and subcellular localization of N-terminal truncated stably expressed in C2C12 cells. (a) Whole-cell proteins were extracted from one control C2C12 cell clone, transfected with pcDNA3 empty vector, and three potential clones of Δ-BTBD1 C2C12 cells, and analysed by Western blot using an anti-FLAG antibody. The specific 40 kDa band corresponding to Δ-BTBD1-FLAG protein is indicated. (b) Protein sublocalization in proliferative Δ-BTBD1 C2C12 cells was determined by immunofluorescence using anti-FLAG antibody. Cytoplasmic and possibly nuclear labelling is indicated (arrows). (c) Control C2C12 cells treated with the anti-FLAG antibody are shown as negative control. Scale bar=10 μm
Δ-BTBD1 expression in C2C12 cells blocks myogenic differentiation
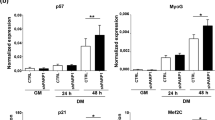
Since the BTBD1 gene was found to be much more expressed in C2C12 myotubes than in C2C12 myoblasts, we decided to study the effect of Δ-BTBD1 overexpression in C2C12 myoblasts under myogenic differentiation conditions. At subconfluence, control and Δ-BTBD1 C2C12 cells were placed in DM. After 4 days, interestingly, no myotubes were detected in Δ-BTBD1 cell culture dishes (Figure 4a), whereas control C2C12 cells were largely fused into myotubes. Δ-BTBD1 C2C12 cells were blocked at a mononucleate state without the cell alignments characteristic of an engagement in cellular fusion. Even after 7 days in DM, Δ-BTBD1 C2C12 cells were unable to fuse and form myotubes. In order to understand this alteration, we studied the expression of two skeletal muscle differentiation markers, that is, MyoD (an early marker) and myogenin (a late marker). Western blot analyses were performed during differentiation. MyoD expression was decreased and myogenin expression was dramatically inhibited in cells overexpressing Δ-BTBD1 (Figure 4b). The strongest effect observed on the late differentiation marker versus the early one suggested that Δ-BTBD1 myoblasts may differentiate in myocytes, but are unable to fuse in myotubes. The inability of Δ-BTBD1 C2C12 cells to differentiate is thus associated with profound modifications in myogenic marker expressions.
Analysis of Δ-BTBD1 C2C12 cell myogenic potential. (a) Morphological analysis of Δ-BTBD1 and control C2C12 cells after 4 days in DM. (b) Effects of Δ-BTBD1 stable expression in C2C12 cells on myogenesis markers. Total proteins were extracted at days 0, 1, 2, 3 and 4 in DM. MyoD and myogenin levels were analysed by Western blot. The total p38 kinase level was used as loading control. Scale bar=20 μm
Δ-BTBD1 C2C12 cells are able to differentiate in adipocytes
Control and Δ-BTBD1 C2C12 confluent cells were treated either with 5 μM rosiglitazone in GM for adipogenesis induction or switched to DM for myogenesis. This experiment was performed to determine the capacity of Δ-BTBD1 C2C12 cells to enter adipocytic differentiation since wild-type C2C12 cells have been shown to positively respond to adipogenic induction.34, 35 After 2 days, in accordance with the above results, Δ-BTBD1 C2C12 cells in DM did not differentiate into myotubes (Figure 5a) and, as expected, myotubes were only detected in control C2C12 cells in DM or with rosiglitazone (Figure 5c and d). In Δ-BTBD1 C2C12 cells treated with rosiglitazone, few rounded cells containing lipid droplets specific to adipocyte structures were detected (Figure 5b). After 7 days, large myotubes were detected in control C2C12 cells (Figure 5g and h), while none were found in Δ-BTBD1 C2C12 cells (Figure 5e and f). Δ-BTBD1 C2C12 cells in DM and control cells treated with rosiglitazone displayed a high number of rounded cells containing oil droplets (Figure 5e–h). The most spectacular result was obtained with Δ-BTBD1 C2C12 cells treated with rosiglitazone: large areas (50% of the culture dish) of adipocytes with the biggest lipid droplets were detected (Figure 5f). These observations were confirmed by Oil red O staining at 7 days of treatment, revealing the presence of neutral triglycerides in all conditions except control C2C12 cells in DM (Figure 5i–p). The findings were further confirmed by a study on the expression of aP2, an adipocyte-specific marker. As expected, aP2 was detected by Western blot in Δ-BTBD1 C2C12 cells after a 7 day treatment with rosiglitazone (data not shown). The presence of neutral lipids in droplets and aP2 expression are characteristic of mature adipocytes. Thus, Δ-BTBD1 overexpression did not alter adipogenesis and BTBD1 did not appear to be essential for this differentiation.
