Abstract
DNA copy number changes were investigated in 51 (19 uterine and 32 nonuterine) primary leiomyosarcomas by comparative genomic hybridization. The aim was to evaluate whether true biological differences exist between uterine and nonuterine leiomyosarcoma and whether changes revealed by comparative genomic hybridization have prognostic value. Genomic imbalances were found in 48 (94%) cases. The most frequent DNA copy number changes were losses in 10q (35%), 13q (57%), and 16q (41%), gains in 1q (41%), and gains and high-level amplifications in 17p (39%). Gains were nearly as frequent as losses in both uterine and nonuterine leiomyosarcoma. Correlation-based tree modeling revealed two clusters that segregated significantly a group of uterine (gains at 1q11–q24) and a group of nonuterine (losses at 13q14–q34, 16q11.1–q24, and 10q21–q26) cases. The nonuterine cluster was associated with subcutaneous origin and a trend toward increased metastasis-free survival. Further explorative analyses identified aberrations associated with shorter metastasis-free survival time, including losses at 2q32.1–q37 and gains at 8q24.1–q24.3, whereas the cases with losses at 6cen-p25 showed longer metastasis-free survival time.
Similar content being viewed by others
Main
Leiomyosarcomas are malignant mesenchymal tumors composed of cells with phenotypic features of smooth muscle differentiation.1, 2, 3 These tumors occur in a wide range of anatomic body sites. Although the histological appearance of all leiomyosarcomas, regardless of the anatomic site, is similar, they are usually divided into site-related groups due to clinical and biological differences.1, 2, 3 Leiomyosarcoma is one of the most common sarcomas of the uterus4, 5 but less frequent in external soft tissues. Subcutaneous and deep-seated leiomyosarcomas of soft tissue occur in approximately equal proportions, and they are supposed to originate from small to medium-sized veins.6, 7, 8, 9, 10 Cutaneous leiomyosarcomas constitute an entity that arises from dermis, most probably from arrectores pilorum muscles, and shows an indolent course of disease in contrast to leiomyosarcomas of other locations.11, 12
Cytogenetic analyses of more than 100 leiomyosarcoma cases have been reported.13, 14 Standard karyotyping and fluorescent in situ hybridization have not revealed any single specific aberration common to all or most leiomyosarcomas, but many nonrandom structural aberrations and numerical changes have been detected. Cytogenetic signs of gene amplification have also been seen in leiomyosarcoma. Most reported karyotypes are complex, contain a high number of chromosomal changes, and show marked heterogeneity within this group of sarcomas.14, 15, 16, 17, 18, 19 Comparative genomic hybridization (CGH) studies have also shown multiple and complex changes, suggesting a genetically advanced disease. So far, CGH has been used to identify gains and losses of DNA copy number changes in less than 200 cases of leiomyosarcoma. Extensive genetic imbalances have been detected in nearly all cases.20, 21, 22, 23, 24, 25, 26, 27, 28
Surgery is the main treatment modality for leiomyosarcoma irrespective of localization. Pre- or postoperative radiation therapy is used in high-grade tumors and in cases where sufficient surgical margins cannot be obtained. In uterine leiomyosarcoma, the primary therapy is extrafascial hysterectomy and bilateral salphingo-oophorectomy with or without pelvic lymphadenectomy.29 Radiation therapy or chemotherapy is used as the only treatment modality when the tumor is thought to be inoperable or widespread.5, 30 The use of cytotoxic drugs as adjuvant therapy has been more common in uterine leiomyosarcoma than in external soft tissue leiomyosarcoma, even when no significant survival benefit has been derived.5, 29, 31 It is uncertain whether this difference in treatment tradition reflects a true difference in biology.
In order to evaluate whether true biological differences exist between uterine and non-uterine leiomyosarcoma, we undertook to study 51 primary leiomyosarcomas, which to our knowledge represent the largest series of leiomyosarcoma cases analyzed by the CGH technique so far. The aims of this study were to compare the genetic changes in these two leiomyosarcoma groups using CGH to obtain a genome-wide pattern of chromosomal gains, losses, and high-level amplifications of small chromosomal areas, and to further evaluate their possible prognostic and/or diagnostic significance.
Materials and methods
Tumor Specimens
The material consisted of 51 primary leiomyosarcoma samples obtained from 51 Finnish patients treated at the Helsinki University Central Hospital, Helsinki between 1981 and 2003. The primary tumors were situated on the extremities, trunk wall, in the superficial head and neck region, and in the uterus. Complete clinical and follow-up data were available. Surgical and adjuvant treatments, and development of local and distant recurrences were documented. Two experienced sarcoma pathologists (TB, RB) performed the histopathological review to confirm the diagnosis and re-evaluate the tumor grading. All cases were histologically clear leiomyosarcomas showing smooth muscle differentiation. Immunohistochemically, the tumors were positive for α-smooth muscle actin and showed at least focal positivity for desmin. The uterine leiomyosarcoma diagnosis was based on the morphological criteria by Bell et al.32 No uterine epitheloid or myxoid leiomyosarcomas were included in the study and also tumors of uncertain malignant potential were excluded. No universally agreed grading system for uterine leiomyosarcoma exists. However, these tumors were categorized to ‘low grade’ or ‘high grade’ based on the degree of cellular atypia, amount of tumor necrosis and number of mitotic figures. A four-grade system based on similar parameters was applied to non-uterine tumors (I–II, low grade; III–IV, high grade). The depth of the superficial (cutaneous vs subcutaneous) non-uterine tumors was specified, and possible connection to a blood vessel was determined microscopically and from the primary pathologist's reports.
