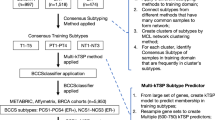
Abstract
Around 25% of hereditary breast and ovarian cancer families have mutations in the BRCA1 and BRCA2 genes. The search for other genes has until now failed, probably because there is not one single BRCAX gene, but rather various genes that may each be responsible for a small number of breast cancer families and/or may interact according to a polygenic model. We have studied 50 tumors from probands belonging to non-BRCA1/2 breast cancer families (BRCAX), using 25 immunohistochemical markers. The objective was to classify these tumors and confirm that they are heterogeneous. Unsupervised cluster analysis showed the existence of the following two main groups of tumors: high-grade and estrogen receptor (ER)-negative tumors (50%), and low-grade and ER-positive tumors (50%). In addition we identified five subgroups, three among the high-grade and two among the low-grade groups; one overexpressing HER-2 (18%); one with a basal-like phenotype (14%); one with a normal breast-like phenotype (18%); a luminal A subgroup (36%), and a luminal B subgroup (14%). Hypermethylation of the BRCA1 gene was observed in 42% of the cases, spread across all five subgroups, but only 37% of those had loss of heterozygosity as well. These latter cases were all clustered in the high-grade group and the majority of them in the basal-like subgroup. Our results show that familial non-BRCA1/2 tumors are heterogeneous and suggest a polygenic model for explaining the majority of BRCAX families. In addition we have defined a subset of them that have somatic inactivation of the BRCA1 gene.
Similar content being viewed by others
Main
More than a decade has passed since BRCA1 and BRCA2 were cloned1, 2 and their association with familial breast and ovarian cancer (FBOC) was established.3 However, recent data indicate that these two genes explain only 25% of these families.4, 5, 6
The large number of families without an identified causative gene mutation has led many a groups to pursue putative BRCAX gene(s) through different approaches, but without success. Several reports have been published since 1995 suggesting linkage of the BRCAX gene to specific chromosomal regions,7, 8 but these data have not been reproduced in larger series.9, 10, 11 Hedenfalk et al12 carried out a study using expression arrays in a small group of non-BRCA1/2 tumors, and concluded that they were heterogeneous and could be split into two main groups; but again, these results have not been reproduced. It has also been suggested that BRCAX families could be explained by a polygenic model, or that they might carry mutations in another gene or genes conferring a moderately increased risk of breast cancer.13, 14, 15
Using paraffin-embedded tissue, we and others have previously demonstrated that BRCA1 and BRCA2 tumors can be differentiated because they have a specific immunohistochemical profile.16, 17, 18, 19, 20 Based on these results, we hypothesized that the use of different immunohistochemical markers might help to group the non-BRCA1/2 tumors, and to confirm that they are heterogeneous. The confirmation of this heterogeneity and their classification would be very important for further studies searching for candidate genes.
Materials and methods
Patients
We collected paraffin-embedded tumor tissues from 50 individuals (mean age 47 years) from 50 different high-risk families, who were studied for mutations in the BRCA1 and BRCA2 genes. These individuals were from families with at least three members affected with breast and/or ovarian cancer, at least one of whom was younger than 50 when diagnosed.4 All 50 individuals were screened for mutations, including large deletions, in the BRCA1 and BRCA2 genes, and no mutations were detected.4, 21 The complete coding sequence and exon–intron boundaries of the BRCA1 and BRCA2 genes were analyzed by a combination of the following different techniques, depending on the center of origin: SSCP, PTT, CSGE, DGGE, and direct sequencing.21
We compared the profiles of non-BRCA1/2 tumors vs a group of 33 tumors from patients carrying a mutation in the BRCA1 gene, selected using the same criteria and studied with the same methodology as described above. Finally, we included a control group of 50 sporadic tumors that were selected because they were diagnosed at similar ages to the non-BRCA1/2 tumors (mean 49 years). In order to confirm that the non-BRCA1/2 and sporadic cases were genetically different, we estimated BRCA1 and BRCA2 carrier probabilities for both the groups using BRCAPRO.22 Data on the majority of the markers assessed in the present study have been previously reported for all three groups of tumors.18
Tissue Microarray Construction
The morphological subtype and grade was assessed in complete sections of each tumor stained with hematoxylin–eosin (H–E). The non-BRCA1/2 tumors consisted of 44 invasive ductal carcinomas, five in situ ductal carcinomas and one invasive lobular carcinoma.