Analysis of Δ-BTBD1 C2C12 cell adipogenic potential. Δ-BTBD1 (a, b, e, f, i, j, m, n) and control (c, d, g, h, k, l, o, p) C2C12 cells were grown either in DM for myogenesis control or in GM containing 5 μM rosiglitazone to study adipogenesis. (a–h) Morphological analysis was performed after 2 and 7 days of treatment. Lipid droplets are indicated in the figure (arrow). (i–p) Adipogenesis was confirmed by Oil red O staining at 7 days of treatment. Lipid droplets containing neutral lipids were stained in red (arrow). Scale bar=10 μm
Δ-BTBD1 C2C12 cells display Topo1 modifications
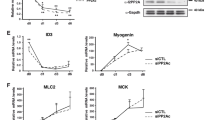
The reported Δ-BTBD1 C2C12 cell alterations in proliferation rate and myogenic differentiation may result from Topo1 deregulation. In order to evaluate this, we studied Topo1 expression levels at proliferation and differentiation stages by Western blotting and cellular sublocalization by immunocytochemistry. We found an almost two-fold lower nuclear Topo1 content in proliferative Δ-BTBD1 C2C12 cells than in control C2C12 cells (Figure 6). When cells were transferred to DM, there was a drastic reduction in nuclear Topo1 content in control C2C12 cells, while no significant reduction was noted in Δ-BTBD1 C2C12 cells during 3 days of differentiation. Furthermore, Topo1 subcellular localization was altered. Besides weak cytoplasmic labelling, Topo1 was mostly observed in a diffuse form in the nuclei of Δ-BTBD1 C2C12 cells (Figure 7a and b) as compared to the main nuclear peripheral localization noted in control C2C12 myoblasts (Figure 7d and e) and myotubes (Figure 7f). The results of quantitative Western blot analysis of Topo1 content in nuclear versus cytoplasmic fractions are shown in Figure 7g. The same number of cells in 100 mm culture dishes were used for Δ-BTBD1 and control C2C12 cells grown in GM close to confluence. Volumes of 1/20 from nuclear and cytoplasmic protein preparations were separated on SDS–PAGE and analysed with anti-Topo1 antibody. The equality of total protein loading in nuclear and cytoplasmic fractions from Δ-BTBD1 and control C2C12 cells was confirmed by the equal Coomassie blue intensity of the protein bands (not shown). Under these conditions, Topo1 was essentially found in the nuclear fractions with a small cytoplasmic amount detected in control C2C12 cells. This confirms that the Topo1 localization modifications observed in Δ-BTBD1 C2C12 cells by immunofluorescence were essentially localization alterations within the nucleus. The absence of Topo1 band in the Δ-BTBD1 C2C12 cell cytoplasm could have been related to the lower amount of Topo1 in these cells. Overall, Δ-BTBD1 overexpression was found to be associated with deregulation of the nuclear Topo1 level and localization.
Study of Topo1 expression in control and Δ-BTBD1 C2C12 cells. Nuclear proteins were extracted from control and Δ-BTBD1 C2C12 cells cultured in GM or after 0, 1, 2 and 3 days in DM. (a) Topo1 protein level was analysed by Western blot and p38 was used for standardization. (b) The values are displayed as histograms. Each band was quantified and Topo1 band intensity values were standardized with the p38 intensity values. Data are means±S.E.M. of three separate experiments
Topo1 subcellular localization in Δ-BTBD1 and control C2C12 cells. Δ-BTBD1 C2C12 cells (a–c) were cultured in GM and control C2C12 cells were cultured in GM (d, e) or for 4 days in DM (f). Cells were fixed and Topo1 was detected by immunofluorescence analysis. Δ-BTBD1 C2C12 cells did not show any labelling with rabbit preimmune serum (c). Nuclei of Δ-BTBD1 C2C12 cells were diffusely labelled (arrow) while control C2C12 cells showed nuclear periphery labelling. The same labelling was found for Δ-BTBD1 C2C12 cells in GM and DM. Weaker cytoplasmic labelling was also observed in all cells, especially in myotubes. (g) Nuclear and cytoplasmic fractions were separated and analysed by Western blots with an anti-Topo1 antibody. Scale bar=10 μm
Discussion
BTBD1 is a new protein that contains several protein–protein interaction motifs. It was discovered at the same time as its homologue BTBD2 by Carim-Todd et al.29 in 2001. Xu et al.28 then showed that BTBD1 and BTBD2 were able to bind. Further studies on BTBD1 and BTBD2 are thus required to fill in the gaps concerning molecular events that regulate Topo1 activity, its involvement in vital biological functions such as mitosis and transcription,3, 36, 37, 38 and in severe pathologies such as cancer and leukaemia.15, 16, 17, 18, 19, 20 Here we attempted to elucidate BTBD1 function in skeletal muscle, where this protein is mostly expressed.
