Abstract
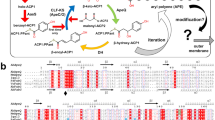
The biosynthesis of penicillin and cephalosporin antibiotics in microorganisms requires the formation of the bicyclic nucleus of penicillin1. Isopenicillin N synthase (IPNS), a non-haem iron-dependent oxidase, catalyses the reaction of a tripeptide, δ-(L-α-aminoadipoyl)- L-cysteinyl-D-valine (ACV), and dioxygen to form isopenicillin N and two water molecules2. Mechanistic studies suggest the reaction is initiated by ligation of the substrate thiolate to the iron centre, and proceeds through an enzyme-bound monocyclic intermediate3,4 (Fig. 1). Here we report the crystal structure of IPNS complexed to ferrous iron and ACV, determined to 1.3 å resolution. Based on the structure, we propose a mechanism for penicillin formation that involves ligation of ACV to the iron centre, creating a vacant iron coordination site into which dioxygen can bind. Subsequently, iron-dioxygen and iron-oxo species remove the requisite hydrogens from ACV without the direct assistance of protein residues (Fig. 2). The crystal structure of the complex with the dioxygen analogue, NO and ACV bound to the active-site iron supports this hypothesis.

AA, L-δ-(α-aminoadipoyl).

sp;IPNS complex.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Baldwin, J. E. & Abraham, E. Biosynthesis of penicillins and cephalosporins. Nat. Prod. Rep. 5, 129– 145 (1988).
Pang, C. P. et al. Purification of isopenicillin N synthetase. Biochem. J. 222, 789–795 ( 1984).
Baldwin, J. E. & Schofield, C. J. in The Chemistry of β-lactams (ed. Page, M. I.) 1–78 (Blackie, London, (1992)).
Que, L. & Ho, R. Y. N. Dioxygen activation by enzymes with mononuclear non-haem iron active sites. Chem. Rev. 96 , 2607–2624 (1996).
Orville, A. M. et al. Thiolate ligation of the active site iron(II) of isopenicillin N synthase derives from substrate rather than endogenous cysteine: spectroscopic studies of site-specific Cys → Ser mutated enzymes. Biochemistry 31, 4602–4612 ( 1992).
Randall, C. R. et al. X-ray absorption studies of the ferrous active site of isopenicillin N synthase and related model complexes. Biochemistry 32, 6664–6673 (1993).
Roach, P. L. et al. Crystal structure of isopenicillin N synthase is the first from a new structural family of enzymes. Nature 375, 700–704 (1995).
Roach, P. L. et al. Anaerobic crystallisation of an isopenicillin N synthase.Fe(II).substrate complex demonstrated by X-ray studies. Eur. J. Biochem. 242, 736–740 (1996).
Borovok, I., Landman, O., Kreisberg-Zakarin, R., Aharonowitz, Y. & Cohen, G. Ferrous active site of isopenicillin N synthase: genetic and sequence analysis of endogenous ligands. Biochemistry 35, 1981–1987 (1996).
Chen, V. J. et al. Spectroscopic studies of isopenicillin N synthase. A mononuclear nonhaem Fe2+oxidase with metal coordination sites for small molecules and substrate. J. Biol. Chem. 264, 21677–21681 (1989).
Baldwin, J. E. et al. Penicillin biosynthesis: active site mapping with aminoadipoylcysteinylvaline variants. J. Chem. Soc. Chem. Commun. 1225– 1227 (1984).
Rowe, C. J. thesis, Oxford Univ. (1995).
Hadfield, A. & Hajdu, J. Afast and portable microspectrophotometer for protein crystallography. J. Appl. Crystallogr. 26 , 839–842 (1993).
Cooper, R. D. G. The enzymes involved in biosynthesis of penicillin and cephalosporin: Their structure and function. Bioorg. Med. Chem. 1, 1–17 (1993).
Baldwin, J. E. et al. Evidence for an insertion-homolysis mechanism for carbon-sulfur bond formation in penicillin biosynthesis; 2. Incubation and interpretation. Tetrahedron 52, 2537–2556 (1996).