Representative areas of the different lesions were carefully selected on H–E sections and marked on individual paraffin blocks. Two tissue cores (1 mm in diameter) were obtained from each specimen. In addition, four cores of normal breast tissue and two cores of tonsil were included as controls. The tissue cores were arrayed onto one independent new paraffin block using a tissue microarray (TMA) workstation (Beecher Instruments, Silver Spring, MD, USA). The final TMA consisted of 106 cores, each 1 mm in diameter, spaced 0.8 mm from each other. A section stained with H–E was studied to confirm the presence of morphologically representative areas of the original lesions. The BRCA1 and sporadic tumors were included in two separate TMAs using the same technique.
Immunohistochemical Studies
Immunohistochemical staining was performed by the Envision method (Dako, Glostrup, Denmark), with a heat-induced antigen retrieval step. Sections from the tissue array were immersed in 10 mM boiling sodium citrate at pH 6.5 for 2 min in a pressure cooker. Antibodies, dilutions and suppliers are listed in Table 1.
Between 150 and 200 cells per core were scored for the percentage of positive nuclei or cytoplasms, depending on the marker. We evaluated nuclear staining for estrogen receptor (ER); progesterone receptor (PR); p53; Ki-67; cyclins D1, D3, E, and A; p16; p27; p21; Skp2; retinoblastoma protein (Rb); E2F6; MDM2; topoisomerase IIα; survivin; and CHEK2, and evaluated cytoplasmic staining for BCL2, vimentin, cytokeratin 5/6 (CK5/6), cytokeratin 8 (CK8), and cyclin B1, as previously described.16, 17, 18 Only the percentage of stained cells was considered (independent of the intensity), and the positivity threshold for each marker is listed in Table 1. We and others have previously used the same threshold in the analysis of these markers.16, 18, 20, 23, 24, 25, 26, 27 HER-2 expression was evaluated according to the four-category (0–3+) DAKO system proposed for the evaluation of the HercepTest, and HER-2 expression of 3+ was the only value considered positive, as previously described.18, 19
Statistical Analysis
Hierarchical unsupervised cluster analysis was performed by means of the UPGMA (unweighted pair-group method using arithmetic averages) method using correlation distance and Euclidean distance between markers. The cluster was displayed using SOTAARRAY28 (software available at http://gepas.bioinfo.cipf.es/). The method was implemented in the GEPAS package.29 Immunohistochemical results were represented by a range of color from green to red, the most green representing the lowest, and the most red the highest percentage of positive cells for each marker. Exceptions were grades which were scaled as 33% ‘expressed’ for grade 1, 66% for grade 2, and 100% for grade 3 and HER-2, which was scaled as for 100% for positive (3+), and 0% for negative (Figure 1). We used the CAAT software based on Silhouette Width for clustering validation (software available at http://gepas.bioinfo.cipf.es/).
Hierarchical clustering of 50 non-BRCA1/2 tumors. White squares correspond to data not available. The percentage of positive cells for each immunohistochemical marker is represented as a range of color between the most green (lowest percentage) and the most red (highest percentage). Intermediate colors represent percentages between the lowest and the highest.
The χ2-test was performed to determine the differences in the distributions of the expression of each antibody and grade between the groups (Tables 2 and 3). The statistical program SPSS 13.0 for Windows (SPSS Inc., Chicago, IL, USA) was used for this analysis. Correction for multiple testing was made by a permutation method in which group membership was randomly assigned, conserving observed proportions, and the distribution of minimum P-values determined over 10 000 permutations. Differences in median BRCAPRO probabilities were tested using the Mann–Whitney rank-sum test.
BRCA1 Promoter Hypermethylation
DNA methylation patterns in the CpG islands of the promoter of the BRCA1 gene were determined by methylation-specific PCR in primary tumors after bisulfite treatment of DNA.30 Placental DNA treated in vitro with SssI bacterial methylase was used as a positive control, and DNA from normal lymphocytes was used as a negative control for methylated alleles of BRCA1.
