Abstract
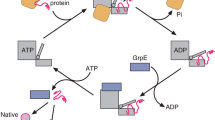
The bacterial protein ClpA, a member of the Hsp100 chaperone family, forms hexameric rings that bind to the free ends of the double-ring serine protease ClpP (refs 1, 2). ClpA directs the ATP-dependent degradation of substrate proteins bearing specific sequences3,4,5, much as the 19S ATPase ‘cap’ of eukaryotic proteasomes functions in the degradation of ubiquitinated proteins6,7,8. In isolation, ClpA and its relative ClpX can mediate the disassembly of oligomeric proteins9,10; another similar eukaryotic protein, Hsp104, can dissociate low-order aggregates11. ClpA has been proposed to destabilize protein structure, allowing passage of proteolysis substrates through a central channel into the ClpP proteolytic cylinder12,13,14. Here we test the action of ClpA on a stable monomeric protein, the green fluorescent protein GFP, onto which has been added an 11-amino-acid carboxy-terminal recognition peptide, which is responsible for recruiting truncated proteins to ClpAP for degradation5,15. Fluorescence studies both with and without a ‘trap’ version of the chaperonin GroEL, which binds non-native forms of GFP16, and hydrogen-exchange experiments directly demonstrate that ClpA can unfold stable, native proteins in the presence of ATP.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kessel,M. et al. Homology in structural organization between E. coli ClpAP protease and the eukaryotic 26 S proteasome. J. Mol. Biol. 250, 587–594 (1995).
Beuron,F. et al. At sixes and sevens: characterization of the symmetry mismatch of the ClpAP chaperone-assisted protease. J. Struct. Biol. 123, 248–259 (1998).
Wickner,S. et al. A molecular chaperone, ClpA, functions like DnaK and DnaJ. Proc. Natl Acad. Sci. USA 91, 12218–12222 (1994).
Levchenko,I., Smith,C. K., Walsh,N. P., Sauer,R. T. & Baker,T. A. PDZ-like domains mediate binding specificity in the Clp/Hsp100 family of chaperones and protease regulatory subunits. Cell 91, 939–947 (1997).
Gottesman,S., Roche,E., Zhou,Y. N. & Sauer,R. T. The ClpXP and ClpAP proteases degrade proteins with carboxy-terminal peptide tails added by the SsrA-tagging system. Genes Dev. 12, 1338–1347 (1998).
DeMartino,G. N. et al. PA700, an ATP-dependent activator of the 20 S proteasome, is an ATPase containing multiple members of a nucleotide-binding protein family. J. Biol. Chem. 269, 20878–20884 (1994).
Glickman,M. H. et al. A subcomplex of the proteasome regulatory particle required for ubiquitin-conjugate degradation and related to the COP9-signalosome and eIF3. Cell 94, 615–623 (1998).
Baumeister,W., Walz,J., Zühl,F. & Seemüller,E. The proteasome: paradigm of a self-compartmentalizing protease. Cell 92, 367–380 (1998).
Pak,M. & Wickner,S. Mechanism of protein remodeling by ClpA chaperone. Proc. Natl Acad. Sci. USA 94, 4901–4906 (1997).
Levchenko,I., Luo,L. & Baker,T. A. Disassembly of the Mu transposase tetramer by the ClpX chaperone. Genes Dev. 9, 2399–2408 (1995).
Glover,J. R. & Lindquist,S. Hsp104, Hsp70, and Hsp40: a novel chaperone system that rescues previously aggregated proteins. Cell 94, 73–82 (1998).
Gottesman,S., Maurizi,M. R. & Wickner,S. Regulatory subunits of energy-dependent proteases. Cell 91, 435–438 (1997).
Larsen,C. N. & Finley,D. Protein translocation channels in the proteasome and other proteases. Cell 91, 431–434 (1997).
Wang,J., Hartling,J. A. & Flanagan,J. M. The structure of ClpP at 2.3 Å resolution suggests a model for ATP-dependent proteolysis. Cell 91, 447–456 (1997).
Keiler,K. C., Waller,P. R. H. & Sauer,R. T. Role of a peptide tagging system in degradation of proteins synthesized from damaged messenger RNA. Science 271, 990–993 (1996).
Weissman,J. S., Rye,H. S., Fenton,W. A., Beechem,J. M. & Horwich,A. L. Characterization of the active intermediate of a GroEL-GroES-mediated protein folding reaction. Cell 84, 481–490 (1996).
Miranker,A., Robinson,C. V., Radford,S. E. & Dobson,C. M. Investigation of protein folding by mass spectrometry. FASEB J. 10, 93–101 (1996).
Shtilerman,M., Lorimer,G. H. & Englander,S. W. Chaperonin function: folding by forced unfolding. Science 284, 822–825 (1999).
Maurizi,M. R., Clark,W. P., Kim, S.-H. & Gottesman,S. ClpP represents a unique family of serine proteases. J. Biol. Chem. 265, 12546–12552 (1990).
Fenton,W. A., Kashi,Y., Furtak,K. & Horwich,A. L. Residues in chaperonin GroEL required for polypeptide binding and release. Nature 371, 614–619 (1994).
Acknowledgements
E.U.W. is a postdoctoral associate of the Jane Coffin Childs Foundation. A.D.M. is a Pew Scholar in the biomedical sciences. This work was supported by the NIH, the Howard Hughes Medical Institute and the Nelson Fund.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Weber-Ban, E., Reid, B., Miranker, A. et al. Global unfolding of a substrate protein by the Hsp100 chaperone ClpA. Nature 401, 90–93 (1999). https://doi.org/10.1038/43481
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/43481
This article is cited by
-
ClpC2 protects mycobacteria against a natural antibiotic targeting ClpC1-dependent protein degradation
Communications Biology (2023)
-
Mapping SP-C co-chaperone binding sites reveals molecular consequences of disease-causing mutations on protein maturation
Nature Communications (2022)
-
Targeted substrate loop insertion by VCP/p97 during PP1 complex disassembly
Nature Structural & Molecular Biology (2021)
-
Time-resolved neutron scattering provides new insight into protein substrate processing by a AAA+ unfoldase
Scientific Reports (2017)
-
Small heat shock proteins sequester misfolding proteins in near-native conformation for cellular protection and efficient refolding
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.