Abstract
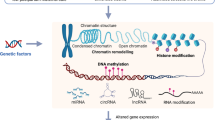
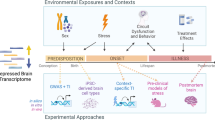
Major depressive disorder (MDD) is a common and highly heterogeneous psychiatric disorder encompassing a spectrum of symptoms involving deficits to a range of cognitive, psychomotor and emotional processes. As is the norm for aetiological studies into the majority of psychiatric phenotypes, particular focus has fallen on the interplay between genetic and environmental factors. There are, however, several epidemiological, clinical and molecular peculiarities associated with MDD that are hard to explain using traditional gene- and environment-based approaches. Our goal in this study is to demonstrate the benefits of looking beyond conventional ‘DNA+environment’ and ‘DNA × environment’ aetiological paradigms. Epigenetic factors – inherited and acquired modifications of DNA and histones that regulate various genomic functions occurring without a change in nuclear DNA sequence – offer new insights about many of the non-Mendelian features of major depression, and provide a direct mechanistic route via which the environment can interact with the genome. The study of epigenetics, especially in complex diseases, is a relatively new field of research, and optimal laboratory techniques and analysis methods are still being developed. Incorporating epigenetic research into aetiological studies of MDD thus presents a number of methodological and interpretive challenges that need to be addressed. Despite these difficulties, the study of DNA methylation and histone modifications has the potential to transform our understanding about the molecular aetiology of complex diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 4th edn. American Psychiatric Association: Washington, DC, 1994.
Murray CJ, Lopez AD . Global mortality, disability, and the contribution of risk factors: global burden of disease study. Lancet 1997; 349: 1436–1442.
Kessler RC, Berglund P, Demler O, Jin R, Koretz D, Merikangas KR et al. The epidemiology of major depressive disorder: results from the national comorbidity survey replication (NCS-R). JAMA 2003; 289: 3095–3105.
Winokur G, Pitts Jr FN . Affective disorder: VI. A family history study of prevalences, sex differences and possible genetic factors. J Psychiatr Res 1965; 3: 113–123.
Stenstedt A . Genetics of neurotic depression. Acta Psychiatr Scand 1966; 42: 392–409.
Reich T, Clayton PJ, Winokur G . Family history studies: V. The genetics of mania. Am J Psychiatry 1969; 125: 1358–1369.
Gershon ES, Dunner DL, Goodwin FK . Toward a biology of affective disorders. Genetic Contributions. Arch Gen Psychiatry 1971; 25: 1–15.
Murphy DL, Wyatt RJ . Neurotransmitter-related enzymes in the major psychiatric disorders: I. Catechol-O-Methyl transferase, monoamine oxidase in the affective disorders, and factors affecting some behaviorally correlated enzyme activities. Res Publ Assoc Res Nerv Ment Dis 1975; 54: 277–288.
Caspi A, Sugden K, Moffitt TE, Taylor A, Craig IW, Harrington H et al. Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science 2003; 301: 386–389.
Sullivan PF, Neale MC, Kendler KS . Genetic epidemiology of major depression: review and meta-analysis. Am J Psychiatry 2000; 157: 1552–1562.
Wender PH, Kety SS, Rosenthal D, Schulsinger F, Ortmann J, Lunde I . Psychiatric disorders in the biological and adoptive families of adopted individuals with affective disorders. Arch Gen Psychiatry 1986; 43: 923–929.
Bierut LJ, Heath AC, Bucholz KK, Dinwiddie SH, Madden PA, Statham DJ et al. Major depressive disorder in a community-based twin sample: are there different genetic and environmental contributions for men and women? Arch Gen Psychiatry 1999; 56: 557–563.
Kendler KS, Gardner CO, Neale MC, Prescott CA . Genetic risk factors for major depression in men and women: similar or different heritabilities and same or partly distinct genes? Psychol Med 2001; 31: 605–616.
Rush AJ, Zimmerman M, Wisniewski SR, Fava M, Hollon SD, Warden D et al. Comorbid psychiatric disorders in depressed outpatients: demographic and clinical features. J Affect Disord 2005; 87: 43–55.
Camp NJ, Cannon-Albright LA . Dissecting the genetic aetiology of major depressive disorder using linkage analysis. Trends Mol Med 2005; 11: 138–144.