Of the 51 patients, 32 (63%) (14 males and 18 females) had non-uterine leiomyosarcoma and 19 (37%) uterine leiomyosarcoma. The non-uterine group consisted of nine cutaneous, 11 subcutaneous and 12 deep-seated tumors. The whole study series comprised 19 (37%) low-grade and 32 (63%) high-grade tumors. The median age of the patients at diagnosis was 59 years (range, 20–91). Tumor sizes varied from 0.7 to >20.0 cm, with a median tumor size of 5.0 cm (non-uterine 3.5 cm and uterine 7.0 cm). None of the patients had received chemo- and/or radiotherapy before surgery. The median follow-up time for all surviving patients was 73 months, ranging from 9 months to 212 months. Metastatic dissemination was observed in five (10%) of the patients at the presentation, and 16 patients developed a metastatic course of disease during the follow-up; altogether 41% of the patients progressed to metastasis. Local recurrence was recorded in 10 (20%) patients. Clinical characteristics of the patients are presented in Table 1.
Comparative Genomic Hybridization
DNA was extracted from paraffin-embedded tissue sections from all 51 tumor samples following the procedure reported by Isola et al.33 CGH was performed using direct fluorochrome-conjugated DNA for all samples as described elsewhere.34, 35 Briefly, tumor and reference DNA (genomic DNA from peripheral blood leukocytes from normal donors) were labeled with fluorescein-iso-thiocyanate (FITC)-conjugated dCTP and dUTP (Dupont, Boston, MA, USA), and Texas red-conjugated dCTP and dUTP (Dupont) by nick translation, respectively, to obtain fragments ranging from 600 to 2000 bp, as published previously.35, 36 The hybridization mixture consisted of 400 ng tumor DNA, 400 ng reference DNA, and 10 μg unlabeled human Cot-1 DNA (Gibco/BRL, Life Technologies, Gaithersburg, MD, USA) dissolved in 10 μl hybridization buffer (50% formamide, 10% dextran sulfate, 2 × SSC). The hybridization mixture was denatured at 75°C for 5 min and hybridized to a slide with normal metaphase spreads denatured in 70% formamide/2 × SSC (pH 7) at 68°C for 2 min. Hybridization was carried out at 37°C for 48 h. Then the slides were washed three times in 50% formamide/2 × SSC (pH 7), twice in 2 × SSC, and once in 0.1 × SSC at 45°C, followed by 2 × SSC, 0.1 M NaH2PO4—0.1 M Na2HPO4—0.1% Nonidet P-40 (pH 8), and distilled water at room temperature for 10 min each. After air-drying, the slides were counterstained with 4′,6-diamidino-2-phenyl-indole-dihydrochloride (DAPI) (Sigma Chemical Co., St Louis, MO, USA) and mounted using an antifading medium (Vectashield®, Vector Laboratories, Burlingame, CA, USA).
Digital Image Analysis
The hybridization was analyzed using an Olympus fluorescence microscope and the ISIS digital image analysis system (MetaSystems GmbH, Altlussheim, Germany), based on an integrated high-sensitivity monochrome charge-coupled device (CCD) camera and automated CGH analysis software. Three-color images (green for tumor DNA, red for reference DNA, and blue for counterstaining) were acquired from 12 metaphases per sample. The chromosomal regions were interpreted as over-represented when the green-to-red ratio exceeded 1.17 (gains) or 1.5 (high-level amplifications), and as under-represented (losses) when the ratio was less than 0.85. In each CGH experiment, a negative (peripheral blood DNA from normal donor) and positive (tumor DNA with known copy number changes) control were included and run simultaneously with the tumor samples. Telomeric and heterochromatinic regions were excluded from the analysis when they appeared as the sole aberration present in the sample, as these regions cannot be evaluated reliably by CGH.36, 37 All results were confirmed using a 99% confidence interval with a 1% error probability. Briefly, intraexperimental s.d. for all positions in the CGH ratio profiles were calculated from the variation of the ratio values of all homologous chromosomes within the experiment. Confidence intervals for the ratio profiles were then computed by combining them with an empirical interexperimental s.d. and by estimating the error probability based on the t-distribution.
Determination of Minimal Common Chromosomal Regions
CGH analyzes at a resolution down to the sub-band level of the chromosomes. CGH data were displayed on chromosome ideograms, where losses and gains of chromosome regions are presented as lines on the left or the right side of the ideogram, respectively. The minimal common regions of DNA copy number changes in the whole series were determined visually from the ideogram (supplementary data at www.helsinki.fi/cmg/cgh_data). The ideograms in Figures 1 and 2 show the difference between the uterine and non-uterine cases.
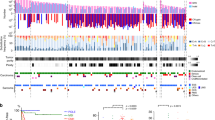
Summary of gains and losses of DNA sequence copy number in 32 primary non-uterine leiomyosarcoma samples analyzed by CGH. Losses are shown on the left side of the chromosome and gains on the right. Each vertical line represents a genetic alteration seen in one sample. High-level amplifications of small chromosomal regions are shown as thick lines.
Summary of gains and losses of DNA sequence copy number in 19 primary uterine leiomyosarcoma samples analyzed by CGH. Losses are shown on the left side of the chromosome and gains on the right. Each vertical line represents a genetic alteration seen in one sample. High-level amplifications of small chromosomal regions are shown as thick lines.
Statistical Analysis
For both univariate and higher order statistical modeling of aberration patterns, a data set was constructed to reflect the presence or absence of aberrations in each individual case. Aberrations with no or limited information were excluded by requiring at least 10.0% of cases to share any particular aberration. Owing to their low overall occurrence, even single cases with high-level amplifications were included in the data set. The data was coded into binary variables, whereby 0 (zero) represented the absence and 1 (one) the presence of an aberration.