This study was performed in mouse, a model that seemed very appropriate since BTBD1, initially identified in humans, was found to be expressed in this species with the same characteristics as in humans in terms of sequence, messenger size, wide tissue distribution, and preferential expression in skeletal muscle and heart. This wide tissue distribution could be due to the expression in fibroblastic structures present in most tissues. This was ruled out in the case of skeletal muscle since BTBD1 expression was observed in muscle cells grown in vitro. The preferential expression in skeletal muscle and heart suggests that BTBD1 has a more specific function in striated muscles.
BTBD1 function was investigated in modified murine C2C12 muscle cells that overexpress a truncated form of BTBD1. Such molecules may act in a dominant-negative fashion. The expression of truncated BTBD1 decreased the proliferation rate of C2C12 cells, showing that a fully functional BTBD1 is important for cellular proliferation.
N-terminal truncated BTBD1 blocks myogenic differentiation
The most spectacular finding of the present study was the significance of BTBD1 in myogenic differentiation. In vitro myogenic C2C12 differentiation is considered to mimic most in vivo steps. Under low serum culture conditions, confluent myoblasts complete a last cell cycle, then differentiate into myocytes, which align contiguously to each other and finally fuse into myotubes. This is the last structure before in vivo terminal myofibre formation. In cell culture conditions, BTBD1 was found to be more expressed in differentiated myotubes than at the myoblast and myocyte stages (Figure 1). BTBD1 may therefore be essentially a terminal differentiation marker or, alternatively, it may contribute to the differentiation process. This latter hypothesis is strongly corroborated by the finding that Δ-BTBD1 C2C12 cells were blocked at the myocyte stage under differentiation culture conditions and lost the capacity to fuse and become myotubes. This indicates that BTBD1 is somehow involved in C2C12 cell myogenic differentiation and therefore possibly in in vivo myogenesis. More specifically, BTBD1 seems to be crucial for the late step of differentiation, that is, progression from myocytes to fused myotubes, since myogenin, which is a late differentiation marker triggering myocyte fusion, is strongly inhibited in Δ-BTBD1 C2C12 cells, while the expression of MyoD, which is an early differentiation marker, is only decreased.
Does BTBD1 function as a general Topo1 degradation regulator?
Regarding the possible mechanisms underlying BTBD1 activities, the first point that we assessed was a potential action on Topo1 since BTBD1 is known to specifically bind to Topo1. In addition, alterations in proliferation as well as protein synthesis (data not shown), together with a complete defect in differentiation can be typical of Topo1 deregulation. A substantial decrease in Topo1 amount was detected in C2C12 cells between the exponential growth phase and proliferation arrest, and then Topo1 amounts remained constant during differentiation. These variations are similar to Topo1 regulation patterns documented during differentiation of human intestinal epithelial cells in a study in which a constant decrease in cytoplasmic Topo1 level was also noted during differentiation.39 Marked alterations were detected in nuclear Topo1 in Δ-BTBD1 C2C12 cells, which express an N-terminus truncated BTBD1. Firstly, proliferating Δ-BTBD1 C2C12 cells contained less Topo1 than control cells. Secondly, no decrease in Topo1 level was detected between proliferating and differentiating modified cells. Finally, the Topo1 subcellular localization differed between control and Δ-BTBD1 C2C12 cells; the nuclear peripheral localization shifted to a diffuse nuclear localization in Δ-BTBD1 C2C12 cells. Together, these results strongly suggest that BTBD1 is essential for normal Topo1 activity, which in turn is essential for cellular proliferation and differentiation. Furthermore, we speculate that BTBD1 may participate in the negative regulation machinery of Topo1. First, the BTBD1 sequence is highly similar to the BTBD2 sequence, which has been shown in vitro to slightly inhibit Topo1 activity on DNA. Moreover, Xu et al.40 recently described the cellular colocalization of BTBD1 and BTBD2 to cytoplasmic bodies with TRIM5δ, which is a member of the TRIM protein family. Interestingly, these proteins have ubiquitin ligase and multimerization activities. One possible function of BTBD1 could therefore be to address Topo1 to the ubiquitin–proteasome pathway. Indeed, ubiquitin-dependent degradation was reported to be one Topo1 degradation pathway, in addition to trypsin-like serine protease-dependent degradation.21, 26 At this point, it is not clear whether the Topo1–BTBD1 interactions take place in the nucleus or in the cytoplasm. Since BTBD1 has been localized in cytoplasmic bodies,40 the simplest hypothesis to explain BTBD1 function is that BTBD1 binds to Topo1 through its C-terminal BTB domain after Topo1 transportation in the cytoplasm. The N-terminal BTB domain would interact with the appropriate proteins to address Topo1 to proteasome. The BTBD1 contribution to Topo1 activity inhibition would thus be mainly to elicit Topo1 degradation. Another nonexclusive hypothesis is that a small pool of BTBD1 molecules could first interact with Topo1 in the nucleus and thereby contribute to Topo1 activity inhibition, and would then participate in Topo1 transport to cytoplasmic proteasome through the N-terminal BTB domain. A full N-terminal BTB domain is missing in the Δ-BTBD1 mutant protein, thus preventing it from achieving these latter functions. According to both hypotheses, the consequence would be alteration of the Topo1 degradation/synthesis balance, which in turn would impair normal functioning of Topo1. Because of the wide spectrum of activities of Topo1, it is at this point impossible to understand exactly how Topo1 can mediate the effects of BTBD1 on cellular proliferation and differentiation. In addition, we cannot rule out that BTBD1 function involves other mechanisms not related to Topo1 activity.