BRCA1 Loss of Heterozygosity
Loss of Heterozygosity (LOH) analysis of the BRCA1 gene was performed using the intronic microsatellite markers D17S1322 and D17S855 that localize to introns 19 and 20, respectively, and D17S1327 that localizes in 17q21.31 outside the BRCA1 gene.31 The forward primer for each set was labeled using the fluorescent dye FAM (Applied Biosystems/PE Biosystems, Foster City, CA, USA). Reactions were cycled as follows: 95°C for 5 min, then 35 cycles at 94°C for 60 s, 55°C for 60 s, and 72°C for 90 s, followed by final elongation at 72°C for 5 min.
Allele sizes were determined using an automated capillary sequencer (ABI Prism™ 310; Applied Biosystems, Perkin Elmer, Warrington, UK) and were analyzed using GeneScan 3.1 software (Applied Biosystems, Warrington, UK). LOH was determined when the difference between peaks representing alleles in the tumor and the corresponding normal DNA reactions exceeded 25%.
Results
The morphological and immunohistochemical profiles of non-BRCA1/2 tumors were established by analyzing grade and 25 immunohistochemical markers in 50 such tumors, and comparing them with 50 sporadic tumors. Non-BRCA1/2 tumors were of lower grade (adjusted P=0.04); 54% were grade 1 vs 20% of sporadic tumors (Table 2). Although there was marginal evidence that p53 and p21 expression differed between the two groups, these associations disappeared after correction for multiple testing. Overall, the expression of markers related to proliferation, cell cycle, apoptosis, hormone receptors, and epithelial proteins showed similar patterns after adjustment for multiple testing (Table 2).
Although the phenotype of both groups was similar, they were genetically very different. Using BRCAPRO we found that the median probability of not being carrier of BRCA1 or BRCA2 mutations was 0.994 (range 0.964–0.997) for the sporadic tumors and 0.519 (range 0.062–0.987) for the non-BRCA1/2 tumors (P<0.0001).
Hierarchical Unsupervised Cluster Analysis
We performed a hierarchical unsupervised cluster analysis of the non-BRCA1/2 tumors based on the 25 immunohistochemical markers and grade, and found that the 50 tumor samples were separated into two main groups with 25 tumors each, differentiated primarily by grade and ER status. The high-grade branch (Figure 1, left) included tumors of grade 2 or 3 (brown and red squares, respectively) that were ER negative and had overexpression of proteins that promote cell cycle progression and proliferation. The low-grade branch (Figure 1, right) included grade 1 tumors (green squares) that were ER positive and showed overexpression of proteins related to the inhibition of the cyclin–CDK complexes, or the overexpression of luminal epithelial proteins such as CK8.
Within the high-grade branch we distinguished the following three different subgroups: one characterized by HER-2 overexpression (Figure 1, blue branch) that corresponded to 18% of all cases; a second basal-like subgroup containing 14% of the cases; and a third group that represents 18% of all cases, defined by low expression of the luminal epithelial marker CK8 and overexpression of other proteins associated with cell cycle progression and proliferation. Furthermore, this latter subgroup contained most of the ER-positive tumors in this predominantly ER-negative branch (Figure 1, green branch), and we have named it the ‘normal breast-like’ group because of its similarity to the group described by Sorlie et al32 with this name.
The low-grade branch could be divided into two further subgroups (Figure 1, pink and brown branches). The brown branch in Figure 1 was the largest (18 tumors) and demonstrated higher expression of ER, BCL2, CK8, and proteins that inhibit cell cycle progression. The second subgroup contained the remaining seven (14%) cases (Figure 1, pink branch) and showed low, or loss of, expression of ER, PR, and BCL2, but conserved the expression of CK8. We have named them the luminal A and B groups, respectively, according to Sorlie's classification.32
We performed an unsupervised cluster analysis with the 50 sporadic breast tumors, using the same immunohistochemical markers, and identified the same five subgroups, with the same pattern of immunohistochemical expression, and similar proportions in each subgroup; 16% HER-2 positive; 15% basal like; 11% normal breast like; 42% luminal A; and 16% luminal B. We validated these results using the Silhouette Width technique (data not shown).