Groves, J. T. & Han, Y. Z. in Cytochrome P-450: Structure, Chemistry and Biochemistry 2nd edn (ed. Ortiz de Montellano, P. R.) 3– 49 (Plenum, New York, (1995)).
Baldwin, J. E., Morris, G. M. & Richards, W. G. Electron transport in cytochromes P-450 by covalent switching. Proc. R. Soc. Lond. B 245, 43– 52 (1991).
Prescott, A. G. Adilemma of dioxygenases (or where biochemistry and molecular biology fail to meet). J. Exp. Bot. 44, 849– 861 (1993).
Hanauske-Abel, H. M. & Guzzler, V. Astereochemical concept for the catalytic mechanism of prolylhydroxylase. J. Theor. Biol. 94, 421–455 ( 1982).
Shu, L. et al. X-ray absorption spectroscopic studies of the Fe(II) active site of catechol 2,3-dioxygenase. Implications for the extradiol cleavage mechanism. Biochemistry 34, 6649–6659 (1995).
Han, S., Eltis, L. D., Timmis, K. N., Muchmore, S. W. & Bolin, J. T. Crystal structure of the biphenyl-cleaving extradiol dioxygenase from a PCB-degrading pseudomonad. Science 270, 976–980 ( 1995).
Senda, T. et al. Three-dimensional structures of free form and two substrate complexes of an extradiol ring-cleavage type dioxygenase, the BphC enzyme from Pseudomonas sp. strain KKS102. J. Mol. Biol. 255, 735–752 (1996).
Otwinowski, Z. in Data Collection and Processing (eds Sawyer, L., Isaacs, N. W. & Bailey, S.) 55–62 (Daresbury Laboratory, Warrington, UK, (1993)).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 ( 1994).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjelgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A. T., Kuriyan, J. & Karplus, M. Crystallographic R factor refinement by molecular dynamics. Science 235, 458–460 (1987).
Konnert, J. H. & Hendrickson, W. A. Arestrained-parameter thermal-factor refinement procedure. Acta Crystallogr. A 36, 344–350 (1980).
Sheldrick, G. M. SHELXL93, Program for Crystal Structure Refinement (Univ. Gottingen, Germany, (1993)).
Leslie, A. G. W. Mosflm (MRC Laboratory of Molecular Biology, Cambridge, ( 1996)).
CCP4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Acknowledgements
We thank K. Harlos, E. Garman, R. Bryan, I. Andersson, R. M. Adlington, R. C. Wilmouth, V. Fülöp, J. P. N. Pitt, A. Howe, S. Lee, J. W. Keeping, B. Rasmussen and A. Thompson for help and discussions. Financial support was provided by the MRC, BBSRC, EPSRC and Zeneca through a DTI link scheme.
Author information
Authors and Affiliations
Author notes
The crystallographic coordinates have been deposited in the Brookhaven Protein Data Bank (accession nos 1IPS, 2IPS and 3IPS) and will be released one year after publication.
Rights and permissions
About this article
Cite this article
Roach, P., Clifton, I., Hensgens, C. et al. Structure of isopenicillinN synthase complexed with substrate and the mechanism ofpenicillin formation. Nature 387, 827–830 (1997). https://doi.org/10.1038/42990
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/42990
This article is cited by
-
Cephalosporin resistance, tolerance, and approaches to improve their activities
The Journal of Antibiotics (2024)
-
Subcellular localization of fungal specialized metabolites
Fungal Biology and Biotechnology (2022)
-
Molecular insights into the unusually promiscuous and catalytically versatile Fe(II)/α-ketoglutarate-dependent oxygenase SptF
Nature Communications (2022)
-
Crystal structure and catalytic mechanism of the MbnBC holoenzyme required for methanobactin biosynthesis
Cell Research (2022)
-
Exploring the Molecular Basis of Substrate and Product Selectivities of Nocardicin Bifunctional Thioesterase
Interdisciplinary Sciences: Computational Life Sciences (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.