To assess the association between each individual chromosomal aberration and relevant clinical parameters, including tumor subtype (uterine vs non-uterine), localization (cutaneous, subcutaneous, deep-seated or uterine), histologic grade (low-grade vs high-grade), and metastasis, a ‘prediction score’ was calculated according to Golub et al.38 The score reflects the degree of correlation between the observed aberration pattern for all cases with an idealized pattern that perfectly discriminates between the categories of the clinical parameter being analyzed. In total, 173 scores were calculated and ranked in descending order, effectively placing aberrations with high discriminatory strength to the top of the list. A χ2 test was subsequently run on the aberrations that generated the top 10 ranking scores to evaluate whether the proportion of cases with and without a particular aberration was statistically significantly different among the categories of the clinical parameter in question. Metastatic status was further explored taking the time variable into account using the log-rank test to assess the differences in metastasis-free survival rates, calculated using the Kaplan–Meier method, between cases with and without a particular aberration. To reduce the likelihood of false positive results that may occur as a consequence of performing multiple individual univariate statistical tests, a conservative approach was adopted in interpreting the results. As chromosomal aberrations cannot be considered independent events, especially when these occur in adjacent loci, the Bonferroni type of correction was deemed inappropriate and an alternative strategy was devised. Only the results with P<0.05 in at least two adjacent chromosomal loci were considered and the reported individual P-value was set to equal the highest (least significant) one.
Correlation-based tree modeling was applied to the data in order to create a correlation matrix, describing the relationship between all possible pair-wise comparisons between individual cases. Case similarities were calculated using the Pearson correlation adjusted to account for values centered to zero. The values of the matrix were used to generate a tree structure or dendrogram, employing an agglomerative algorithm using unweighted pair-group average linkage amalgamation rules. Branch lengths in the resulting tree reflect the degree of correlation, while joining nodes depict the pairs of cases with the highest correlation. Cases with similar patterns of aberrations will achieve a high degree of correlation and thus appear close in the dendrogram, whereas cases with little in common with respect to aberration pattern will end up far apart. The same algorithm was employed to cluster the aberrations with respect to case patterns. The resulting tree-models representing correlations between the cases (vertical orientation) and correlations between the aberrations (horizontal orientation) were depicted in a 2-dimensional graph that displayed also the pattern (as absence or presence) of any particular aberration and case. Associations between the branches at any level of the tree and the clinical parameters mentioned above, that is, tumor subtype, localization, histologic grade and metastasis, were assessed by overlaying the respective information on the 2-D dendrogram and analyzing their relation to the vertical branching pattern. The statistical evaluation was performed by comparing the distribution of cases representing the different levels of a given clinical parameter to the distribution observed in all 51 cases. The F or binomial distributions were used in the calculations according to the number of categories in the tested parameter. Aberrations that contributed most significantly to the identified branching patterns were calculated in a similar fashion, using Fisher's exact test to compare the observed and expected distributions. Lists ranking the relative importance of aberrations for a given cluster in the tree were obtained by taking the negative logarithm of the P-values from Fisher's exact test.
Results
Overview of DNA Sequence Copy Number Changes Revealed by CGH
Of the 51 leiomyosarcoma samples studied, 48 (94%) had changes with a mean value of 11. 02±0.96 aberrations per sample (range, 1–25). Three samples (6%) did not show any aberrations. The absence of detectable changes could be attributable to non-neoplastic DNA extracts, intratumor heterogeneity, or the underlying shortcomings in the resolution of CGH.
Gains of DNA copy number changes were as frequent as losses (gains:losses=1.0:1.1) with a mean value of 4.88±0.51 (range, 0–15) and 5.40±0.55 (range, 0–14) aberrations per sample, respectively. High-level amplifications of small chromosomal regions were found in 23 out of the 51 (45%) tumors analyzed with a mean value of 0.75±0.15 aberrations per sample (range, 0–5).
Of the 32 non-uterine tumors (nos 1–32), 30 (94%) had changes with a mean value of 10.53±1.03 aberrations per sample (range, 2–22). Two low-grade tumors (6%) did not show any aberrations (nos 1 and 4). Gains were as frequent as losses (gains:losses=1.0:1.1) with a mean value of 4.67±0.51 (range, 0–11) and 5.10±0.63 (range, 0–13) aberrations per sample, respectively. High-level amplifications were found in 16 out of the 32 tumors analyzed with a mean value of 0.77±0.17 aberrations per sample (range, 0–5).
Among the 30 non-uterine tumors with DNA copy number changes, gains were frequent in chromosomes 1 (40%), 5 (37%), 8 (27%), 9 (27%), 15 (27%), 17 (40%), 19 (33%), 20 (30%), and 22 (33%). High-level amplifications were found in 16 out of the 30 (53%) abnormal tumors analyzed, with the highest frequency in chromosome 17 (33%). Losses frequently affected chromosomes 2 (37%), 6 (40%), 8 (30%), 10 (50%), 13 (70%), 16 (40%), and X (37%). Figure 1 summarizes all chromosomal regions with increased or decreased DNA sequence copy number changes in the non-uterine leiomyosarcoma samples.
Of the 19 uterine tumors (nos 33–51), 18 (95%) had changes with a mean value of 11.83±1.93 aberrations per sample (range, 1–25) while only one (5%) of the low-grade tumors (no 33) did not reveal any DNA copy number changes. Gains were as frequent as losses (gains:losses=1.0:1.1) with a mean value of 5.22±1.10 (range, 0–15) and 5.89±1.03 (range, 0–14) aberrations per sample, respectively. High-level amplifications were found in 7 out of the 19 tumors analyzed with a mean value of 0.72±0.30 aberrations per sample (range, 0–5).
Among the 18 uterine tumors with DNA copy number changes, gains were frequent in chromosomes 1 (56%), 3 (28%), 5 (28%), 8 (44%), 12 (28%), 16 (28%), 17 (33%), and 20 (28%). High-level amplifications were found in seven out of the 18 (39%) abnormal tumors analyzed, with the highest frequency in chromosome 1 (17%). Losses frequently affected chromosomes 2 (50%), 4 (28%), 6 (39%), 7 (33%), 11 (44%), 13 (44%), 14 (39%), and 16 (50%). Figure 2 summarizes all chromosomal regions with increased or decreased DNA sequence copy number changes in the uterine leiomyosarcoma samples. Table 2 shows the copy number karyotypes of the 51 primary leiomyosarcoma samples analyzed by CGH.
