Abstract
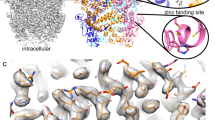
Despite close structural similarity, vardenafil (Levitra®) is 32-fold more potent than sildenafil (Viagra®) to inhibit cGMP-binding cGMP-specific PDE (PDE5); this is due to differences between their heterocyclic rings. In co-crystals with PDE5, one of the rings of vardenafil or sildenafil interacts with Tyr612, a catalytic site AA, via (1) a hydrogen bond with a water molecule and (2) hydrophobic bonds. For mutant PDE5Y612F, which ablates hydrogen-bonding potential, vardenafil or sildenafil inhibition was strengthened (2.2- or 3.0-fold, respectively), implying that the Tyr612 hydroxyl is a negative determinant for these inhibitors. For mutant PDE5Y612A, which ablates both hydrogen bonding and hydrophobic-bonding potential, vardenafil inhibition was weakened much more than sildenafil inhibition (122- and 26-fold, respectively), suggesting that hydrophobic bonds involving Tyr612 are stronger for vardenafil than for sildenafil.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 8 print issues and online access
$259.00 per year
only $32.38 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Abbreviations
- PDE:
-
3′,5′-cyclic nucleotide phosphodiesterase
- PDE5A:
-
cGMP-binding cGMP-specific PDE
- KPEM:
-
10 mM potassium phosphate
- pH 6.8:
-
1 mM EDTA, and 25 mM β-mercaptoethanol
- IBMX:
-
3-isobutyl-1-methylxanthine
- DTT:
-
DL-dithiothreitol
References
Hamet P, Coquil JF, Bousseau-Lafortune S, Franks DJ, Tremblay J . Cyclic GMP binding and phosphodiesterase: implication for platelet function. Adv Cyclic Nucleotide Protein Phosphorylation Res 1984; 16: 119–136.
Corbin JD, Francis SH . Cyclic GMP phosphodiesterase-5: target of sildenafil. J Biol Chem 1999; 274: 13729–13732.
Francis SH, Turko IV, Corbin JD . Cyclic nucleotide phosphodiesterases: relating structure and function. Prog Nucleic Acid Res Mol Biol 2001; 65: 1–52.
Corbin JD, Francis SH, Webb DJ . Phosphodiesterase type 5 as a pharmacologic target in erectile dysfunction. Urology 2002; 60: 4–11.
Rotella DP . Phosphodiesterase 5 inhibitors: current status and potential applications. Nat Rev Drug Discov 2002; 1: 674–682.
Montorsi F, Salonia A, Deho F, Cestari A, Guazzoni G, Rigatti P et al. Pharmacological management of erectile dysfunction. BJU Int 2003; 91: 446–454.
Hellstrom WJ, Gittelman M, Karlin G, Segerson T, Thibonnier M, Taylor T et al. Sustained efficacy and tolerability of vardenafil, a highly potent selective phosphodiesterase type 5 inhibitor, in men with erectile dysfunction: results of a randomized, double-blind, 26-week placebo-controlled pivotal trial. Urology 2003; 61: 8–14.
Lue TF . Erectile dysfunction. N Engl J Med 2000; 342: 1802–1813.
Porst H, Padma-Nathan H, Giuliano F, Anglin G, Varanese L, Rosen R . Efficacy of tadalafil for the treatment of erectile dysfunction at 24 and 36 hours after dosing: a randomized controlled trial. Urology 2003; 62: 121–125; discussion 125–126.
Goldstein I, Lue TF, Padma-Nathan H, Rosen RC, Steers WD, Wicker PA . Oral sildenafil in the treatment of erectile dysfunction. Sildenafil Study Group. N Engl J Med 1998; 338: 1397–1404.
Michelakis ED, Tymchak W, Noga M, Webster L, Wu XC, Lien D et al. Long-term treatment with oral sildenafil is safe and improves functional capacity and hemodynamics in patients with pulmonary arterial hypertension. Circulation 2003; 108: 2066–2069.
Ghofrani HA, Wiedemann R, Rose F, Schermuly RT, Olschewski H, Weissmann N et al. Sildenafil for treatment of lung fibrosis and pulmonary hypertension: a randomised controlled trial. Lancet 2002; 360: 895–900.
Kloner RA, Brown M, Prisant LM, Collins M . Effect of sildenafil in patients with erectile dysfunction taking antihypertensive therapy. Sildenafil Study Group. Am J Hypertens 2001; 14: 70–73.