Abkevich V, Camp NJ, Hensel CH, Neff CD, Russell DL, Hughes DC et al. Predisposition locus for major depression at chromosome 12q22–12q23.2. Am J Hum Genet 2003; 73: 1271–1281.
Arango V, Underwood MD, Mann JJ . Serotonin brain circuits involved in major depression and suicide. Prog Brain Res 2002; 136: 443–453.
Lotrich FE, Pollock BG . Meta-analysis of serotonin transporter polymorphisms and affective disorders. Psychiatr Genet 2004; 14: 121–129.
Lesch KP, Bengel D, Heils A, Sabol SZ, Greenberg BD, Petri S et al. Association of anxiety-related traits with a polymorphism in the serotonin transporter gene regulatory region. Science 1996; 274: 1527–1531.
Lim JE, Papp A, Pinsonneault J, Sadee W, Saffen D . Allelic expression of serotonin transporter (SERT) MRNA in human pons: lack of correlation with the polymorphism SERTLPR. Mol Psychiatry 2006; 11: 649–662.
Gizatullin R, Zaboli G, Jonsson EG, Asberg M, Leopardi R . Haplotype analysis reveals tryptophan hydroxylase (TPH) 1 gene variants associated with major depression. Biol Psychiatry 2006; 59: 295–300.
Schumacher J, Jamra RA, Becker T, Ohlraun S, Klopp N, Binder EB et al. Evidence for a relationship between genetic variants at the brain-derived neurotrophic factor (bdnf) locus and major depression. Biol Psychiatry 2005; 58: 307–314.
Strauss J, Barr CL, George CJ, Devlin B, Vetro A, Kiss E et al. Brain-derived neurotrophic factor variants are associated with childhood-onset mood disorder: confirmation in a hungarian sample. Mol Psychiatry 2005; 10: 861–867.
Massat I, Souery D, Del Favero J, Nothen M, Blackwood D, Muir W et al. Association between COMT (Val158Met) functional polymorphism and early onset in patients with major depressive disorder in a European multicenter genetic association study. Mol Psychiatry 2005; 10: 598–605.
Papadimitriou GN, Dikeos DG, Souery D, Del Favero J, Massat I, Avramopoulos D et al. Genetic association between the phospholipase A2 gene and unipolar affective disorder: a multicentre case–control study. Psychiatr Genet 2003; 13: 211–220.
van Rossum EF, Binder EB, Majer M, Koper JW, Ising M, Modell S et al. Polymorphisms of the glucocorticoid receptor gene and major depression. Biol Psychiatry 2006; 59: 681–688.
Arias B, Catalan R, Gasto C, Gutierrez B, Fananas L . Evidence for a combined genetic effect of the 5-HT(1A) receptor and serotonin transporter genes in the clinical outcome of major depressive patients treated with citalopram. J Psychopharmacol 2005; 19: 166–172.
Levinson DF . The genetics of depression: a review. Biol Psychiatry 2006; 60: 84–92.
Bruce ML . Psychosocial risk factors for depressive disorders in late life. Biol Psychiatry 2002; 52: 175–184.
Kendler KS, Prescott CA . A population-based twin study of lifetime major depression in men and women. Arch Gen Psychiatry 1999; 56: 39–44.
Turkheimer E, Waldron M . Nonshared environment: a theoretical, methodological, and quantitative review. Psychol Bull 2000; 126: 78–108.
Bouchard Jr TJ, Lykken DT, McGue M, Segal NL, Tellegen A . Sources of human psychological differences: the minnesota study of twins reared apart. Science 1990; 250: 223–228.
Bouchard Jr TJ, McGue M . Genetic and environmental influences on human psychological differences. J Neurobiol 2003; 54: 4–45.
Gottesman II . Schizophrenia Genesis. Freeman: New York, 1991.
Plomin R, Lichtenstein P, Pedersen NL, McClearn GE, Nesselroade JR . Genetic influence on life events during the last half of the life span. Psychol Aging 1990; 5: 25–30.
Kendler KS, Neale M, Kessler R, Heath A, Eaves L . A twin study of recent life events and difficulties. Arch Gen Psychiatry 1993; 50: 789–796.
Kendler KS, Karkowski-Shuman L . Stressful life events and genetic liability to major depression: genetic control of exposure to the environment? Psychol Med 1997; 27: 539–547.