Minimal Common Regions of DNA Sequence Copy Number Changes
The minimal common regions of DNA copy number changes in the whole series of 51 leiomyosarcomas were determined. The minimal common regions of recurrent gains were 1q11–q31 (24–33%), 5cen-p15.3 (22–25%), 8q21.3–q24.3 (22–25%), 15q (14–20%), 17cen.p12 (22–24%), 17cen.q21 (24%), 19p (22–24%), 20q (25%), and 22q (24–27%), and the most frequent high-level amplifications were 1q21 (8%), 5p13.p15.2 (8%), and 17p (14–18%). The regions of recurrent losses were 2q32.1–q37 (22–29%), 6cen.p25 (22–24%), 6cen.q22 (24–25%), 9p21.p24 (22%), 10cen.p15 (22–24%), 10cen.q26 (27–35%), 13q (31–57%), 14q (8–18%), 16q (35–41%), and Xq21-q28 (27–29%).
The minimal common regions in all leiomyosarcoma samples are presented at our web site (supplementary data www.helsinki.fi/cmg/cgh_data).
Results from Statistical Analyses
To evaluate whether specific aberration patterns could be associated with clinically relevant parameters, such as tumor subtype (uterine vs non-uterine), localization (cutaneous, subcutaneous, deep-seated, or uterine), histologic grade (low vs high), and metastatic recurrence, the CGH data was studied in greater detail using both univariate and higher order statistical modeling. In order to focus the analysis on the most prominent and representative aberrations, simultaneously achieving the aim of minimizing the occurrence of noise in the data, inclusion cut-offs were defined. Gains and losses were included in the analysis when at least 10.0% of the cases shared the aberration, whereas the considerably less frequent high-level amplifications were included even when single cases were affected. Consequently, the number of aberrations came down from 387 loci to 173 aberrations, the minimal common regions in all 51 tumors (supplementary data at www.helsinki.fi/cmg/cgh_data).
The significance of individual aberrations was explored in greater detail by ranking the aberrations with respect to their power to discriminate between the categories of clinical parameters, based on the calculation of a ‘prediction strength’ score,38 followed by univariate χ2 testing of the top ranking aberrations. No statistically significant results were found for tumor subtype, localization or grade, but for metastatic status significant differences were found in the cases with and without losses at 6cen-p25 and 2q32.1–q37, or gains at 8q24.1-q24.3, with adjusted P-values of 0.017, 0.035, and 0.035, respectively. Further analysis, which took the time variable into account, used the log-rank test to evaluate the differences in metastasis-free survival rates with regard to aberration status, confirmed the importance of these aberrations and resulted in adjusted P-values of 0.012, 0.009, and 0.009 (order as above). The cases with these aberrations demonstrated a decreased survival rate, with the notable exception of cases with losses at 6p, for which the opposite was found, suggesting that losses at 6p confer a protective effect resulting in significantly higher metastasis-free survival rates as compared to cases without them (Figure 3).
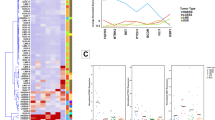
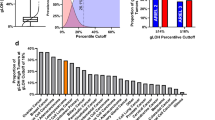
Higher order modeling of the data, both with respect to correlations between cases and correlations between aberrations, are summarized in a 2-dimensional dendrogram (Figure 4a). Vertical branching pattern did not identify any clear-cut division into clusters that would exclusively represent individual categories according to any of the tested clinical parameters, that is, tumor subtype, localization, histologic grade, and metastasis. However, closer scrutiny of the branching structure by overlaying information of the leiomyosarcoma subtype (uterine vs non-uterine) revealed two branches, a few levels down from the top of the dendrogram, that clearly segregated the two subtypes. The larger branch, denoted cluster A (Figure 4a, b), consisted of 11 cases, of which 10 (91%) were non-uterine, whereas the smaller branch, cluster B (Figure 4a, c), included six cases, of which five (83%) were uterine. Statistically both of these proportions were significantly different from the distribution expected to be observed by chance, P=0.044 and 0.030 for clusters A and B, respectively. Further characterization of the clusters disclosed some interesting features regarding the cases in cluster A. Compared to the overall distribution of cases with regard to tumor localization (cutaneous, subcutaneous, deep-seated, or uterine), cluster A had a significantly higher proportion of subcutaneous tumors and a significantly lower proportion of deep-seated tumors (P=0.024). A trend indicating a lower proportion of metastatic recurrence among these cases was also observed (P=0.098), which was additionally substantiated when the time variable was included in the log-rank test for difference in the Kaplan–Meier curves (P=0.088). Cases in cluster B did not exhibit similar significant differences when compared to the whole study group (Figure 4d). In an effort to determine the aberrations underlying the observed segregation into the two clusters, the ability of each aberration to predict cluster membership was evaluated using Fisher's exact test. A list ranking the aberrations with respect to prediction strength was obtained by taking the negative logarithm of the P-values from Fisher's exact test. Among the highest ranked aberrations were losses in 13q14-q34, 16q11.1-q24, and 10q21-q26 as well as gains in 1q11-q24 (Table 3). All the above losses were specific to cluster A, whereas gains in 1q were specific to cluster B.
(a) Two-dimensional correlation-based tree modeling of 51 leiomyosarcoma cases and 173 chromosomal aberrations. Each column represents a case and each row an aberration. Red in the matrix indicates presence and gray absence of a particular aberration. The dendrogram at the top shows the degree of similarity between the cases, and the dendrogram on the left side shows similarities between aberrations. The clusters that segregate non-uterine cases (cluster A) and uterine cases (cluster B), and the aberrations that most significantly contribute to the segregation pattern are highlighted. (b and c) Sub-dendrograms of clusters A and B in panels b and c, respectively, show non-uterine cases in red and uterine in green. (d) Kaplan–Meier curve shows differences in metastasis-free survival between cluster A (P=0.088) (red curve), cluster B (green curve) and the whole study group (black curve).
