Abstract
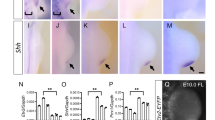
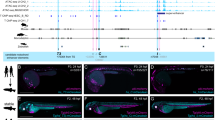
Hox genes control regional identity during segmentation of the vertebrate hindbrain into rhombomeres1–3. Here we use trans-genic analysis to investigate the upstream mechanisms for regulation ofHoxb-3 in rhombomere(r)5. We identified enhancers from the mouse and chick genes sufficient for r5-restricted expression. Sequence comparisons revealed two blocks of similarity (of 19 and 45 base pairs), which each containin vitro binding sites for the kreisler protein (Kmrl1), a Maf/b-Zip protein expressed in r5 and r6 (ref. 4). Both sites are required for r5 activity, suggesting that Hoxb-3 is a direct target ofkreisler. Multimers of the 19-base-pair (bp) block recreate a Krml1-like pattern in r5/r6, but the 45-bp block mediates expression only in r5. Therefore elements within the 45-bp block restrict the response to Krml1. We identified additional sequences that contain an Ets-related activation site, required for both the activation and restriction to r5. These studies demonstrate thatKrml1directly activates expression of Hoxb-3 in r5 in combination with an Ets-related activation site, and suggest that kreisler plays a primary role in regulating segmental identity through Hox genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
1. Lumsden, A. & Krumlauf, R. Patterning the vertebrate neuraxis. Science 274, 1109-1115 (1996). 2. Studer, M., Lumsden, A., Ariza-McNaughton, L., Bradley, A. & Krumlauf, R. Altered segmental identity and abnormal migration of motor neurons in mice lacking Hoxb-1. Nature 384, 630-634 (1996). 3. Goddard, J., Rossel, M., Manley, N. & Capecchi, M. Mice with targeted disruption of Hoxbl fail to form the motor nucleus of the VTIth nerve. Development 122, 3217-3228 (1996). 4. Cordes, S. P. & Barsh, G. S. The mouse segmentation gene kr encodes a novel basic domain-leucine zipper transcription factor. Cell 79, 1025-1034 (1994). 5. Hunt, P. et al A distinct Hox code for the branchial region of the head. Nature 253, 861-864 (1991). 6. Deol, M. The abnormalities of the inner ear in kreisler mice. /. Embryol. Exp. Morphol. 12, 475-490 (1964). 7. Frohman, M. A., Martin, G. R., Cordes, S., Halamek, L. P. & Barsh, G. S. Altered rhombomere-specific gene expression and hyoid bone differentiation in the mouse segmentation mutant kreiser (kr). Development 117, 925-936 (1993). 8. McKay, I. J. et al. The kreisler mouse: a hindbrain segmentation mutant that lacks two rhombomeres. Development 120, 2199-2211 (1994). 9. McKay, I. & Lumsden, A. Motor neuron defects in the kreisler mutant mouse. Eur. J. Neurosci. (in the press). 10. Swiatek, P. J. & Gridley, T. Perinatal lethality and defects in hindbrain development in mice homozygous for a targeted mutation of the zinc finger gene Krox-20. Genes Dev. 7,2071-2084 (1993). 11. Schneider-Maunoury, S. et al. Disruption of Krox-20 results in alteration of rhombomeres 3 and 5 in the developing hindbrain. Cell 75, 1199-1214 (1993). 12. Carpenter, E. M., Goddard, J. M., Chisaka, O., Manley, N. R. & Capecchi, M. R. Loss ofHoxa-1 (Hox-1.6) function results in the reorganization of the murine hindbrain. Development 118,1063-1075 (1993). 13. Dolle, P. et al. Local alterations of Krox-20 and Hox gene expression in the hindbrain of Hox-1 (Hox-1.6) homozygote null mutant embryos. Proc. Natl Acad. Sci. USA 90, 7666-7670 (1993). 14. Mark, M. et al. Two rhombomeres are altered in Hoxa-1 mutant mice. Development 119, 319-338 (1993). 15. Nonchev, S. et al. The conserved role of Krox-20 in directing Hox gene expression during vertebrate hindbrain segmentation. Proc. Natl Acad. Sci. USA 93, 9339-9345 (1996). 16. Kataoka, K., Noda, N. & Nishizawa, M. Maf nuclear oncoprotein recognizes sequences related to an AP-1 sites and forms heterodimers with both Fos and Jun. Mol. Cell. Biol. 14, 700-712 (1994). 17. Kataoka, K., Fujiwara, K. T., Noda, M. & Nishizawa, M. MafB, a new Maf family transcription activator that can associate with Maf and Fos but not with Jun. Mol. Cell. Biol. 14, 7581-7591 (1994). 18. Nye, J., Petersen, J., Gunther, C., Jonsen, M. & Graves, B. Interaction of murine Ets-1 with GGA-binding sites establishes the ETS domain as a new DNA binding motif. Genes Dev. 6,975-990 (1992). 19. Shore, P. et al. Determinants of DNA binding specificity of ETS-domain transcription factors. Mol. Cell. Biol. 16, 3338-3349 (1996). 20. Wasylyk, B. et al. The c-ets proto-oncogenes encode transcription factors that cooperate with c-Fos and c-Jun for transcriptional activation. Nature 346, 191-193 (1990). 21. Treier, M., Bohmann, D. & Mlozik, M. JUN cooperates with the Ets domain protein Pointed to induce photoreceptor R7 fate in the Drosophila eye. Cell 83, 753-776 (1995). 22. Sieweke, M., Tekotte, H., Frampton, J. & Graf, T. MafB is an interaction partner and repressor of Ets-1 that inhibits erythroid differentiation. Cell 85, 49-60 (1996). 23. Maroulakou, L, Papas, T. & Green, J. Differential expression of ets-1 and ets-2 proto-oncogenes during murine embryogenesis. Oncogene 9, 1551-1565 (1994). 24. Whiting, J. et al. Multiple spatially specific enhancers are required to reconstruct the pattern of Hox-2.6 gene expression. Genes Dev. 5, 2048-2059 (1991). 25. Yee, S.-P. & Rigby, P. W. }. The regulation of myogenin gene expression during the embryonic development of the mouse. Genes Dev. 7, 1277-1289 (1993). 26. Sham, M.-H. et al. Analysis of the murine Hox-2.7 gene: conserved alternative transcripts with differential distributions in the nervous system and the potential for shared regulatory regions. EMBO J. 11, 1825-1836 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Manzanares, M., Cordes, S., Kwan, CT. et al. Segmental regulation of Hoxb-3 by kreisler. Nature 387, 191–195 (1997). https://doi.org/10.1038/387191a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/387191a0
This article is cited by
-
The gene regulatory networks underlying formation of the auditory hindbrain
Cellular and Molecular Life Sciences (2015)
-
Development of oculomotor circuitry independent of hox3 genes
Nature Communications (2014)
-
A Hox regulatory network of hindbrain segmentation is conserved to the base of vertebrates
Nature (2014)
-
Evolution of anterior Hox regulatory elements among chordates
BMC Evolutionary Biology (2011)
-
Patterning and axon guidance of cranial motor neurons
Nature Reviews Neuroscience (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.