Paykel E, Abbott R, Jenkins R, Brugha T, Meltzer H . Urban–rural mental health differences in great britain: findings from the national morbidity survey. Int Rev Psychiatry 2003; 15: 97–107.
Wang JL . Rural-urban differences in the prevalence of major depression and associated impairment. Soc Psychiatry Psychiatr Epidemiol 2004; 39: 19–25.
Whitfield JB, Zhu G, Heath AC, Martin NG . Choice of residential location: chance, family influences, or genes? Twin Res Hum Genet 2005; 8: 22–26.
Willemsen G, Posthuma D, Boomsma DI . Environmental factors determine where the Dutch live: results from the Netherlands twin register. Twin Res Hum Genet 2005; 8: 312–317.
Kendler KS, Kuhn JW, Vittum J, Prescott CA, Riley B . The interaction of stressful life events and a serotonin transporter polymorphism in the prediction of episodes of major depression: a replication. Arch Gen Psychiatry 2005; 62: 529–535.
Wilhelm K, Mitchell PB, Niven H, Finch A, Wedgwood L, Scimone A et al. Life events, first depression onset and the serotonin transporter gene. Br J Psychiatry 2006; 188: 210–215.
Moffitt TE, Caspi A, Rutter M . Strategy for investigating interactions between measured genes and measured environments. Arch Gen Psychiatry 2005; 62: 473–481.
Henikoff S, Matzke MA . Exploring and explaining epigenetic effects. Trends Genet 1997; 13: 293–295.
Jaenisch R, Bird A . Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 2003; 33: 245–254.
Jenuwein T, Allis CD . Translating the histone code. Science 2001; 293: 1074–1080.
Jones PL, Veenstra GJ, Wade PA, Vermaak D, Kass SU, Landsberger N et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat Genet 1998; 19: 187–191.
Robertson KD, Wolffe AP . DNA methylation in health and disease. Nat Rev Genet 2000; 1: 11–19.
Riggs AD, Xiong Z, Wang L, LeBon JM . Methylation dynamics, epigenetic fidelity and X chromosome structure. Novartis Found Symp. 1998; 214: 214–225.
Ushijima T, Watanabe N, Okochi E, Kaneda A, Sugimura T, Miyamoto K . Fidelity of the methylation pattern and its variation in the genome. Genome Res 2003; 13: 868–874.
Rakyan VK, Preis J, Morgan HD, Whitelaw E . The marks, mechanisms and memory of epigenetic states in mammals. Biochem J 2001; 356: 1–10.
Richards EJ . Inherited epigenetic variation – revisiting soft inheritance. Nat Rev Genet 2006; 7: 395–401.
Nicholls RD, Knepper JL . Genome organization, function, and imprinting in Prader–Willi and Angelman syndromes. Annu Rev Genomics Hum Genet 2001; 2: 153–175.
Xu GL, Bestor TH, Bourc'his D, Hsieh CL, Tommerup N, Bugge M et al. chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature 1999; 402: 187–191.
Amir RE, Zoghbi HY . Rett Syndrome: Methyl-CpG-binding protein 2 mutations and phenotype–genotype correlations. Am J Med Genet 2000; 97: 147–152.
Gibbons RJ, McDowell TL, Raman S, O'Rourke DM, Garrick D, Ayyub H et al. Mutations in ATRX, encoding a SWI/SNF-like protein, cause diverse changes in the pattern of DNA methylation. Nat Genet 2000; 24: 368–371.
Tapscott SJ, Klesert TR, Widrow RJ, Stoger R, Laird CD . Fragile-X syndrome and myotonic dystrophy: parallels and paradoxes. Curr Opin Genet Dev 1998; 8: 245–253.
Jones PA, Baylin SB . The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002; 3: 415–428.
Wong AH, Gottesman II, Petronis A . Phenotypic differences in genetically identical organisms: the epigenetic perspective. Hum Mol Genet 2005; 14: R11–R18.
Kato T, Iwamoto K, Kakiuchi C, Kuratomi G, Okazaki Y . Genetic or epigenetic difference causing discordance between monozygotic twins as a clue to molecular basis of mental disorders. Mol Psychiatry 2005; 10: 622–630.