When the branching pattern was analyzed for the horizontal tree-model based on correlations between aberrations, it was apparent that correlation strength associated with physical closeness. With only a few exceptions, likely due to small number of occurrences limiting the accuracy of the calculations, clusters of aberrations consisted of events affecting adjacent loci on the same chromosome segment. There were thus no indications of physically unrelated aberrations co-occurring in the study group.
Discussion
Our study reports a CGH analysis performed on a large series of primary leiomyosarcomas comprising 19 uterine tumors and 32 non-uterine tumors of various external soft tissue sites. The aim was to evaluate whether true biological differences exist between uterine and non-uterine leiomyosarcomas. We obtained a genome-wide pattern of chromosomal gains, losses, and high-level amplifications of small chromosomal areas, and evaluated their possible prognostic and/or diagnostic significance using high-order statistical modeling. Two clusters clearly segregated uterine samples from non-uterine samples and the non-uterine cluster showed a trend of increased metastasis-free survival time. Furthermore, we identified some DNA copy number changes that are associated with higher metastatic potential.
Similarities and Differences in DNA Sequence Copy Number Changes of Uterine and Non-Uterine Leiomyosarcoma
The CGH analysis showed genomic imbalances in 48 (94%) of the 51 leiomyosarcomas studied. We found some general similarities between uterine and non-uterine leiomyosarcomas, such as gains of DNA copy number changes were nearly as frequent as losses in both uterine and non-uterine samples. In accordance with previous CGH studies, we found highly complex karyotypic changes in both subtypes. None of the most frequent gains or losses in either tumor group were unique and no specific aberration was restricted to any subtype of leiomyosarcoma. However, we found some typical and nearly consistent DNA copy number changes, including high frequency of losses in 10q (35% of all 51 tumors), 13q (57%) and 16q (41%), gains in 1q (41%), and gains and high-level amplifications in 17p (39%); aberrations that have been reported in previous CGH studies of leiomyosarcoma.21, 23, 24, 25, 26, 27, 28 The high frequency of chromosome imbalances suggests the presence of pathogenetically relevant oncogenes or tumor suppressor genes within these chromosome regions.
The purpose of our study was to evaluate possible differences between chromosomal aberrations in uterine and non-uterine leiomyosarcoma. To our knowledge, no previous study has focused on this issue or analyzed CGH results of leiomyosarcoma using correlation-based tree modeling, which was possible on this a large series of cases with complex chromosome imbalances. However, similar type of unsupervised clustering analysis, which is commonly used for the analysis of gene expression arrays, has also been applied to analysis of CGH data of sarcoma.39 Although no pathognomonic aberrations for non-uterine or uterine leiomyosarcoma, or clear-cut division into clusters exclusively representing any of the clinical parameters (subtype, localization, grade, and metastasis) tested were found, we revealed two clusters that segregated significantly a group of uterine tumors with gains at 1q11–q24 and a group of non-uterine tumors with losses at 13q14–q34, 16q11.1–q24, and 10q21–q26.
In the larger cluster, 10 of 11 tumors were non-uterine, and in the smaller cluster 5 of six tumors were uterine. The non-uterine cluster showed a trend indicating decreased metastatic recurrence, and the proportion of subcutaneous tumors was significantly higher than expected. As statistical testing of differences in proportion of low-grade or small tumors in the non-uterine cluster with favorable survival rates did not show significance, these parameters did not explain the finding (data not shown). The highest ranked aberrations in the non-uterine cluster, that is, losses in 13q, 16q, and 10q, have previously been connected and associated with decreased metastatic rate. Our results are in accord with the cytogenetic study by the CHAMP Study Group,18 which showed that losses of 10q and 13q segments were more frequent among patients without metastases. Interestingly, El-Rifai et al21 found frequent losses in 10q and 13q, and 71% of cases with loss in 13q had losses in 10q, suggesting a cumulative effect of deletions of several tumor suppressor genes in leiomyosarcoma. Furthermore, Hu et al27 studied exclusively uterine leiomyosarcoma and found that loss of 13q (without loss of 10q) was associated with longer survival time. They suggested that loss of tumor suppressor gene in 13q may be an early event in the development of leiomyosarcoma, and activation or inactivation of other oncogenes and tumor suppressor genes may be responsible for tumor progression.