Carson CC . Erectile dysfunction: evaluation and new treatment options. Psychosom Med 2004; 66: 664–671.
Boolell M, Allen MJ, Ballard SA, Gepi-Attee S, Muirhead GJ, Naylor AM et al. Sildenafil: an orally active type 5 cyclic GMP-specific phosphodiesterase inhibitor for the treatment of penile erectile dysfunction. Int J Impotence Res 1996; 8: 47–52.
Jeremy JY, Ballard SA, Naylor AM, Miller MA, Angelini GD . Effects of sildenafil, a type-5 cGMP phosphodiesterase inhibitor, and papaverine on cyclic GMP and cyclic AMP levels in the rabbit corpus cavernosum in vitro. Br J Urol 1997; 79: 958–963.
Saenz de Tejada I, Angulo J, Cuevas P, Fernandez A, Moncada I, Allona A et al. The phosphodiesterase inhibitory selectivity and the in vitro and in vivo potency of the new PDE5 inhibitor vardenafil. Int J Impot Res 2001; 13: 282–290.
Sung BJ, Yeon Hwang K, Ho Jeon Y, Lee JI, Heo YS, Hwan Kim J, Moon J et al. Structure of the catalytic domain of human phosphodiesterase 5 with bound drug molecules. Nature 2003; 425: 98–102.
Turko IV, Francis SH, Corbin JD . Potential roles of conserved amino acids in the catalytic domain of the cGMP-binding cGMP-specific phosphodiesterase. J Biol Chem 1998; 273: 6460–6466.
Turko IV, Ballard SA, Francis SH, Corbin JD . Inhibition of cyclic GMP-binding cyclic GMP-specific phosphodiesterase (Type 5) by sildenafil and related compounds. Mol Pharmacol 1999; 56: 124–130.
Francis SH, Sekhar KR, Rouse AB, Grimes KA, Corbin JD . Single step isolation of sildenafil from commercially available Viagra tablets. Int J Impot Res 2003; 15: 369–372.
Francis SH, Lincoln TM, Corbin JD . Characterization of a novel cGMP binding protein from rat lung. J Biol Chem 1980; 255: 620–626.
Wilkinson AJ, Fersht AR, Blow DM, Winter G . Site-directed mutagenesis as a probe of enzyme structure and catalysis: tyrosyl-tRNA synthetase cysteine-35 to glycine-35 mutation. Biochemistry 1983; 22: 3581–3586.
Andrews PR, Craik DJ, Martin JL . Functional group contributions to drug–receptor interactions. J Med Chem 1984; 27: 1648–1657.
Thomas MK, Francis SH, Corbin JD . Characterization of a purified bovine lung cGMP-binding cGMP phosphodiesterase. J Biol Chem 1990; 265: 14964–14970.
Turko IV, Haik TL, McAllister-Lucas LM, Burns F, Francis SH, Corbin JD . Identification of key amino acids in a conserved cGMP-binding site of cGMP-binding phosphodiesterases. A putative NKXnD motif for cGMP binding. J Biol Chem 1996; 271: 22240–22244.
Blount MA, Beasley A, Zoraghi R, Sekhar KR, Bessay EP, Francis SH et al. Binding of tritiated sildenafil, tadalafil, or vardenafil to the phosphodiesterase-5 catalytic site displays potency, specificity, heterogeneity, and cGMP stimulation. Mol Pharmacol 2004; 66: 144–152.
Corbin JD, Beasley A, Blount MA, Francis SH . Vardenafil: structural basis for higher potency over sildenafil in inhibiting cGMP-specific phosphodiesterase-5 (PDE5). Neurochem Int 2004; 45: 859–863.
Francis SH, Corbin JD . Molecular mechanisms and pharmacokinetics of phosphodiesterase-5 antagonists. Curr Urol Rep 2003; 4: 457–465.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Corbin, J., Francis, S. & Zoraghi, R. Tyrosine-612 in PDE5 contributes to higher affinity for vardenafil over sildenafil. Int J Impot Res 18, 251–257 (2006). https://doi.org/10.1038/sj.ijir.3901411
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ijir.3901411
Keywords
This article is cited by
-
Use of the KlADH3 promoter for the quantitative production of the murine PDE5A isoforms in the yeast Kluyveromyces lactis
Microbial Cell Factories (2017)