Specific role of BTBD1 in myogenic differentiation
C2C12 cells have been described as having pluripotent mesenchymal precursor cell characteristics. Indeed, these cells treated with rosiglitazone or ‘bone morphogenetic protein 2’ are able to differentiate in adipocytes34, 35 or osteoblasts,41 respectively. Δ-BTBD1 C2C12 cells were treated with rosiglitazone, a member of the glitazone family known to induce adipogenesis in various cell types, and no effect of BTBD1 was detectable on adipogenic differentiation of C2C12 cells. It would therefore be worthwhile investigating why BTBD1 effects on differentiation are specific to myogenesis although BTBD1 and Topo1 expressions have a wide tissue distribution and Topo1 is likely involved in non-tissue-specific cellular differentiation. In the present work, we showed that N-terminal truncated BTBD1 alters the expression of MyoD and especially myogenin, which are key myogenic regulatory factors. It is clear that they may mediate a specific role of BTBD1 in myogenic differentiation. However, further investigation will be necessary to elucidate the molecular mechanisms linking BTBD1, MyoD and myogenin. Such mechanisms imply that there are particular links between myogenic regulatory factors and BTBD1 and/or Topo1. Indeed, BTBD1 may interact either directly or through its Topo1 interactions with these factors, but no such interactions have been documented to date. Interestingly, many lines of evidence have shown that the myogenic regulation cascade functions via chromatin-modifying enzymes. More precisely, myogenic regulatory factors, such as MyoD, are known to recruit chromatin-remodelling proteins, as recently reviewed by McKinsey et al.42 This is the case particularly with respect to ATP-dependent SWI/SNF enzymes, which were shown to be essential for the induction of muscle-specific genes by some myogenic regulatory factors43 and the class II histone deacetylases HDAC4 and HDAC5.44 In fact, these latter compounds inhibit members of the myocyte enhancer factor-2 (MEF2) transcription factor family that associate with myogenic regulatory factors to activate skeletal myogenesis. The importance of enzymes that remodel chromatin for myogenesis is highlighted by the role of HDACs as targets for signalling pathways controlled by insulin-like growth factor and CaM kinase during myogenic differentiation.45 Structural domains that participate in chromatin reorganization and independent of the transcription activation domain have even been described in MyoD molecules.46 To our knowledge, no role of Topo1 has been discovered in modifications of chromatin associated with molecular muscle differentiation mechanisms, but it would be reasonable to assume that this role exists. By this hypothesis, the impact of BTBD1 on myogenesis would be due to its interactions with Topo1 and therefore to the possible effects on the regulation of chromatin structure. This is the working hypothesis we favoured for further investigations.
This study suggests that N-terminal truncated BTBD1 acts in a dominant-negative manner to prevent muscle cell differentiation and therefore that the BTBD1 function may be required for normal muscle cell differentiation, which could be related to its high expression in skeletal muscle. Several lines of evidence suggest that the already known interactions of BTBD1 with Topo1 could be part of the myogenic role of BTBD1. This raises many questions on the molecular events and interactions involved in Topo1 activity regulation and their impact on myogenic mechanisms.
Materials and Methods
Materials
Restriction enzymes were purchased from Invitrogen (Cergy Pontoise, France) and cell culture reagents were from Sigma (Saint Quentin Fallavier, France). Rosiglitazone (BRL-49653-C) was obtained from GlaxoSmithKline (Worthing, UK). Antibodies were purchased from the following vendors: anti-MyoD monoclonal antibody (M-318) and anti-Topo1 rabbit polyclonal antibody (H-300) from Santa Cruz (Heidelberg, Germany), anti-FLAG monoclonal antibody from Euromedex (Mundolsheim, France), anti-myogenin monoclonal antibody (F5D) from Pharmingen (Le Pont de Claix, France), anti-p38 rabbit polyclonal antibody from Sigma (Saint Quentin Fallavier, France), swine anti-rabbit IgG and rabbit HRP- or FITC-conjugated anti-mouse IgG from Dako (Trappes, France).
Animals and tissues
All procedures were approved by the local Centre National de la Recherche Scientifique ethics committee. C57BL/6 animals (10-week-old) were used for mouse tissue RNA preparations. All tissues were immediately frozen in liquid nitrogen after excision.