In the present study, deletions of chromosome 13 involved several minimal overlapping regions of 13q. The region at 13q14, which contains the tumor suppressor gene RB, showed losses in 55% of the cases. Alterations of RB are frequent in a wide range of malignancies, in leiomyosarcoma as well as in other soft-tissue sarcomas.40, 41 Analyses of the genes and proteins in the Rb-cyclin D pathway have revealed frequent abnormalities in leiomyosarcoma.23, 42, 43
Standard cytogenetic methods have rarely revealed rearrangements of 10q in leiomyosarcoma.16 By CGH, we observed recurrent losses at 10q with the highest rate at 35% in the most frequent minimal common region of 10q23.q25. This finding is consistent with previous results reported in leiomyosarcoma and a wide range of other malignancies.21, 23, 25, 27, 41, 44, 45 The tumor suppressor gene PTEN is mapped to 10q23.45 This chromosomal region is commensurate with the frequent loss of heterozygosity of chromosome 10 demonstrated in uterine leiomyosarcoma but not in leiomyomas.46 Moreover, MXI1, a negative regulator of the MYC oncoprotein having a tumor suppressor function is mapped to 10q24–25.41, 47
About 41% of our cases showed losses in 16q. This observation is in agreement with the findings that Hu et al27 reported in uterine leiomyosarcoma. Several putative tumor suppressor genes are assigned to this region. Among them are RB2/p130 (16q12.2), CDH1 (16q22.1), and CDH13 (16q24.2.q24.3).41, 48
Gains in chromosome 1q were specific for the small uterine cluster. Furthermore, 33% of our cases with gains in chromosome 1q shared a minimal common overlapping region of 1q21. Also high-level amplification of this minimal common region was observed. Skubitz et al49 demonstrated that the expression of cellular retinoic acid-binding protein gene type II (CRABP2), located at 1q21.3, is five- to 10-fold in leiomyosarcoma as compared to normal myometrium. Increased sequence copy number at 1q21.q23 has been detected frequently in several malignancies (see Knuutila et al50 and references therein), but especially in sarcomas, including leiomyosarcoma, liposarcoma, osteosarcoma, chondrosarcoma, and malignant fibrous histiocytoma.21, 24, 26, 27, 34, 44, 51, 52, 53, 54, 55, 56 This region harbors several genes of potential significance.56, 57, 58
Our study showed gains and high-level amplifications in 17p in approximately 40% of the cases. A difference in the frequency of high-level amplifications affecting 17p was observed between non-uterine (28%) and uterine tumors (5.3%). Thus our data support previous CGH analyses, which have reported high-level amplifications in 17p in 20–30% of extra-uterine cases and, sporadically, in uterine cases.21, 23, 24, 26, 27 One of the best known tumor suppressor genes, TP53, is located at 17p13, and p53 and p53 protein overexpression has been described in leiomyosarcoma.42, 59, 60, 61, 62 Gains as well as high-level amplifications at 17p11.p12 seem to be frequently involved in other sarcomas, including osteosarcoma, chondrosarcoma, and malignant fibrous histiocytoma.21, 23, 27, 34, 44, 51, 52, 53, 63, 64 This suggests that different sarcoma entities most probably share a common pathogenic pathway.
In the present study, we found high-level amplifications in 1q, 5p, and Xp, which have been sporadically found in previous leiomyosarcoma studies.20, 21, 22, 23, 25 Recurrent high-level amplifications on chromosome arms 5p, 17p, and Xp have also been observed in osteosarcoma, malignant fibrous histiocytoma, and liposarcoma.51, 53, 65, 66 Interestingly, the frequently amplified regions we observed on chromosome arms 5p, 17p, and Xp all harbor genes involved in the ubiquitin-mediated protein degradation pathway.67 Otaño-Joos et al23 have described a similar pattern of chromosomal involvement.
DNA Copy Number Changes Associated with Metastasis in Leiomyosarcoma
Comparison of our findings to clinicopathological factors showed no significant correlation between any single CGH aberration and tumor subtype (uterine vs non-uterine), localization (cutaneous, subcutaneous, deep-seated, or uterine), or grade (low vs high). However, statistical analysis of metastasis-free survival revealed a significantly shorter survival time for cases with losses at 2q32.1–q37 or gains at 8q24.1–24.3. In contrast, cases with losses at 6cen-p25 showed significantly longer metastasis-free survival time. We further analyzed whether cases with and without the aberration in question differed with respect to proportion of cases with respective tumor localizations or grade. We revealed a statistically significant increase in the proportion of high-grade tumors among cases with gain at 8q24.1 (P=0.020) and a similar strong trend for losses at 2q33–q34 and 2q36–q37 (P=0.052) (data not shown). Non-significant values were observed for the remainder of aberrations and with regard to tumor localization.
Losses at 2q32.1–q37 and 6cen-p25 have previously not been associated with metastasis in leiomyosarcoma. However, losses of regions at 2q have been detected in osteosarcoma, with minimal common region 2q34-qter,51 and a study of Ewing sarcoma and related tumors cases with gains at 6p21.1-pter showed a statistically significant decrease in metastasis-free and overall survival.68 The number of previous studies on leiomyosarcoma is small and only few associations between DNA copy number changes in leiomyosarcoma and prognosis have been observed.18, 21, 27, 28
In the present study, we found that gains at 8q24.1–q24.3 were significantly associated with metastatic potential. The 8q24.1 region contains the proto-oncogene c-MYC, which is known to be amplified in several tumors. Frequent gains and high-level amplifications in 8q, with the minimal common overlapping region of 8q24.1, have been described in large (5–20 cm) and very large (>20 cm) leiomyosarcomas, but these findings were not tested statistically for metastasis.21 In synovial sarcoma, gains of 8q were significantly overrepresented in large tumors.69 Furthermore, in Ewing sarcoma and related tumors, copy number increases of chromosome 8 were associated with trends towards worse metastasis-free and overall survival,68 in osteosarcoma with a statistically significant poor metastasis-free survival, and in chondrosarcoma with shorter overall survival.65 Our findings suggest that copy number increase at 8q24.1–q24.3 might reflect the aggressiveness and dissemination capacity of the tumor.
In conclusion, we found some typical and nearly consistent DNA copy number changes, for example, losses in 10q, 13q, and 16q, gains in 1q, and gains and high-level amplifications in 17p. We revealed two clusters that significantly segregated uterine and non-uterine leiomyosarcomas and found that the non-uterine cluster showed a trend towards increased metastasis-free survival. Further explorative analyses identified aberrations associated with shorter metastasis-free survival, that is losses at 2q32.1–q37 and gains at 8q24.1–q24.3. In contrast, the cases with losses at 6cen-p25 showed longer metastasis-free survival. The true prognostic impact of these aberrations should be verified on an independent sample set.
References
Weiss SW, Goldblum JR . Leiomyosarcoma. In: Weiss SW, Goldblum JR (eds). Enzinger and Weiss's Soft Tissue Tumors, 4th edn. Mosby Harcourt: Philadelphia, 2001, pp 727–748.
Miettinen M . Smooth muscle tumors. In: Miettinen M (ed). Diagnostic soft tissue pathology. 1st edn. Churchill Livingstone: Philadelphia, 2003, pp 241–258.