Fraga MF, Ballestar E, Paz MF, Ropero S, Setien F, Ballestar ML et al. Epigenetic differences arise during the lifetime of monozygotic twins. Proc Natl Acad Sci USA 2005; 102: 10604–10609.
Petronis A, Gottesman II, Kan P, Kennedy JL, Basile VS, Paterson AD et al. Monozygotic twins exhibit numerous epigenetic differences: clues to twin discordance? Schizophr Bull 2003; 29: 169–178.
Mill J, Dempster E, Caspi A, Williams B, Moffitt T, Craig I . Evidence for monozygotic twin (MZ) discordance in methylation level at two CpG Sites in the promoter region of the Catechol-O-Methyltransferase (COMT) gene. Am J Med Genet B Neuropsychiatr Genet 2006; 141: 421–425.
Rakyan VK, Blewitt ME, Druker R, Preis JI, Whitelaw E . Metastable epialleles in mammals. Trends Genet 2002; 18: 348–351.
Petronis A . The origin of schizophrenia: genetic thesis, epigenetic antithesis, and resolving synthesis. Biol Psychiatry 2004; 55: 965–970.
Sutherland JE, Costa M . Epigenetics and the environment. Ann N Y Acad Sci 2003; 983: 151–160.
Waterland RA, Jirtle RL . Early nutrition, epigenetic changes at transposons and imprinted genes, and enhanced susceptibility to adult chronic diseases. Nutrition 2004; 20: 63–68.
Cooney CA, Dave AA, Wolff GL . Maternal methyl supplements in mice affect epigenetic variation and DNA methylation of offspring. J Nutr 2002; 132: 2393S–2400S.
Numachi Y, Yoshida S, Yamashita M, Fujiyama K, Naka M, Matsuoka H et al. Psychostimulant alters expression of DNA methyltransferase MRNA in the rat brain. Ann N Y Acad Sci 2004; 1025: 102–109.
Weaver IC, Cervoni N, Champagne FA, D'Alessio AC, Sharma S, Seckl JR et al. Epigenetic programming by maternal behavior. Nat Neurosci 2004; 7: 847–854.
Kendler KS, Gatz M, Gardner CO, Pedersen NL . A Swedish national twin study of lifetime major depression. Am J Psychiatry 2006; 163: 109–114.
Vaillant GE, Batalden M, Orav J, Roston D, Barrett JE . Evidence for a possibly X-linked trait related to affective illness. Aust N Z J Psychiatry 2005; 39: 730–735.
Thomson PA, Wray NR, Thomson AM, Dunbar DR, Grassie MA, Condie A et al. Sex-specific association between bipolar affective disorder in women and GPR50, an X-linked orphan G protein-coupled receptor. Mol Psychiatry 2005; 10: 470–478.
Faraone SV, Lyons MJ, Tsuang MT . Sex differences in affective disorder: genetic transmission. Genet Epidemiol 1987; 4: 331–343.
Craig IW, Harper E, Loat CS . The genetic basis for sex differences in human behaviour: role of the sex chromosomes. Ann Hum Genet 2004; 68: 269–284.
Plenge RM, Stevenson RA, Lubs HA, Schwartz CE, Willard HF . Skewed X-chromosome inactivation is a common feature of X-linked mental retardation disorders. Am J Hum Genet 2002; 71: 168–173.
Willard HF . The sex chromosomes and X chromosome inactivation. In: Scriver CR, Beaudet AL, Sly WS, Valle D, Childs B, Vogelstein B (eds). The Metabolic and Molecular Bases of Inherited Disease. McGraw-Hill: New York, 2000, pp 1191–1221.
Belmont JW . Genetic control of X inactivation and processes leading to X-inactivation skewing. Am J Hum Genet 1996; 58: 1101–1108.
Loat CS, Asbury K, Galsworthy MJ, Plomin R, Craig IW . X inactivation as a source of behavioural differences in monozygotic female twins. Twin Res 2004; 7: 54–61.
Carrel L, Willard HF . X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 2005; 434: 400–404.
Zubenko GS, Maher B, Hughes III HB, Zubenko WN, Stiffler JS, Kaplan BB et al. L. genome-wide linkage survey for genetic loci that influence the development of depressive disorders in families with recurrent, early onset, major depression. Am J Med Genet B Neuropsychiatr Genet 2003; 123: 1–18.