Plasmid
A cDNA encoding a BTBD1 large fragment truncated from the 108 N-terminal amino-acid residues, named Δ-BTBD1, was FLAG tagged at the C-terminus by PCR using Advantage Klen Taq polymerase (Ozyme, Saint Quentin Yvelines, France) and mouse BTBD1 ORF cDNA as template. Forward and reverse primer sequences were 5′-GAGAGGGATCCTTTGACGCCATGTTCAACGGCGGCATGGCCACCACGTCG-3′ and 5′-CTCTCGCGGCCGTTAGTTAGTTATTTGTCATCATCGTCCTTATAGTCTGTATAAAATATGATTTCTGGAAT-3′, respectively. The Δ-BTBD1 PCR products were gel-purified and cloned into pGEM-T Easy vector (Promega, Charbonnieres, France). Inserts were excised with BamHI and NotI endonucleases and subcloned into BamHI and NotI sites of the pcDNA3 vector (Invitrogen). Plasmid constructions were checked by automatic sequencing, performed by Genome Express SA (Grenoble, France).
Cell culture
The human myoblast clone was a generous gift from Dr. L Bernheim and isolated from single satellite cells, as previously described.33 The C2C12 mouse skeletal muscle cell line (CRL 1772) was obtained from the American Type Culture Collection. Wild-type and transfected cell lines were grown at 37°C in a 5% CO2 atmosphere in a GM composed of Dulbecco's modified Eagle's medium with 4.5 g/l glucose, 10% fetal calf serum and 1% v/v penicillin/streptomycin. Myotube C2C12 differentiation was induced by withdrawing fetal calf serum from confluent cells and adding 10 μg/ml insulin, 5 μg/ml transferrin and 2% horse serum; this medium was named DM. Adipogenesis was induced by adding rosiglitazone (5 μM final concentration) in GM to confluent cells. Only experiments reproducible on three independent modified cell lines were considered because of the clonal heterogeneity of C2C12 cells. Cell counts were performed with the CASY®1 Cell Counter Model TT from Schärfe System Gmbh (Reutligen, Germany).
Stable transfection
Cell transfection was performed using the calcium phosphate procedure. C2C12 cells were transfected with 20 μg of pcDNA3-Δ-BTBD1-FLAG construct (Δ-BTBD1 C2C12 cell line) or pcDNA3 empty vector alone (control C2C12 cell line). C2C12 cells were diluted about 103-fold the day after transfection and grown in 0.8 μg/ml of G418 until selection was complete. Viable colonies were further screened for transgene expression by Western blotting.
RNA analysis
Total RNA was prepared with the RNAInstaPure kit (Eurogentec, Seraing, Belgium). For Northern blot analysis, approximately 50 μg of total RNA was separated and transferred onto nylon membranes according to standard procedures. Hybridizations were performed at 42°C in the presence of 50% formamide as previously described.47 PCR-amplified products of mouse BTBD1 ORF and 18S rRNA were used as probes and labelled with [α-32P]dCTP using the Prime-a-gene labelling system (Promega). Complete mouse BTBD1 ORF cDNA was amplified with forward 5′-GTTTTGTCCATAAGTGGGC-3′ and reverse 5′-TGAAGGTGAAAAAGCTGTGC-3′ primers. The 18S rRNA cDNA fragment was amplified with forward 5′-AGTTGGTGGAGCGATTTGTC-3′ and reverse 5′-GGCCTCACTAAACCATCCAA-3′ primers. The myogenin cDNA fragment was amplified with forward 5′-GCGCAGGCT CAAGAAAGTGAAT-3′ and reverse 5′-TTTCGTCTGGGAAGGCAACA-3′ primers. For RT-PCR analysis, reverse transcription was carried out with M-MLV reverse transcriptase (Invitrogen) and oligodT on 1 μg of total RNA. PCR was performed with Taq polymerase (Invitrogen). Routine PCR amplifications were analysed after 30 cycles. Agarose electrophoresis bands were quantified with Pcbas software in the linear amplification range and normalized with 18S rRNA amplification. PCR-amplified fragment sequences were controlled with several restriction enzymes.
Western blot analysis
Whole-cell, nuclear and cytoplasmic protein extracts were prepared by Triton lysis as previously described.39, 48 Equal amounts of proteins (25 μg or 1/20 of preparations for subcellular analysis) were separated by SDS–PAGE on 6 or 10% acrylamide gels. Proteins were transferred to Hybond-C extra membranes (Amersham Biosciences, Buckinghamshire, UK) and stained with Ponceau red to monitor transfers. Blots were then incubated for 1 h at room temperature in TBS (0.05 M Tris, 0.15 M NaCl, pH 7.4) with 0.1% Tween-20 and 5% nonfat powdered milk. Membranes were probed overnight at 4°C with a primary antibody. Antibodies were used at the following dilution in TBS: 0.1% Tween-20, 3% BSA: anti-p38 at 1 : 10 000, anti-FLAG and anti-myogenin at 1 : 5000, anti-MyoD at 1 : 400 and anti-Topo1 at 1 : 200. The membranes were then probed for 1 h at room temperature with an HRP-conjugated secondary antibody. After each incubation, membranes were washed three times in TBS and 0.1% Tween-20 for 10 min. Proteins were then visualized using the Amersham ECL system.
