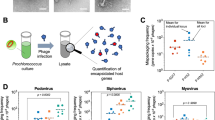
A bacteriophage may protect itself and its host against a deadly effect of bright sunlight.
Abstract
Cyanobacteria contribute to the overall photosynthetic production of oxygen in the oceans, but they are susceptible to infection by viruses and also to photo-inhibition when sunlight is too intense. Here we show that the genomic sequence of one such virus, a bacteriophage known as S-PM2, encodes the D1 and D2 proteins that are key components of one of the photosynthetic reaction centres (photosystem II, PSII), which are crucial sites of damage in photo-inhibition. The presence of this virus, and others like it, in the ocean may ensure that photo-inhibition is prevented in infected cells, allowing photosynthesis to continue and therefore provide the energy needed by the virus for its replication.
Similar content being viewed by others
Main
Primary production in the oligotrophic regions of the oceans is dominated by the cyanobacterial components of the picophytoplankton organisms Synechococcus and Prochlorococcus1. The amounts of visible and ultraviolet solar radiation in oceanic ecosystems may be sufficient, particularly in surface waters, to cause photo-inhibition2. The primary cause of photo-inhibition in chloroplasts and cyanobacteria is damage to PSII, a large protein–pigment complex that catalyses the light-dependent oxidation of water to molecular oxygen.
At the core of PSII is a dimer of two related proteins, D1 and D2, which binds the pigments and co-factors necessary for the complex's primary photochemistry. During photosynthesis, D1 and, to a lesser extent, D2 turn over rapidly as a result of light-induced damage and are replaced by newly synthesized polypeptides in a repair cycle. When the rate of photo-inactivation and damage of D1 exceeds the capacity for repair, photo-inhibition occurs, resulting in a reduction in the maximum efficiency of PSII photochemistry3.
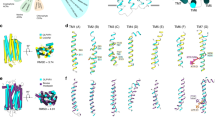
Viruses in general, and bacteriophages (viruses that infect bacteria) in particular, are abundant in marine ecosystems and are thought to exert major biogeochemical and ecological effects on the marine environment4. We analysed the genome sequence of S-PM2 (Fig. 1), a bacteriophage that infects marine Synechococcus strains5 and is about 194 kilobases long6, and found that it encodes the D1 and D2 proteins of PSII.
The genome contains a region of about 3.8 kilobases (GenBank accession no. AY329638) that extends from just over 100 base pairs upstream of the D1 gene (psbA), through two genes that are unrelated to photosynthesis, to a point some 100 base pairs downstream of the carboxy terminus of the D2 gene (psbD). The two intervening genes encode a homologue of the bacteriophage T4 gp49 (also known as recombination endonuclease VII) and a protein that has some similarity to a putative membrane protein present in the bacterium Escherichia coli.
The psbA gene appears to be interrupted, as the translated product of the gene aligns with other cyanobacterial and plant D1 proteins up to residue 276 (see supplementary information). There is then a region of 212 base pairs that encodes an amino-acid sequence without any similarity to known D1 proteins; this is followed by a region that encodes the remaining 25 amino acids of D1. This suggests that the psbA gene in S-PM2 contains a self-splicing intron. Introns occur in the psbA genes of the protozoan Chlamydamonas, and there is evidence for light/redox-regulated splicing of psbA precursor-messenger RNAs7. We detected copies of psbA genes after amplification by the polymerase chain reaction in five out of eight other Synechococcus viruses, although these genes seem to lack the putative intron.
The complete D1 protein of S-PM2 is similar to the D1 proteins of the marine Synechococcus sp. WH8102 (see supplementary information). There is homology in the DNA sequences, indicating that S-PM2 might have acquired the gene horizontally from its Synechococcus host. Presumably, psbD was acquired independently, given the presence of two unrelated intervening genes.
The expression of virus-encoded D1 and D2 proteins in infected cells would allow a repair cycle to operate in PSII after the host's protein synthesis had been shut down, thereby maintaining the cells' photosynthetic activity and the concomitant evolution of oxygen, and ensuring the provision of energy for the continued replication of the virus. This survival strategy resembles one used by a virus that infects the green alga Chlorella, which enhances the mechanism used by the host cell to rid itself of surplus light energy to avoid photo-inhibition8.
References
Partensky, F., Blanchot, J. & Vaulot, D. in Marine Cyanobacteria (eds Charpy, L. & Larkum, A. W. D.) Vol. 19, 457–475 (Institut Océanographique, Monaco, 1999).
Piazena, H., Perez-Rodrigues, E., Häder, D.-P. & Lopez-Figueroa, F. Deep Sea Res. II 49, 3513–3528 (2002).
Aro, E. M., Virgin, I. & Andersson, B. Biochim. Biophys. Acta 1143, 113–134 (1993).
Fuhrman, J. A. Nature 399, 541–548 (1999).
Wilson, W. H., Joint, I. R., Carr, N. G. & Mann, N. H. Appl. Environ. Microbiol. 59, 3736–3743 (1993).
Hambly, E. et al. Proc. Natl Acad. Sci. USA 98, 11411–11416 (2001).
Deshpande, N. N., Bao, Y. & Herrin, D. L. RNA 3, 37–48 (1997).
Seaton, G. G. R., Hurry, V. M. & Rohozinsky, J. FEBS Lett. 389, 319–323 (1996).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Mann, N., Cook, A., Millard, A. et al. Bacterial photosynthesis genes in a virus. Nature 424, 741 (2003). https://doi.org/10.1038/424741a
Issue Date:
DOI: https://doi.org/10.1038/424741a
This article is cited by
-
COBRA improves the completeness and contiguity of viral genomes assembled from metagenomes
Nature Microbiology (2024)
-
High-resolution metagenomic reconstruction of the freshwater spring bloom
Microbiome (2023)
-
Genomic and transcriptomic insights into complex virus–prokaryote interactions in marine biofilms
The ISME Journal (2023)
-
Lysogenic bacteriophages encoding arsenic resistance determinants promote bacterial community adaptation to arsenic toxicity
The ISME Journal (2023)
-
Heterogeneity of soil bacterial and bacteriophage communities in three rice agroecosystems and potential impacts of bacteriophage on nutrient cycling
Environmental Microbiome (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.