Abstract
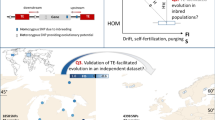
Population geneticists have long sought to estimate the distribution of selection intensities among genes of diverse function across the genome. Only recently have DNA sequencing and analytical techniques converged to make this possible. Important advances have come from comparing genetic variation within species (polymorphism) with fixed differences between species (divergence)1,2. These approaches have been used to examine individual genes for evidence of selection. Here we use the fact that the time since species divergence allows combination of data across genes. In a comparison of amino-acid replacements among species of the mustard weed Arabidopsis with those among species of the fruitfly Drosophila, we find evidence for predominantly beneficial gene substitutions in Drosophila but predominantly detrimental substitutions in Arabidopsis. We attribute this difference to the Arabidopsis mating system of partial self-fertilization, which corroborates a prediction of population genetics theory3,4,5,6 that species with a high frequency of inbreeding are less efficient in eliminating deleterious mutations owing to their reduced effective population size.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
McDonald, J. H. & Kreitman, M. Adaptive protein evolution at the Adh locus in Drosophila. Nature 351, 652–654 (1991).
Sawyer, S. A. & Hartl, D. L. Population genetics of polymorphism and divergence. Genetics 132, 1161–1176 (1992).
Charlesworth, D., Morgan, M. T. & Charlesworth, B. The effect of deleterious mutations on neutral molecular variation. J. Hered. 84, 321–325 (1993).
Kondrashov, A. S. Muller's ratchet under epistatic selection. Genetics 136, 1469–1473 (1994).
Caballero, A. & Santiago, E. Response to selection from new mutation and effective size of partially inbred populations. I. Theoretical results. Genet. Res. 66, 213–225 (1995).
Wang, J. L., Hill, W. G., Charlesworth, D. & Charlesworth, B. Dynamics of inbreeding depression due to deleterious mutations in small populations: Mutation parameters and inbreeding rate. Genet. Res. 74, 165–178 (1999).
Hartl, D. L., Moriyama, E. N. & Sawyer, S. A. Selection intensity for codon bias. Genetics 138, 227–234 (1994).
Akashi, H. Inferring weak selection from patterns of polymorphism and divergence at ‘silent’ sites in Drosophila DNA. Genetics 139, 1067–1076 (1995).
Gelman, A., Carlin, J. S., Stern, H. S. & Rubin, D. B. Bayesian Data Analysis (Chapman & Hall, London, 1997).
Carlin, B. P. & Louis, T. A. Bayes and Empirical Bayes Methods for Data Analysis (Chapman & Hall, London, 2000).
Gilks, R., Richardson, S. & Spiegelhalter, D. J. Markov Chain Monte Carlo in Practice (Chapman & Hall, London, 1996).
Metropolis, N., Rosenbluth, A. W., Rosenbluth, M. N., Teller, A. H. & Teller, E. Equations of state calculations by fast computing machines. J. Chem. Phys. 21, 1087–1091 (1953).
Geman, S. & Geman, D. Stochastic relaxation, Gibbs distributions and the Bayesian restoration of images. IEEE Trans. Pattern Anal. Machine Intell. 6, 721–741 (1984).
Gelman, A. & Rubin, D. B. Inference from iterative simulation using multiple sequences. Statist. Sci. 7, 457–511 (1992).
Abbott, R. J. & Gomes, M. F. Population genetic structure and the outcrossing rate of Arabidopsis thaliana. Heredity 62, 411–418 (1989).
Savolainen, O., Langley, C. H., Lazzaro, B. P. & Freville, H. Contrasting patterns of nucleotide polymorphism at the Adh locus in the outcrossing Arabidopsis lyrata and the selfing Arabidopsis thaliana. Mol. Biol. Evol. 17, 645–655 (2000).
Kusaba, M. et al. Self-incompatibility in the genus Arabidopsis: Characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana. Plant Cell 13, 627–643 (2001).
Caccone, A., Amato, G. D. & Powell, J. R. Rates and patterns of scnDNA and mtDNA divergence within the Drosophila melanogaster subgroup. Genetics 118, 671–683 (1988).
Acknowledgements
We thank D. Weinreich and D. Rand for providing the Drosophila data, and A. Kondrashov for numerous suggestions for improving the presentation. This work was supported by grants from the US Public Health Service, the US National Science Foundation, Howard Hughes and Marshall Sherfield Fellowships to C.D.B., and an Alfred P. Sloan Foundation Young Investigator Award to M.D.P.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests
Supplementary information
Supplementary information
Additional supplementary information accompanies this paper on the authors' website http://www.oeb.harvard.edu/hartl/lab. (TXT 5 kb)
Rights and permissions
About this article
Cite this article
Bustamante, C., Nielsen, R., Sawyer, S. et al. The cost of inbreeding in Arabidopsis. Nature 416, 531–534 (2002). https://doi.org/10.1038/416531a
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/416531a
This article is cited by
-
Molecular evolution of glutathione peroxidase (Gpx) family genes in desert poplar (Populus euphratica Oliv.)
Tree Genetics & Genomes (2017)
-
Geographically disjunct populations and widespread genets in an endangered halophilic plant, the Amargosa niterwort (Nitrophila mohavensis)
Conservation Genetics (2013)
-
Genomic consequences of transitions from cross- to self-fertilization on the efficacy of selection in three independently derived selfing plants
BMC Genomics (2012)
-
Rates of evolution in stress-related genes are associated with habitat preference in two Cardamine lineages
BMC Evolutionary Biology (2012)
-
Absence of Positive Selection on Centromeric Histones in Tetrahymena Suggests Unsuppressed Centromere-Drive in Lineages Lacking Male Meiosis
Journal of Molecular Evolution (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.