Immunocytochemistry
Cells were grown on 22 mm glass coverslips. They were rinsed once in sterile phosphate-buffered saline (PBS) and fixed as previously described49 for 15 min at room temperature in 10% formaldehyde, followed by methanol permeabilization for 10 min at −20°C. All of the following steps were performed at room temperature unless otherwise specified. Cells were washed in PBS. Blocking was performed for 30 min in PBS 3% BSA. Cells were incubated for 2 h at 37°C with mouse anti-FLAG or anti-Topo1 antibody, at 1 : 500 and 1 : 100 dilution in PBS 3% BSA, respectively. After rinses with PBS, bound antibodies were detected with FITC-conjugated secondary antibody for 1 h at 37°C. Cells were washed with PBS and distilled water, then mounted in Mowiol (Calbiochem, La Jolla, USA) for visualization on an Olympus BH2 epifluorescence microscope. Photographs were made with a Nikon Coolpix 990 digital camera.
Oil red O staining
Staining was performed as previously described.50 Cells were mounted with 70% glycerol for visualization on a Leitz Wetzlar light microscope. Photographs were made with a Nikon Coolpix 990 digital camera.
Abbreviations
- BTB:
-
broad-complex, Tramtrack and bric à brac
- DM:
-
differentiation medium
- GM:
-
growth medium
- Topo1:
-
DNA topoisomerase 1
References
Champoux JJ (2001) DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70: 369–413
Nitiss JL (1998) Investigating the biological functions of DNA topoisomerases in eukaryotic cells. Biochim. Biophys. Acta 1400: 63–81
Wang JC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat. Rev. Mol. Cell Biol. 3: 430–z440
Wang JC (1996) DNA topoisomerases. Annu. Rev. Biochem. 65: 635–692
Castano IB, Brzoska PM, Sadoff BU, Chen H and Christman MF (1996) Mitotic chromosome condensation in the rDNA requires TRF4 and DNA topoisomerase I in Saccharomyces cerevisiae. Genes Dev. 10: 2564–2576
Shykind BM, Kim J, Stewart L, Champoux JJ and Sharp PA (1997) Topoisomerase I enhances TFIID–TFIIA complex assembly during activation of transcription. Genes Dev. 11: 397–407
Merino A, Madden KR, Lane WS, Champoux JJ and Reinberg D (1993) DNA topoisomerase I is involved in both repression and activation of transcription. Nature 365: 227–232
Rossi F, Labourier E, Forne T, Divita G, Derancourt J, Riou JF, Antoine E, Cathala G, Brunel C and Tazi J (1996) Specific phosphorylation of SR proteins by mammalian DNA topoisomerase I. Nature 381: 80–82
Straub T, Grue P, Uhse A, Lisby M, Knudsen BR, Tange TO, Westergaard O and Boege F (1998) The RNA-splicing factor PSF/p54 controls DNA-topoisomerase I activity by a direct interaction. J. Biol. Chem. 273: 26261–26264
Alsner J, Svejstrup JQ, Kjeldsen E, Sorensen BS and Westergaard O (1992) Identification of an N-terminal domain of eukaryotic DNA topoisomerase I dispensable for catalytic activity but essential for in vivo function. J. Biol. Chem. 267: 12408–12411
Bharti AK, Olson MO, Kufe DW and Rubin EH (1996) Identification of a nucleolin binding site in human topoisomerase I. J. Biol. Chem. 271: 1993–1997
Bjornsti MA, Benedetti P, Viglianti GA and Wang JC (1989) Expression of human DNA topoisomerase I in yeast cells lacking yeast DNA topoisomerase I: restoration of sensitivity of the cells to the antitumor drug camptothecin. Cancer Res. 49: 6318–6323
Lee MP, Brown SD, Chen A and Hsieh TS (1993) DNA topoisomerase I is essential in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 90: 6656–6660
Onishi Y, Azuma Y, Sato Y, Mizuno Y, Tadakuma T and Kizaki H (1993) Topoisomerase inhibitors induce apoptosis in thymocytes. Biochim. Biophys. Acta 1175: 147–154
Giaccone G, van Ark-Otte J, Scagliotti G, Capranico G, van der Valk P, Rubio G, Dalesio O, Lopez R, Zunino F, Walboomers J and Pinedo HM (1995) Differential expression of DNA topoisomerases in non-small cell lung cancer and normal lung. Biochim. Biophys. Acta 1264: 337–346
Goodlad GA and Clark CM (1993) Altered levels of histone H1(0) and DNA topoisomerase activity in the liver of the tumour-bearing rat. Biochem. Mol. Biol. Int. 30: 1043–1052
Goodlad GA and Clark CM (1995) H1 histone sub-type distribution and DNA topoisomerase activity in skeletal muscle of tumour-bearing rats. Cancer Lett. 98: 111–114