Evans HL, Shipley J . Leiomyosarcoma. In: Fletcher CDM, Unni KK, Mertens F (eds). World Health Organization classification of tumours. Pathology and genetics of tumours of soft tissue and bone. IARC Press: Lyon, 2002, pp 131–134.
Nordal RR, Thoresen SO . Uterine sarcomas in Norway 1956–1992: Incidence, survival and mortality. Eur J Cancer 1997;33:907–911.
Benoit L, Arnould L, Cheynel N, et al. The role of surgery and treatment trends in uterine sarcoma. Eur J Surg Oncol 2005;31:434–442.
Hashimoto H, Daimaru Y, Tsuneyoshi M, et al. Leiomyosarcoma of the external soft tissues. A clinicopathologic, immunohistochemical, and electron microscopic study. Cancer 1986;57:2077–2088.
Gustafson P, Willén H, Baldetorp B, et al. Soft tissue leiomyosarcoma. A population-based epidemiologic and prognostic study of 48 patients, including cellular DNA content. Cancer 1992;70:114–119.
Farshid G, Pradhan M, Goldblum J, et al. Leiomyosarcoma of somatic soft tissues. A tumor of vascular origin with multivariate analysis of outcome in 42 cases. Am J Surg Pathol 2002;26:14–24.
Mankin HJ, Casas-Ganem J, Kim JI, et al. Leiomyosarcoma of somatic soft tissues. Clin Orthop Relat Res 2004;1:225–231.
Miyajima K, Oda Y, Oshiro Y, et al. Clinicopathological prognostic factors in soft tissue leiomyosarcoma: A multivariate analysis. Histopathology 2002;40:353–359.
Jensen ML, Jensen OM, Michalski W, et al. Intradermal and subcutaneous leiomyosarcoma: A clinicopathological and immunohistochemical study of 41 cases. J Cutan Pathol 1996;23:458–463.
Kaddu S, Beham A, Cerroni L, et al. Cutaneous leiomyosarcoma. Am J Surg Pathol 1997;21:979–987.
Sandberg AA . Updates on the cytogenetics and molecular genetics of bone and soft tissue tumors: Leiomyosarcoma. Cancer Genet Cytogenet 2005;161:1–19.
Mitelman F, Johansson B, Mertens F, (eds). Mitelman database of chromosome aberrations in cancer. 2006; http://cgap.nci.nih.gov/Chromosomes/Mitelman.
Boghosian L, Dal Cin P, Turc-Carel C, et al. Three possible cytogenetic subgroups of leiomyosarcoma. Cancer Genet Cytogenet 1989;43:39–49.
Sreekantaiah C, Davis JR, Sandberg AA . Chromosomal abnormalities in leiomyosarcomas. Am J Pathol 1993;142:293–305.
Mertens F, Fletcher CD, Dal Cin P, et al. Cytogenetic analysis of 46 pleomorphic soft tissue sarcomas and correlation with morphologic and clinical features: a report of the CHAMP study group. Genes Chromosomes Cancer 1998;22:16–25.
Mandahl N, Fletcher CD, Dal Cin P, et al. Comparative cytogenetic study of spindle cell and pleomorphic leiomyosarcomas of soft tissues: a report from the CHAMP study group. Cancer Genet Cytogenet 2000;116:66–73.
Miyajima K, Oda Y, Tamiya S, et al. Cytogenetic and clinicopathological analysis of soft-tissue leiomyosarcomas. Pathol Int 2003;53:163–168.
Packenham JP, du Manoir S, Schrock E, et al. Analysis of genetic alterations in uterine leiomyomas and leiomyosarcomas by comparative genomic hybridization. Mol Carcinog 1997;19:273–279.
El-Rifai W, Sarlomo-Rikala M, Knuutila S, et al. DNA copy number changes in development and progression in leiomyosarcomas of soft tissues. Am J Pathol 1998;153:985–990.
Parente F, Grosgeorge J, Coindre JM, et al. Comparative genomic hybridization reveals novel chromosome deletions in 90 primary soft tissue tumors. Cancer Genet Cytogenet 1999;115:89–95.
Otaño-Joos M, Mechtersheimer G, Ohl S, et al. Detection of chromosomal imbalances in leiomyosarcoma by comparative genomic hybridization and interphase cytogenetics. Cytogenet Cell Genet 2000;90:86–92.
Levy B, Mukherjee T, Hirschhorn K . Molecular cytogenetic analysis of uterine leiomyoma and leiomyosarcoma by comparative genomic hybridization. Cancer Genet Cytogenet 2000;121:1–8.
Wang R, Lu YJ, Fisher C, et al. Characterization of chromosome aberrations associated with soft-tissue leiomyosarcomas by twenty-four-color karyotyping and comparative genomic hybridization analysis. Genes Chromosomes Cancer 2001;31:54–64.
Derré J, Lagace R, Nicolas A, et al. Leiomyosarcomas and most malignant fibrous histiocytomas share very similar comparative genomic hybridization imbalances: an analysis of a series of 27 leiomyosarcomas. Lab Invest 2001;81:211–215.
Hu J, Khanna V, Jones M, et al. Genomic alterations in uterine leiomyosarcomas: potential markers for clinical diagnosis and prognosis. Genes Chromosomes Cancer 2001;31:117–124.
Wang R, Titley JC, Lu YJ, et al. Loss of 13q14-q21 and gain of 5p14-pter in the progression of leiomyosarcoma. Mod Pathol 2003;16:778–785.
Giuntoli 2nd RL, Metzinger DS, DiMarco CS, et al. Retrospective review of 208 patients with leiomyosarcoma of the uterus: prognostic indicators, surgical management, and adjuvant therapy. Gynecol Oncol 2003;89:460–469.
