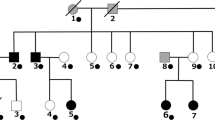
Abstract
We introduced a new genotyping method, fluorescence resonance energy transfer-based melting curve analysis on the LightCycler, for the analysis of the gene, DUSP6 (dual specificity MAP kinase phosphatase 6), in affective disorder patients. The DUSP6 gene is located on chromosome 12q22–23, which overlaps one of the reported bipolar disorder susceptibility loci. because of its role in intracellular signalling pathways, the gene may be involved in the pathogenesis of affective disorders not only on the basis of its position but also of its function. we performed association analysis using a t>G polymorphism that gives rise to a missense mutation (Leu114Val). No evidence for a significant disease-causing effect was found in Japanese unipolars (n = 132) and bipolars (n = 122), when compared with controls (n = 299). More importantly, this study demonstrates that melting curve analysis on the LightCycler is an accurate, rapid and robust method for discriminating genotypes from biallelic markers. This strategy has the potential for use in high throughput scanning for and genotyping of single nucleotide polymorphisms (SNPs).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Yoshikawa T, Padigaru M, Karkera JD, Sharma M, Berrettini W, Esterling LE et al. Genomic structure and novel variants of myo-inositol monophosphatase 2 (IMPA2) Mol Psychiatry 2000; 5: 165–171
Hinney A, Bornscheuer A, Depenbusch M, Mierke B, Tölle A, Middeke K et al. No evidence for involvement of the leptin gene in anorexia nervosa, bulimia nervosa, underweight or early onset extreme obesity: identification of two novel mutations in the coding sequence and a novel polymorphism in the leptin gene linked upstream region Mol Psychiatry 1998; 3: 539–543
Laruelle M, Gelernter J, Innis RB . D2 receptors binding potential is not affected by Taq1 polymorphism at the D2 receptor gene Mol Psychiatry 1998; 3: 261–265
Cavanagh G, Dunn AN, Chapman CE, Metcalfe P . HPA genotyping by PCR sequence-specific priming (PCR-SSP): a streamlined method for rapid routine investigations Transfus Med 1997; 7: 41–45
Hacia JG, Brody LC, Collins FS . Applications of DNA chips for genomic analysis Mol Psychiatry 1998; 3: 483–492
Smith A, Price C, Cullen M, Muda M, King A, Ozanne B et al. Chromosomal localization of three human dual specificity phosphatase genes (DUSP4, DUSP6, and DUSP7) Genomics 1997; 42: 524–527
Ewald H, Degn B, Mors O, Kruse TA . Significant linkage between bipolar affective disorder and chromosome 12q24 Psychiatr Genet 1998; 8: 131–140
Morissette J, Villeneuve A, Bordeleau L, Rochette D, Laberge C, Gagne B et al. Genome-wide search for linkage of bipolar affective disorders in a very large pedigree derived from a homogeneous population in Quebec points to a locus of major effect on chromosome 12q23–q24 Am J Med Genet 1999; 88: 567–587
Jacobsen NJO, Lyons I, Hoogendoorn B, Burge S, Kwok P-Y, O'Donovan MC et al. ATP2A2 mutations in Darier's disease and their relationships to neuropsychiatric phenotypes Hum Mol Genet 1999; 8: 1631–1636
Cowley S, Paterson H, Kemp P, Marshall CJ . Activation of MAP kinase kinase is necessary and sufficient for PC12 differentiation and for transformation of NIH 3T3 cells Cell 1994; 77: 841–852
Furukawa T, Yatsuoka T, Youssef EM, Abe T, Yokoyama T, Fukushige S et al. Genomic analysis of DUSP6, a dual specificity MAP kinase phosphatase, in pancreatic cancer Cytogenet Cell Genet 1998; 82: 156–159
Aslanidis C, Schmitz G . High-speed apolipoprotein E genotyping and apolipoprotein B3500 mutation detection using real-time fluorescence PCR and melting curves Clin Chem 1999; 45: 1094–1097
Nauck MS, Gierens H, Nauck MA, Marz W, Wieland H . Rapid genotyping of human platelet antigen 1 (HPA-1) with fluorophore-labelled hybridization probes on the LightCycler Br J Haematol 1999; 105: 803–810
Sham PC, Curtis D . Monte-Carlo tests for association between disease and alleles at highly polymorphic loci Ann Hum Genet 1995; 59: 97–105
Craddock N, Owen M, Burge S, Kurian B, Thomas P, McGuffin P . Familial cosegregation of major affective disorder and Darier's disease (keratosis follicularis) Br J Psychiatry 1994; 164: 355–358
Wakem P, Ikeda S, Haake A, Polakowska R, Ewing N, Sarret Y et al. Localization of the Darier disease gene to a 2-cM portion of 12q23–24.1 J Invest Dermatol 1996; 106: 365–367
Sakuntabhai A, Ruiz-Perez V, Carter S, Jacobsen N, Burge S, Monk S et al. Mutations in ATP2A2, encoding a Ca2+ pump, cause Darier disease Nat Genet 1999; 21: 271–277
Dawson E, Parfitt E, Roberts Q, Daniels J, Lim L, Sham P et al. Linkage studies of bipolar disorder in the region of the Darier's disease gene on chromosome 12q23–24.1 Am J Med Genet 1995; 60: 94–102
Jacobsen NJO, Franks EKE, Owen MJ, Craddock NJ . Mutational analysis of phospholipase A2A: a positional candidate susceptibility gene for bipolar disorder Mol Psychiatry 1999; 4: 274–279
Kimura M, Furukawa T, Abe T, Yatsuoka T, Youssef EM, Yokoyama T et al. Identification of two common regions of allelic loss in chromosome arm 12q in human pancreatic cancer Cancer Res 1998; 58: 2456–2460
Passik SD, Breitbart WS . Depression in patients with pancreatic carcinoma. Diagnostic and treatment issues Cancer 1996; 78: 615–626
Kennedy JL, Bradwejn J, Koszycki D, King N, Crowe R, Vincent J et al. Investigation of cholecystokinin system genes in panic disorder Mol Psychiatry 1999; 4: 284–285
Skogen B, Bellissimo DB, Hessner MJ, Santoso S, Aster RH, Newman PJ et al. Rapid determination of platelet alloantigen genotypes by polymerase chain reaction using allele-specific primers Transfusion 1994; 34: 955–960
Wang DG, Fan JB, Siao CJ, Berno A, Young P, Sapolsky R et al. Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome Science 1998; 280: 1077–1082
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Toyota, T., Watanabe, A., Shibuya, H. et al. Association study on the DUSP6 gene, an affective disorder candidate gene on 12q23, performed by using fluorescence resonance energy transfer-based melting curve analysis on the LightCycler. Mol Psychiatry 5, 489–494 (2000). https://doi.org/10.1038/sj.mp.4000748
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.mp.4000748