Somatic Inactivation of the BRCA1 Gene
We observed promoter hypermethylation of BRCA1 in 21 (42%) of the 50 cases, and these were distributed evenly over the two main groups (Figure 1). There was no evidence of an association between hypermethylation and grade, or any of the immunohistochemical markers studied.
We tested for LOH at the BRCA1 locus in 19 of the 21 cases with promoter hypermethylation of BRCA1 (there was no DNA from lymphocytes available for the other 2 cases with promoter hypermethylation), and observed LOH in seven (37%). All seven tumors with double somatic BRCA1 inactivation were in the high-grade branch and four (57%) of those were in the basal-like group (Figure 1).
Discussion
Immunohistochemical analysis of BRCA1 and BRCA2 tumors has previously shown a good correlation between genotype and phenotype,16, 19, 20 but there are very few studies on non-BRCA1/2 tumors.18, 20, 27, 33 We have previously reported that non-BRCA1/2 tumors tend to be grade 1–2 and ER and PR positive, have a low proliferative index, and express p53 to a similar extent to BRCA2 and sporadic tumors, but less than BRCA1 tumors.18, 19, 20 The present study has confirmed our previous results with new markers related to cell cycle, apoptosis, proliferation, stromal, and epithelial markers. We can say that non-BRCA1/2 tumors are of lower grade than sporadic tumors, but they are quite similar with respect to the immunohistochemical markers studied (Table 2).
Immunohistochemistry Classification of Familial Non-BRCA1/2 Tumors
By using a hierarchical unsupervised cluster analysis with 25 markers and grade, we have established a classification of the non-BRCA1/2 tumors that demonstrates their heterogeneity. Non-BRCA1/2 tumors can be divided into two main groups primarily according to their ER status and grade. The first group is characterized by higher grade; ER negativity; and the expression of proteins related to proliferation and cell cycle progression, and the second group by low grade; ER positivity; and overexpression of some cyclin–CDK complex inhibitors, antiapoptotic, and luminal (CK8) proteins (Figure 1).
The two main groups of our analysis can be further divided into five subgroups that are consistent with the classification system established by Sorlie et al32 in sporadic breast cancer, using a cDNA array study, as follows: (1) HER-2 positive; (2) basal like; (3) normal breast like, (4) luminal A, and (5) luminal B. In addition, we performed an unsupervised cluster analysis of 50 sporadic tumors using the same immunohistochemical markers used in the non-BRCA1/2 group, and obtained similar subgroups. Thus, we conclude that non-BRCA1/2 tumors are heterogeneous and that the immunohistochemical classification is very similar to that found for sporadic tumors using both expression arrays32 and immunohistochemical markers. We obtained consistent results after excluding families with any ovarian or male breast cancers (data not shown).
Somatic Inactivation of the BRCA1 Gene
We found that 42% of the non-BRCA1/2 tumors presented promoter hypermethylation of the BRCA1 gene, a percentage that is higher than in sporadic cases, where it ranges between 11 and 30%.34, 35 However these tumors were evenly distributed across both groups and hypermethylation was not associated with estrogen or progesterone receptor status, or (higher) grade, as has been suggested by some authors.35, 36 Seven (37%) of the non-BRCA1/2 tumors with promoter hypermethylation also had LOH at, and therefore total inactivation of, BRCA1. All seven were all in the high-grade group and the majority showed the basal-like phenotype (Figure 1), the latter being characteristic of BRCA1 tumors.37, 38, 39, 40 Although the sample size was small, we compared this subgroup with the rest of the non-BRCA1/2 tumors and found significant differences for the majority of the markers based on unadjusted P-values (Table 3). Consistent differences were observed when 33 BRCA1 tumors were compared with the latter group using previously published data.18 However, the same comparison between the seven BRCA1/2 tumors with somatic BRCA1 inactivation and the 33 BRCA1 tumors did not reveal significant differences even before correction for multiple testing (Table 3). In addition, six out of the seven tumors with BRCA1 promoter hypermethylation and LOH showed morphologic characteristics of medullary carcinoma, a subtype associated with BRCA1 tumors.41 All these observations suggest that in the majority of cases, a double somatic ‘hit’ in the BRCA1 gene is necessary for non-BRCA1/2 tumors to generate a BRCA1 phenotype; the first hit would be promoter hypermethylation and the second hit an LOH of the wild-type allele, and both would occur early in tumorigenesis in order to be able to mimic the immunohistochemical profile of BRCA1 tumors.