Csordas A, Puschendorf B, Grunicke H . Increased acetylation of histones at an early stage of oestradiol-mediated gene activation in the liver of immature chicks. J Steroid Biochem 1986; 24: 437–442.
Pasqualini JR, Mercat P, Giambiagi N . Histone acetylation decreased by estradiol in the MCF-7 human mammary cancer cell line. Breast Cancer Res Treat 1989; 14: 101–105.
Yokomori N, Moore R, Negishi M . Sexually dimorphic DNA demethylation in the promoter of the Slp (sex-limited protein) gene in mouse liver. Proc Natl Acad Sci USA 1995; 92: 1302–1306.
Saluz HP, Jiricny J, Jost JP . Genomic Sequencing reveals a positive correlation between the kinetics of strand-specific DNA Demethylation of the overlapping estradiol/glucocorticoid-receptor binding sites and the rate of avian vitellogenin MRNA synthesis. Proc Natl Acad Sci USA 1986; 83: 7167–7171.
McMahon FJ, Stine OC, Meyers DA, Simpson SG, DePaulo JR . Patterns of maternal transmission in bipolar affective disorder. Am J Hum Genet 1995; 56: 1277–1286.
Zill P, Engel R, Baghai TC, Zwanzger P, Schule C, Minov C et al. Analysis of polymorphisms in the olfactory G-Protein Golf in major depression. Psychiatr Genet 2002; 12: 17–22.
Schiffer HH, Heinemann SF . Association of the human kainate receptor GluR7 Gene (GRIK3) with recurrent major depressive disorder. Am J Med Genet B Neuropsychiatr Genet 2006 [E-pub ahead of print].
Davies W, Isles AR, Wilkinson LS . Imprinted gene expression in the brain. Neurosci Biobehav Rev 2005; 29: 421–430.
Preis JI, Downes M, Oates NA, Rasko JE, Whitelaw E . Sensitive flow cytometric analysis reveals a novel type of parent-of-origin effect in the mouse genome. Curr Biol 2003; 13: 955–959.
Sakatani T, Wei M, Katoh M, Okita C, Wada D, Mitsuya K et al. Epigenetic heterogeneity at imprinted loci in normal populations. Biochem Biophys Res Commun 2001; 283: 1124–1130.
Lewis SJ, Lawlor DA, Davey Smith G, Araya R, Timpson N, Day IN et al. The thermolabile variant of MTHFR is associated with depression in the British women's heart and health study and a meta-analysis. Mol Psychiatry 2006; 11: 352–360.
Zogel C, Bohringer S, Gross S, Varon R, Buiting K, Horsthemke B . Identification of Cis- and trans-acting factors possibly modifying the risk of epimutations on chromosome 15. Eur J Hum Genet 2006; 14: 752–758.
Polesskaya OO, Aston C, Sokolov BP . Allele C-specific methylation of the 5-HT2A receptor gene: evidence for correlation with its expression and expression of DNA methylase DNMT1. J Neurosci Res 2006; 83: 362–373.
Dempster EL, Mill J, Craig IW, Collier DA . The quantification of COMT MRNA in post mortem cerebellum tissue: diagnosis, genotype, methylation and expression. BMC Med Genet 2006; 7: 10.
Flanagan JM, Popendikyte V, Pozdniakovaite N, Sobolev M, Assadzadeh A, Schumacher A et al. Intra- and interindividual epigenetic variation in human germ cells. Am J Hum Genet 2006; 79: 67–84.
Frommer M, McDonald LE, Millar DS, Collis CM, Watt F, Grigg GW et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc Natl Acad Sci USA 1992; 89: 1827–1831.
Paul CL, Clark SJ . Cytosine methylation: quantitation by automated genomic sequencing and GENESCAN analysis. Biotechniques 1996; 21: 126–133.
Tost J, Dunker J, Gut IG . Analysis and quantification of multiple methylation variable positions in CpG islands by pyrosequencing. Biotechniques 2003; 35: 152–156.
Mill J, Yazdanpanah S, Gukel E, Ziegler S, Kaminsky Z, Petronis A . Whole genome amplification of sodium bisulfite-treated DNA allows the accurate estimate of methylated cytosine density in limited DNA resources. Biotechniques 2006; 41: 603–607.
