Abstract
The group of enzymes known as TOR (for 'target of rapamycin') kinases regulates cell growth and proliferation in response to nutrients and hormone-dependent mitogenic signals1,2. Here we show that TOR deficiency in the nematode Caenorhabditis elegans more than doubles its natural lifespan. This new function for TOR signalling in ageing control may represent a link between nutrition, metabolism and longevity.
Similar content being viewed by others
Main
In C. elegans, the absence of LET-363/TOR activity causes developmental arrest at the L3 larval stage3. We examined nematodes bred as let-363/CeTor genetic null mutants and nematodes that had been depleted of TOR by using RNA interference to block let-363 expression (termed let-363-(RNAi) worms), and found that these animals had a strikingly extended mean lifespan (Fig. 1a, squares and triangles, respectively). At 25.5 °C, the mean lifetime was 25 days in let-363 mutants compared with 10 days in wild-type animals. This is all the more intriguing in light of the fact that TOR-deficient worms existed as arrested L3 larvae. In comparison, L3 larval arrest induced by starvation persisted for only 14 days on average in wild-type animals (Fig. 1a, diamonds).
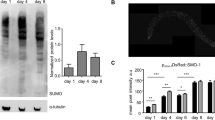
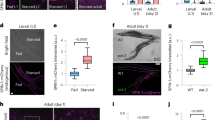
a, Lifespan of TOR-deficient worms compared with the wild type at 25.5 °C: wild type (filled circles); daf-16(mg50) (open circles); dpy-5(e61) unc-13(e450) double mutant (shaded circles); TOR-deficient triple mutants let-363(h114) dpy-5(e61) unc-13(e450) (open squares), let-363(h111) dpy-5(e61) unc-13(e450) (filled squares) and let-363(h131) dpy-5(e61) unc-13(e450) (shaded squares); let-363-RNAi-treated worms from hatching (open triangles) or from the first day of adulthood (shaded triangles); let-363-RNAi-treated daf-16(mg50) worms (filled triangles); starving-arrested wild-type L3 larvae (filled diamonds). Disruption of TOR by RNAi (triangles) seems to be incomplete, as lifespan is not extended as much as in let-363 mutants (squares). b, Nile Red staining of lipid droplets in a wild-type L3 larva (top) and an L3 larva arrested by let-363-(RNAi) treatment (bottom). Images were obtained with the same exposure time.
Strong inhibition of mitochondrial respiration also arrests development at the L3 stage, whereas weaker inhibition permits growth to adulthood and extends adult lifespan, but only if it occurs during larval development4. In contrast, treatment with let-363 double-stranded RNA starting from the first day of adulthood lengthens lifespan to a comparable extent when RNAi treatment is initiated at hatching (Fig. 1a, open triangles). This indicates that TOR has a role in ageing control during adulthood and that the long-lived phenotype of let-363-(RNAi) adults cannot be explained by reduced mitochondrial activity.
Longevity in C. elegans is controlled hormonally by a conserved signalling pathway that involves insulin and insulin-like growth factor (IGF)5,6. Mutants with reduced DAF-2/ IGF signalling activity live twice as long as the wild type5,6. The DAF-2/IGF cascade also acts during adulthood to influence ageing7. The remarkable similarity in the developmental stage at which ageing rate is affected, and our finding that the extended lifespan of daf-2(e1370) mutants is not increased further by treatment with let-363 RNAi (results not shown) — as it is with RNAi blocking expression of respiratory-chain components4 — raise the possibility that TOR and the DAF-2/IGF pathway are related in controlling lifespan.
This idea is compatible with results indicating that the insulin/IGF cascade regulates protein synthesis and cell growth in mammals and Drosophila through the activity of nutrient-sensing TOR (reviewed in refs 1, 2, 8). We have also noted that let-363-(RNAi) animals share certain features of the pleiotropic Daf-2(−) phenotype, such as lipid accumulation mainly in intestinal cells9 (Fig. 1b), as well as reduced fertility10 (mean brood sizes: let-363-(RNAi) adults, 68±6.4; wild type, 191±14.5) and reduced viability10 (embryonic/early larval arrest: let-363-(RNAi), 40.3%; wild type, 5.4%).
Strong mutations in DAF-2/IGF signalling cause a long-lived phenotype, together with a state of developmental diapause known as dauer that is triggered by starvation and crowding in the wild type11. According to our results (not shown), let-363(h111) animals bearing the thermosensitive daf-2(e1370ts) mutation were able to form dauers at the restrictive temperature. Furthermore, let-363-(RNAi) enhanced dauer formation in daf-2(e1370) animals. At 20 °C, only 4.6% (29 out of 630) of daf-2(e1370) mutants entered into the dauer stage, compared with 17.9% (146 out of 817) of daf-2(e1370); let-363-(RNAi) animals (results not shown). This indicates a genetic interaction between let-363/CeTor and daf-2. These results show that in C. elegans the TOR and DAF-2/IGF signalling pathways could be related in controlling ageing, metabolism and reproductive growth.
Lifespan extension in daf-2(e1370) mutants requires the activity of the forkhead transcription factor DAF-16 (refs 5, 6). Mutations in daf-16, however, do not suppress the long-lived phenotype of let-363-(RNAi) worms (Fig. 1a, filled triangles), indicating that TOR may be acting downstream or independently of DAF-16, and that it is interacting with the insulin endocrine system. Although the detailed signalling connections require clarification, our findings point to TOR as a possible mediator of lifespan regulation by insulin signalling and nutrient sensing.
References
Abraham, R. T. Cell 111, 9–12 (2002).
Oldham, S. & Hafen, E. Trends Cell Biol. 13, 79–85 (2003).
Long, X. et al. Curr. Biol. 12, 1448–1461 (2002).
Dillin, A. et al. Science 298, 2398–2401 (2002).
Kenyon, C. et al. Nature 366, 461–464 (1993).
Kenyon, C. in C. elegans II (eds Riddle, D. L., Blumenthal, T., Meyer, B. J. & Priess, J. R.) 791–814 (Cold Spring Harbor Lab. Press, Cold Spring Harbor, New York, 1997).
Dillin, A. et al. Science 298, 830–834 (2002).
Saltiel, A. R. & Kahn, C. R. Nature 414, 799–806 (2001).
Ashrafi, K. et al. Nature 421, 268–272 (2003).
Tissenbaum, H. A. & Ruvkun, G. Genetics 148, 703–717 (1998).
Riddle, D. L. & Albert, P. S. in C. elegans II (eds Riddle, D. L. et al.) 739–768 (Cold Spring Harbor Lab. Press, Cold Spring Harbor, New York, 1997).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Vellai, T., Takacs-Vellai, K., Zhang, Y. et al. Influence of TOR kinase on lifespan in C. elegans. Nature 426, 620 (2003). https://doi.org/10.1038/426620a
Issue Date:
DOI: https://doi.org/10.1038/426620a
This article is cited by
-
The rapid proximity labeling system PhastID identifies ATP6AP1 as an unconventional GEF for Rheb
Cell Research (2024)
-
mTOR Plays an Important Role in the Stemness of Human Fetal Cartilage Progenitor Cells (hFCPCs)
Tissue Engineering and Regenerative Medicine (2024)
-
Longevity and anti-aging effects of curcumin supplementation
GeroScience (2024)
-
Glycerol 3-phosphate phosphatase/PGPH-2 counters metabolic stress and promotes healthy aging via a glycogen sensing-AMPK-HLH-30-autophagy axis in C. elegans
Nature Communications (2023)
-
Pregnane X receptor agonist nomilin extends lifespan and healthspan in preclinical models through detoxification functions
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.