Genetic Implications of the New Familial Non-BRCA1/2 Tumor Classification
The classification by immunohistochemistry of non-BRCA1/2 tumors into five subgroups confirms that non-BRCA1/2 tumors constitute a heterogeneous group of tumors, and it supports the hypothesis that the majority of familial non-BRCA1/2 tumors might be explained by a polygenic model (that is, multiple low-penetrance genes),15 rather than by a single BRCAX gene. That is, our results could represent a practical validation of this hypothesis because we have found that non-BRCA1/2 tumors have the same immunohistochemical profile as that described in sporadic breast tumors, a type of tumor classically associated with a polygenetic model. This concept does not exclude the existence of some genes that could each explain a small number of families,13, 14 as was recently shown for CHEK2.42, 43 In fact, our classification could be very useful in defining more homogenous groups for linkage and other studies designed to identify such genes.
Clinical Implications of the BRCA1-like Group
The same mechanisms of somatic inactivation of the BRCA1 gene that we have described in 14% of our cases was recently observed in a group of sporadic breast tumors,35 and these tumors were also high grade and ER negative. These data confirm the specific characteristics (high grade and ER negative) of this group of cases that exhibit what has generically been named ‘BRCAness’, and that both sporadic44 and familial breast tumors can present as ‘BRCA like’. Some experimental studies using demethylating agents are now being conducted on different tumors with good results,45 and this opens up a new avenue for the treatment of tumors with an allele inactivated by hypermethylation that has to be explored further. On the other hand, the identification of this group of tumors with ‘BRCAness’ also provides a basis for new therapeutic strategies based on the sensitivity to DNA-damaging agents that BRCA1 tumors present.46 DNA repair protein PARP inhibitors and DNA crosslinking agents (cisplatin, mytomycin C, diepoxibutane) seem to affect tumor cells with BRCA1 mutations by inhibiting the DNA repair of single strand breaks and increasing their non-viability, while leaving normal cells or cells with a functional BRCA1 allele unaffected.47 Therefore, BRCA1 tumors occurring as a result of constitutional or somatic mutations could represent a new group for targeted therapeutic strategies.
References
Miki Y, Swensen J, Shattuck-Eidens D, et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994;266:66–71.
Wooster R, Bignell G, Lancaster J, et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 1995;378:789–792.
Easton DF, Bishop DT, Ford D, et al. Genetic linkage analysis in familial breast and ovarian cancer: results from 214 families. The breast cancer linkage consortium. Am J Hum Genet 1993;52:678–701.
Osorio A, Barroso A, Martinez B, et al. Molecular analysis of the BRCA1 and BRCA2 genes in 32 breast and/or ovarian cancer Spanish families. Br J Cancer 2000;82:1266–1270.
Shih HA, Couch FJ, Nathanson KL, et al. BRCA1 and BRCA2 mutation frequency in women evaluated in a breast cancer risk evaluation clinic. J Clin Oncol 2002;20:994–999.
van Beers EH, van Welsem T, Wessels LF, et al. Comparative genomic hybridization profiles in human BRCA1 and BRCA2 breast tumors highlight differential sets of genomic aberrations. Cancer Res 2005;65:822–827.
Seitz S, Rohde K, Bender E, et al. Strong indication for a breast cancer susceptibility gene on chromosome 8p12-p22: linkage analysis in German breast cancer families. Oncogene 1997;14:741–743.
Kainu T, Juo SH, Desper R, et al. Somatic deletions in hereditary breast cancers implicate 13q21 as a putative novel breast cancer susceptibility locus. Proc Natl Acad Sci USA 2000;97:9603–9608.
Smith P, McGuffog L, Easton DF, et al. A genome wide linkage search for breast cancer susceptibility genes. Genes Chromosomes Cancer 2006;45:646–655.
Rahman N, Teare MD, Seal S, et al. Absence of evidence for a familial breast cancer susceptibility gene at chromosome 8p12–p22. Oncogene 2000;19:4170–4173.