Yatabe Y, Tavare S, Shibata D . Investigating stem cells in human colon by using methylation patterns. Proc Natl Acad Sci USA 2001; 98: 10839–10844.
Schumacher A, Kapranov P, Kaminsky Z, Flanagan J, Assadzadeh A, Yau P et al. Microarray-based DNA methylation profiling: technology and applications. Nucleic Acids Res 2006; 34: 528–542.
Zhou D, Qiao W, Wan Y, Lu Z . Microarray-based methylation analysis using dual-color fluorescence hybridization. J Biochem Biophys Methods 2006; 66: 33–43.
Mirnics K . Microarrays in brain research: the good, the bad and the ugly. Nat Rev Neurosci 2001; 2: 444–447.
O'Neill LP, VerMilyea MD, Turner BM . Epigenetic characterization of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nat Genet 2006; 38: 835–841.
Huang HS, Matevossian A, Jiang Y, Akbarian S . Chromatin immunoprecipitation in postmortem brain. J Neurosci Methods 2006; 156: 284–292.
Veldic M, Guidotti A, Maloku E, Davis JM, Costa E . In psychosis, cortical interneurons overexpress DNA-methyltransferase 1. Proc Natl Acad Sci USA 2005; 102: 2152–2157.
Bahn S, Augood SJ, Ryan M, Standaert DG, Starkey M, Emson PC . Gene expression profiling in the post-mortem human brain – no cause for dismay. J Chem Neuroanat 2001; 22: 79–94.
Cui H, Cruz-Correa M, Giardiello FM, Hutcheon DF, Kafonek DR, Brandenburg S et al. Loss of IGF2 imprinting: a potential marker of colorectal cancer risk. Science 2003; 299: 1753–1755.
Suter CM, Martin DI, Ward RL . Germline epimutation of MLH1 in individuals with multiple cancers. Nat Genet 2004; 36: 497–501.
Weksberg R, Shuman C, Caluseriu O, Smith AC, Fei YL, Nishikawa J et al. Discordant KCNQ1OT1 imprinting in sets of monozygotic twins discordant for beckwith-wiedemann syndrome. Hum Mol Genet 2002; 11: 1317–1325.
Eckhardt F, Beck S, Gut IG, Berlin K . Future potential of the human epigenome project. Expert Rev Mol Diagn 2004; 4: 609–618.
Rakyan VK, Hildmann T, Novik KL, Lewin J, Tost J, Cox AV et al. DNA methylation profiling of the human major histocompatibility complex: a pilot study for the human epigenome project. PLoS.Biol 2004; 2: e405.
Eckhardt F, Lewin J, Cortese R, Rakyan VK, Attwood J, Burger M et al. DNA methylation profiling of human chromosomes 6, 20 and 22. Nat Genet 2006; 38: 1378–1385.
Kalow W, Tang BK, Endrenyi L . Hypothesis: comparisons of inter- and intra-individual variations can substitute for twin studies in drug research. Pharmacogenetics 1998; 8: 283–289.
Acknowledgements
Research in the Krembil Family Epigenetics laboratory is supported by the National Institute of Mental Health (R01MH074127-01), the Canadian Institutes of Health Research (CIHR), and NARSAD. JM is supported by a CIHR postdoctoral fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mill, J., Petronis, A. Molecular studies of major depressive disorder: the epigenetic perspective. Mol Psychiatry 12, 799–814 (2007). https://doi.org/10.1038/sj.mp.4001992
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.mp.4001992
Keywords
This article is cited by
-
Early antidepressant treatment response prediction in major depression using clinical and TPH2 DNA methylation features based on machine learning approaches
BMC Psychiatry (2023)
-
Chronic mild stress paradigm as a rat model of depression: facts, artifacts, and future perspectives
Psychopharmacology (2022)
-
The Effect of Chronic Mild Stress and Venlafaxine on the Expression and Methylation Levels of Genes Involved in the Tryptophan Catabolites Pathway in the Blood and Brain Structures of Rats
Journal of Molecular Neuroscience (2020)
-
Genome-wide profiling of DNA methylome and transcriptome in peripheral blood monocytes for major depression: A Monozygotic Discordant Twin Study
Translational Psychiatry (2019)
-
Capsaicin upregulates HDAC2 via TRPV1 and impairs neuronal maturation in mice
Experimental & Molecular Medicine (2018)