Abstract
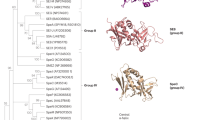
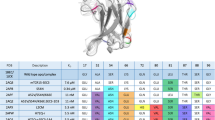
SUPERANTIGENS (SAgs) are viral or bacterial proteins that act as potent T-cell stimulants and have been implicated in a number of human diseases, including toxic shock syndrome1,2, diabetes mellitus3 and multiple sclerosis4. The interaction of SAgs with the T-cell receptor (TCR) and major histocompatibility complex (MHC) proteins results in the stimulation of a disproportionately large fraction of the T-cell population2. We report here the crystal structures of the β-chain of a TCR complexed with the Staphylococcus aureus enterotoxins C2 and C3 (SEC2, SEC3). These enterotoxins, which cause both toxic shock and food poisoning, bind in an identical way to the TCR β-chain. The complementarity-determining region 2 (CDR2) of the β-chain and, to lesser extents, CDR1 and hypervariable region 4 (HV4), bind in a cleft between the two domains of the SAgs. Thus, there is considerable overlap between the SAg-binding site and the peptide/MHC-binding sites of the TCR. A model of a TCR–SAg–MHC complex constructed from the crystal structures of (1) the β-chain–SEC3 complex, (2) a complex between staphylococcal enterotoxin B (SEB) and an MHC molecule5, and (3) a TCR Vα domain6, reveals that the SAg acts as a wedge between the TCR and MHC to displace the antigenic peptide away from the TCR combining site. In this way, the SAg is able to circumvent the normal mechanism for T-cell activation by specific peptide/MHC complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Bohach, G. A., Fast, D. J., Nelson, R. D. & Schlievert, P. M. Crit. Rev. Microbiol. 17, 251–272 (1990).
Kotzin, B. L., Leung, D. Y. M., Kappler, J. & Marrack, P. Adv. Immunol. 54, 99–166 (1993).
Conrad, B. et al. Nature 371, 351–355 (1994).
Brocke, S. et al. Nature 365, 642–644 (1993).
Jardetzky, T. S. et al. Nature 368, 711–718 (1994).
Fields, B. A. et al. Science 270, 1821–1824 (1995).
Bentley, G. A., Boulot, G., Karjalainen, K. & Mariuzza, R. A. Science 267, 1984–1987 (1995).
Bentley, G. A. & Mariuzza, R. A. Annu. Rev. Immunol. 14, 563–590 (1996).
Papageorgiou, A. C. et al. Structure 3, 769–779 (1995).
Hoffmann, M. L. et al. Infect. Immun. 62, 3396–3407 (1994).
Malchiodi, E. L. et al. J. Exp. Med. 182, 1833–1845 (1995).
Patten, P. A. et al. J. Immunol. 150, 2281–2294 (1993).
Webb, S. R. & Gascoigne, N. R. J. Curr. Opin. Immunol. 6, 467–475 (1994).
Swaminathan, S., Furey, W., Pletcher, J. & Sax, M. Nature 359, 801–806 (1992).
Deringer, J. R., Ely, R. J., Stauffacher, C. V. & Bohach, G. A. Mol. Microbiol. (in the press).
Fremont, D. H., Matsumura, M., Stura, E. A., Peterson, P. & Wilson, I. A. Science 257, 919–927 (1992).
Arden, B., Clark, S. P., Kabelitz, D. & Mak, T. W. Immunogenetics 42, 455–500 (1995).
Hong, S.-C., Waterbury, G., Kanner, E. M. & Janeway, C. A. J. Exp. Med. 183, 1437–1446 (1996).
Woodland, D. L. & Blackman, M. A. Immunol. Today 14, 208–212 (1993).
Fremont, D. H., Hendrickson, W. A., Marrack, P. & Kappler, J. Science 272, 1001–1004 (1996).
Labrecque, N., Thibodeau, J., Mourad, W. & Sekaly, R.-P. J. Exp. Med. 180, 1921–1929 (1994).
Deckhut, A. M., Chien, Y., Blackman, M. A. & Woodland, D. L. J. Exp. Med. 180, 1931–1935 (1994).
Seth, A. et al. Nature 369, 324–327 (1994).
Kim, J., Urban, R. G., Strominger, J. L. & Wiley, D. C. Science 266, 1870–1874 (1995).
Blomster-Hautamaa, D. A. & Schlievert, P. M. Methods Enzymol. 165, 37–42 (1988).
Otwinowski, Z. in Data Collection and Processing: Proc. CCP4 Study Weekend (eds Sawyer, L., Isaacs, N. & Bailey, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK, 1993).
Navaza, J. Acta Crystallogr. A 50, 157–163 (1994).
Brünger, A. T. X-PLOR Version 3.1. A System for X-ray Crystallography and NMR (Yale Univ. Press, New Haven, London, 1992).
Kleywegt, G. J. Acta Crystallogr. D 52, 842–857 (1996).
Furey, W. & Swaminathan, S. Methods Enzymol (in the press).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. J. Appl. Crystallogr. 26, 283–291 (1993).
Garcia, K. C. et al. Science 274, 209–219 (1996).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fields, B., Malchiodi, E., Li, H. et al. Crystal structure of a T-cell receptor β-chain complexed with a superantigen. Nature 384, 188–192 (1996). https://doi.org/10.1038/384188a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/384188a0
This article is cited by
-
Novel insights into the immune response to bacterial T cell superantigens
Nature Reviews Immunology (2024)
-
The γδTCR combines innate immunity with adaptive immunity by utilizing spatially distinct regions for agonist selection and antigen responsiveness
Nature Immunology (2018)
-
Two common structural motifs for TCR recognition by staphylococcal enterotoxins
Scientific Reports (2016)
-
SEC2-induced superantigen and antitumor activity is regulated through calcineurin
Applied Microbiology and Biotechnology (2013)
-
The structure of superantigen complexed with TCR and MHC reveals novel insights into superantigenic T cell activation
Nature Communications (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.