Thompson D, Szabo CI, Mangion J, et al. Evaluation of linkage of breast cancer to the putative BRCA3 locus on chromosome 13q21 in 128 multiple case families from the breast cancer linkage consortium. Proc Natl Acad Sci USA 2002;99:827–831.
Hedenfalk I, Ringner M, Ben-Dor A, et al. Molecular classification of familial non-BRCA1/BRCA2 breast cancer. Proc Natl Acad Sci USA 2003;100:2532–2537.
Seal S, Thompson D, Renwick A, et al. Truncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility alleles. Nat Genet 2006;38:1239–1241.
Rahman N, Seal S, Thompson D, et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat Genet 2007;39:165–167.
Antoniou AC, Pharoah PD, McMullan G, et al. A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br J Cancer 2002;86:76–83.
Honrado E, Osorio A, Palacios J, et al. Immunohistochemical expression of DNA repair proteins in familial breast cancer differentiate BRCA2-associated tumors. J Clin Oncol 2005;23:7503–7511.
Palacios J, Honrado E, Osorio A, et al. Phenotypic characterization of BRCA1 and BRCA2 tumors based in a tissue microarray study with 37 immunohistochemical markers. Breast Cancer Res Treat 2005;90:5–14.
Palacios J, Honrado E, Osorio A, et al. Immunohistochemical characteristics defined by tissue microarray of hereditary breast cancer not attributable to BRCA1 or BRCA2 mutations: differences from breast carcinomas arising in BRCA1 and BRCA2 mutation carriers. Clin Cancer Res 2003;9:3606–3614.
Lakhani SR, Van De Vijver MJ, Jacquemier J, et al. The pathology of familial breast cancer: predictive value of immunohistochemical markers estrogen receptor, progesterone receptor, HER-2, and p53 in patients with mutations in BRCA1 and BRCA2. J Clin Oncol 2002;20:2310–2318.
Eerola H, Heikkila P, Tamminen A, et al. Histopathological features of breast tumours in BRCA1-, BRCA2- and mutation-negative breast cancer families. Breast Cancer Res 2005;7:R93–R100.
Diez O, Osorio A, Duran M, et al. Analysis of BRCA1 and BRCA2 genes in Spanish breast/ovarian cancer patients: a high proportion of mutations unique to Spain and evidence of founder effects. Hum Mutat 2003;22:301–312.
Parmigiani G, Berry D, Aguilar O . Determining carrier probabilities for breast cancer-susceptibility genes BRCA1 and BRCA2. Am J Hum Genet 1998;62:145–158.
Chappuis PO, Kapusta L, Begin LR, et al. Germline BRCA1/2 mutations and p27(Kip1) protein levels independently predict outcome after breast cancer. J Clin Oncol 2000;18:4045–4052.
Rodriguez-Pinilla SM, Sarrio D, Honrado E, et al. Prognostic significance of basal-like phenotype and fascin expression in node-negative invasive breast carcinomas. Clin Cancer Res 2006;12:1533–1539.
Abd El-Rehim DM, Pinder SE, Paish CE, et al. Expression of luminal and basal cytokeratins in human breast carcinoma. J Pathol 2004;203:661–671.
Nielsen TO, Hsu FD, Jensen K, et al. Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 2004;10:5367–5374.
Oldenburg RA, Kroeze-Jansema K, Meijers-Heijboer H, et al. Characterization of familial non-BRCA1/2 breast tumors by loss of heterozygosity and immunophenotyping. Clin Cancer Res 2006;12:1693–1700.
Herrero J, Valencia A, Dopazo J . A hierarchical unsupervised growing neural network for clustering gene expression patterns. Bioinformatics 2001;17:126–136.
Herrero J, Al-Shahrour F, Diaz-Uriarte R, et al. GEPAS: a web-based resource for microarray gene expression data analysis. Nucleic Acids Res 2003;31:3461–3467.
Herman JG, Graff JR, Myohanen S, et al. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 1996;93:9821–9826.
Smith TM, Lee MK, Szabo CI, et al. Complete genomic sequence and analysis of 117 kb of human DNA containing the gene BRCA1. Genome Res 1996;6:1029–1049.
Sorlie T, Perou CM, Tibshirani R, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 2001;98:10869–10874.