Husain I, Mohler JL, Seigler HF and Besterman JM (1994) Elevation of topoisomerase I messenger RNA, protein, and catalytic activity in human tumors: demonstration of tumor-type specificity and implications for cancer chemotherapy. Cancer Res. 54: 539–546
Rowinsky EK, Adjei A, Donehower RC, Gore SD, Jones RJ, Burke PJ, Cheng YC, Grochow LB and Kaufmann SH (1994) Phase I and pharmacodynamic study of the topoisomerase I-inhibitor topotecan in patients with refractory acute leukemia. J. Clin. Oncol. 12: 2193–2203
van der Zee AG, de Jong S, Keith WN, Hollema H, Boonstra H and de Vries EG (1994) Quantitative and qualitative aspects of topoisomerase I and II alpha and beta in untreated and platinum/cyclophosphamide treated malignant ovarian tumors. Cancer Res. 54: 749–755
Chen HJ, Hwong CL, Wang CH and Hwang J (2000) Degradation of DNA topoisomerase I by a novel trypsin-like serine protease in proliferating human T lymphocytes. J. Biol. Chem. 275: 13109–13117
Gobert C, Skladanowski A and Larsen AK (1999) The interaction between p53 and DNA topoisomerase I is regulated differently in cells with wild-type and mutant p53. Proc. Natl. Acad. Sci. USA 96: 10355–10360
Karayan L, Riou JF, Seite P, Migeon J, Cantereau A and Larsen CJ (2001) Human ARF protein interacts with topoisomerase I and stimulates its activity. Oncogene 20: 836–848
Wang Z and Roeder RG (1998) DNA topoisomerase I and PC4 can interact with human TFIIIC to promote both accurate termination and transcription reinitiation by RNA polymerase III. Mol. Cell 1: 749–757
Desai SD, Li TK, Rodriguez-Bauman A, Rubin EH and Liu LF (2001) Ubiquitin/26S proteasome-mediated degradation of topoisomerase I as a resistance mechanism to camptothecin in tumor cells. Cancer Res. 61: 5926–5932
Desai SD, Liu LF, Vazquez-Abad D and D'Arpa P (1997) Ubiquitin-dependent destruction of topoisomerase I is stimulated by the antitumor drug camptothecin. J. Biol. Chem. 272: 24159–24164
Hwong CL, Chen CY, Shang HF and Hwang J (1993) Increased synthesis and degradation of DNA topoisomerase I during the initial phase of human T lymphocyte proliferation. J. Biol. Chem. 268: 18982–18986
Xu L, Yang L, Hashimoto K, Anderson M, Kohlhagen G, Pommier Y and D'Arpa P (2002) Characterization of BTBD1 and BTBD2, two similar BTB-domain-containing Kelch-like proteins that interact with Topoisomerase I. BMC Genomics 3: 1
Carim-Todd L, Sumoy L, Andreu N, Estivill X and Escarceller M (2001) Identification and characterization of BTBD1, a novel BTB domain containing gene on human chromosome 15q24. Gene 262: 275–281
Zollman S, Godt D, Prive GG, Couderc JL and Laski FA (1994) The BTB domain, found primarily in zinc finger proteins, defines an evolutionarily conserved family that includes several developmentally regulated genes in Drosophila. Proc. Natl. Acad. Sci. USA 91: 10717–10721
Bardwell VJ and Treisman R (1994) The POZ domain: a conserved protein–protein interaction motif. Genes Dev. 8: 1664–1677
Pietu G, Eveno E, Soury-Segurens B, Fayein NA, Mariage-Samson R, Matingou C, Leroy E, Dechesne C, Krieger S, Ansorge W, Reguigne-Arnould I, Cox D, Dehejia A, Polymeropoulos MH, Devignes MD and Auffray C (1999) The genexpress IMAGE knowledge base of the human muscle transcriptome: a resource of structural, functional, and positional candidate genes for muscle physiology and pathologies. Genome Res. 9: 1313–1320
Baroffio A, Aubry JP, Kaelin A, Krause RM, Hamann M and Bader CR (1993) Purification of human muscle satellite cells by flow cytometry. Muscle Nerve 16: 498–505
Yeow K, Phillips B, Dani C, Cabane C, Amri EZ and Derijard B (2001) Inhibition of myogenesis enables adipogenic trans-differentiation in the C2C12 myogenic cell line. FEBS Lett. 506: 157–162
Grimaldi PA, Teboul L, Inadera H, Gaillard D and Amri EZ (1997) Trans-differentiation of myoblasts to adipoblasts: triggering effects of fatty acids and thiazolidinediones. Prostaglandins Leukot. Essent. Fatty Acids 57: 71–75
Desai SD, Zhang H, Rodriguez-Bauman A, Yang JM, Wu X, Gounder MK, Rubin EH and Liu LF (2003) Transcription-dependent degradation of topoisomerase I-DNA covalent complexes. Mol. Cell. Biol. 23: 2341–2350