Lakhani SR, Gusterson BA, Jacquemier J, et al. The pathology of familial breast cancer: histological features of cancers in families not attributable to mutations in BRCA1 or BRCA2. Clin Cancer Res 2000;6:782–789.
Esteller M, Silva JM, Dominguez G, et al. Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J Natl Cancer Inst 2000;92:564–569.
Wei M, Grushko TA, Dignam J, et al. BRCA1 promoter methylation in sporadic breast cancer is associated with reduced BRCA1 copy number and chromosome 17 aneusomy. Cancer Res 2005;65:10692–10699.
Catteau A, Harris WH, Xu CF, et al. Methylation of the BRCA1 promoter region in sporadic breast and ovarian cancer: correlation with disease characteristics. Oncogene 1999;18:1957–1965.
Foulkes WD, Stefansson IM, Chappuis PO, et al. Germline BRCA1 mutations and a basal epithelial phenotype in breast cancer. J Natl Cancer Inst 2003;95:1482–1485.
Palacios J, Honrado E, Osorio A, et al. Re: germline BRCA1 mutations and a basal epithelial phenotype in breast cancer. J Natl Cancer Inst 2004;96:712–714.
Sorlie T, Tibshirani R, Parker J, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 2003;100:8418–8423.
Lakhani SR, Reis-Filho JS, Fulford L, et al. Prediction of BRCA1 status in patients with breast cancer using estrogen receptor and basal phenotype. Clin Cancer Res 2005;11:5175–5180.
Lakhani SR, Easton DF, Stratton MR, et al. Pathology of familial breast cancer: differences between breast cancers in carriers of BRCA1 or BRCA2 mutations and sporadic cases. Breast cancer linkage consortium. Lancet 1997;349:1505–1510.
Meijers-Heijboer H, van den Ouweland A, Klijn J, et al. Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in non-carriers of BRCA1 or BRCA2 mutations. Nat Genet 2002;31:55–59.
Consortium TCBCC-C . CHEK2*1100delC and susceptibility to breast cancer: a collaborative analysis involving 10,860 breast cancer cases and 9065 controls from 10 studies. Am J Hum Genet 2004;74:1175–1182.
Turner N, Tutt A, Ashworth A . Hallmarks of ‘BRCAness’ in sporadic cancers. Nat Rev Cancer 2004;4:814–819.
Missiaglia E, Donadelli M, Palmieri M, et al. Growth delay of human pancreatic cancer cells by methylase inhibitor 5-aza-2′-deoxycytidine treatment is associated with activation of the interferon signalling pathway. Oncogene 2005;24:199–211.
Turner N, Tutt A, Ashworth A . Targeting the DNA repair defect of BRCA tumours. Curr Opin Pharmacol 2005;5:388–393.
Farmer H, McCabe N, Lord CJ, et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 2005;434:917–921.
Acknowledgements
This work was partially financed by grants from the Ministry of Health and Ministry of Science, FIS04-2224 and SAF03-02497, respectively. EH was financed by the Asociación Española Contra el Cáncer (AECC). We thank the Spanish National Tumor Bank Network for providing samples; Lydia Sánchez, Ana Díez, Ma Jesús Acuña, and Raquel Pajares for their technical assistance; and Jaime Huertas for kind help with CAAT.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Honrado, E., Osorio, A., Milne, R. et al. Immunohistochemical classification of non-BRCA1/2 tumors identifies different groups that demonstrate the heterogeneity of BRCAX families. Mod Pathol 20, 1298–1306 (2007). https://doi.org/10.1038/modpathol.3800969
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800969
Keywords
This article is cited by
-
Molecular characterization and clinical interpretation of BRCA1/BRCA2 variants in families from Murcia (south-eastern Spain) with hereditary breast and ovarian cancer: clinical–pathological features in BRCA carriers and non-carriers
Familial Cancer (2017)
-
Tumor characteristics and prognosis in familial breast cancer
BMC Cancer (2016)
-
Molecular Subtyping of Serous Ovarian Cancer Based on Multi-omics Data
Scientific Reports (2016)
-
RNA profiling reveals familial aggregation of molecular subtypes in non-BRCA1/2 breast cancer families
BMC Medical Genomics (2014)
-
Etiology of familial breast cancer with undetected BRCA1 and BRCA2 mutations: clinical implications
Cellular Oncology (2014)