Kimura K and Hirano T (1997) ATP-dependent positive supercoiling of DNA by 13S condensin: a biochemical implication for chromosome condensation. Cell 90: 625–634
Wang JC and Giaever GN (1988) Action at a distance along a DNA. Science 240: 300–304
Ulukan H, Muller MT and Swaan PW (2001) Downregulation of topoisomerase I in differentiating human intestinal epithelial cells. Int. J. Cancer 94: 200–207
Xu L, Yang L, Moitra PK, Hashimoto K, Rallabhandi P, Kaul S, Meroni G, Jensen JP, Weissman AM and D'Arpa P (2003) BTBD1 and BTBD2 colocalize to cytoplasmic bodies with the RBCC/tripartite motif protein, TRIM5delta. Exp. Cell Res. 288: 84–93
Lee KS, Kim HJ, Li QL, Chi XZ, Ueta C, Komori T, Wozney JM, Kim EG, Choi JY, Ryoo HM and Bae SC (2000) Runx2 is a common target of transforming growth factor beta1 and bone morphogenetic protein 2, and cooperation between Runx2 and Smad5 induces osteoblast-specific gene expression in the pluripotent mesenchymal precursor cell line C2C12. Mol. Cell. Biol. 20: 8783–8792
McKinsey TA, Zhang CL and Olson EN (2002) Signaling chromatin to make muscle. Curr. Opin. Cell Biol. 14: 763–772
Roy K, de la Serna IL and Imbalzano AN (2002) The myogenic basic helix–loop–helix family of transcription factors shows similar requirements for SWI/SNF chromatin remodeling enzymes during muscle differentiation in culture. J. Biol. Chem. 277: 33818–33824
McKinsey TA, Zhang CL, Lu J and Olson EN (2000) Signal-dependent nuclear export of a histone deacetylase regulates muscle differentiation. Nature 408: 106–111
Lu J, McKinsey TA, Zhang CL and Olson EN (2000) Regulation of skeletal myogenesis by association of the MEF2 transcription factor with class II histone deacetylases. Mol. Cell 6: 233–244
Gerber AN, Klesert TR, Bergstrom DA and Tapscott SJ (1997) Two domains of MyoD mediate transcriptional activation of genes in repressive chromatin: a mechanism for lineage determination in myogenesis. Genes Dev. 11: 436–450
Tkatchenko AV, Pietu G, Cros N, Gannoun-Zaki L, Auffray C, Leger JJ and Dechesne CA (2001) Identification of altered gene expression in skeletal muscles from Duchenne muscular dystrophy patients. Neuromuscular Disord. 11: 269–277
Cabane C, Englaro W, Yeow K, Ragno M and Derijard B (2003) Regulation of C2C12 myogenic terminal differentiation by MKK3/p38alpha pathway. Am. J. Physiol. Cell. Physiol. 284: C658–C666
Brunet A, Roux D, Lenormand P, Dowd S, Keyse S and Pouyssegur J (1999) Nuclear translocation of p42/p44 mitogen-activated protein kinase is required for growth factor-induced gene expression and cell cycle entry. EMBO J. 18: 664–674
Aubert J, Dessolin S, Belmonte N, Li M, McKenzie FR, Staccini L, Villageois P, Barhanin B, Vernallis A, Smith AG, Ailhaud G and Dani C (1999) Leukemia inhibitory factor and its receptor promote adipocyte differentiation via the mitogen-activated protein kinase cascade. J. Biol. Chem. 274: 24965–24972
Acknowledgements
We thank Dr. Geneviève Piétu and Dr. Charles Auffray for providing the initial GENX-702 cDNA clone. We are grateful to David Manley for the English revision of the manuscript. D.F. Pisani is a recipient of grants from the Fondation pour la Recherche Médicale. This work was supported by the Centre National de la Recherche Scientifique and the Association Française contre les Myopathies.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by M Blagosklonny
Rights and permissions
About this article
Cite this article
Pisani, D., Cabane, C., Derijard, B. et al. The topoisomerase 1-interacting protein BTBD1 is essential for muscle cell differentiation. Cell Death Differ 11, 1157–1165 (2004). https://doi.org/10.1038/sj.cdd.4401479
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401479
Keywords
This article is cited by
-
Novel Cul3 binding proteins function to remodel E3 ligase complexes
BMC Cell Biology (2014)
-
PED/PEA-15 induces autophagy and mediates TGF-beta1 effect on muscle cell differentiation
Cell Death & Differentiation (2012)
-
Enhancement of Myogenic and Muscle Repair Capacities of Human Adipose–derived Stem Cells With Forced Expression of MyoD
Molecular Therapy (2009)