Dinh TA, Oliva EA, Fuller Jr AF, et al. The treatment of uterine leiomyosarcoma. Results from a 10-year experience (1990–1999) at the Massachusetts General Hospital. Gynecol Oncol 2004;92:648–652.
Sutton G, Blessing J, Hanjani P, et al. Phase II evaluation of liposomal doxorubicin (Doxil) in recurrent or advanced leiomyosarcoma of the uterus: a Gynecologic Oncology Group study. Gynecol Oncol 2005;96:749–752.
Bell SW, Kempson RL, Hendrickson MR . Problematic uterine smooth muscle neoplasms. A clinicopathologic study of 213 cases. Am J Surg Pathol 1994;18:535–558.
Isola J, DeVries S, Chu L, et al. Analysis of changes in DNA sequence copy number by comparative genomic hybridization in archival paraffin-embedded tumor samples. Am J Pathol 1994;145:1301–1308.
Larramendy ML, Tarkkanen M, Valle J, et al. Gains, losses, and amplifications of DNA sequences evaluated by comparative genomic hybridization in chondrosarcomas. Am J Pathol 1997;150:685–691.
El-Rifai W, Larramendy ML, Björkqvist AM, et al. Optimization of comparative genomic hybridization using fluorochrome conjugated to dCTP and dUTP nucleotides. Lab Invest 1997;77:699–700.
Larramendy ML, El-Rifai W, Knuutila S . Comparison of fluorescein isothiocyanate- and Texas red-conjugated nucleotides for direct labeling in comparative genomic hybridization. Cytometry 1998;31:174–179.
Kallioniemi OP, Kallioniemi A, Piper J, et al. Optimizing comparative genomic hybridization for analysis of DNA sequence copy number changes in solid tumors. Genes Chromosomes Cancer 1994;10:231–243.
Golub TR, Slonim DK, Tamayo P, et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 1999;286:531–537.
Chibon F, Mariani O, Mairal A, et al. The use of clustering software for the classification of comparative genomic hybridization data. An analysis of 109 malignant fibrous histiocytomas. Cancer Genet Cytogenet 2003;141:75–78.
Karpeh MS, Brennan MF, Cance WG, et al. Altered patterns of retinoblastoma gene product expression in adult soft-tissue sarcomas. Br J Cancer 1995;72:986–991.
Knuutila S, Aalto Y, Autio K, et al. DNA copy number losses in human neoplasms. Am J Pathol 1999;155:683–694.
Dei Tos AP, Maestro R, Doglioni C, et al. Tumor suppressor genes and related molecules in leiomyosarcoma. Am J Pathol 1996;148:1037–1045.
Stratton MR, Williams S, Fisher C, et al. Structural alterations of the RB1 gene in human soft tissue tumours. Br J Cancer 1989;60:202–205.
Lee J, Li S, Torbenson M, Liu QZ, et al. Leiomyosarcoma of the breast: a pathologic and comparative genomic hybridization study of two cases. Cancer Genet Cytogenet 2004;149:53–57.
Li J, Yen C, Liaw D, et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 1997;275:1943–1947.
Quade BJ, Pinto AP, Howard DR, et al. Frequent loss of heterozygosity for chromosome 10 in uterine leiomyosarcoma in contrast to leiomyoma. Am J Pathol 1999;154:945–950.
Wechsler DS, Shelly CA, Dang CV . Genomic organization of human MXI1, a putative tumor suppressor gene. Genomics 1996;32:466–470.
Mayol X, Grana X, Baldi A, et al. Cloning of a new member of the retinoblastoma gene family (pRb2) which binds to the E1A transforming domain. Oncogene 1993;8:2561–2566.
Skubitz KM, Skubitz AP . Differential gene expression in leiomyosarcoma. Cancer 2003;98:1029–1038.
Knuutila S, Björkqvist AM, Autio K, et al. DNA copy number amplifications in human neoplasms: review of comparative genomic hybridization studies. Am J Pathol 1998;152:1107–1123.
Tarkkanen M, Karhu R, Kallioniemi A, et al. Gains and losses of DNA sequences in osteosarcomas by comparative genomic hybridization. Cancer Res 1995;55:1334–1338.
Forus A, Weghuis DO, Smeets D, et al. Comparative genomic hybridization analysis of human sarcomas: II. Identification of novel amplicons at 6p and 17p in osteosarcomas. Genes Chromosomes Cancer 1995;14:15–21.
Larramendy ML, Tarkkanen M, Blomqvist C, et al. Comparative genomic hybridization of malignant fibrous histiocytoma reveals a novel prognostic marker. Am J Pathol 1997;151:1153–1161.
Szymanska J, Tarkkanen M, Wiklund T, et al. Gains and losses of DNA sequences in liposarcomas evaluated by comparative genomic hybridization. Genes Chromosomes Cancer 1996;15:89–94.
Armengol G, Tarkkanen M, Virolainen M, et al. Recurrent gains of 1q, 8 and 12 in the Ewing family of tumours by comparative genomic hybridization. Br J Cancer 1997;75:1403–1409.
Forus A, Berner JM, Meza-Zepeda LA, et al. Molecular characterization of a novel amplicon at 1q21-q22 frequently observed in human sarcomas. Br J Cancer 1998;78:495–503.
Forus A, Weghuis DO, Smeets D, et al. Comparative genomic hybridization analysis of human sarcomas: I. Occurrence of genomic imbalances and identification of a novel major amplicon at 1q21–q22 in soft tissue sarcomas. Genes Chromosomes Cancer 1995;14:8–14.
Forus A, D'Angelo A, Henriksen J, et al. Amplification and overexpression of PRUNE in human sarcomas and breast carcinomas-a possible mechanism for altering the nm23-H1 activity. Oncogene 2001;20:6881–6890.
Patterson H, Gill S, Fisher C, et al. Abnormalities of the p53 MDM2 and DCC genes in human leiomyosarcomas. Br J Cancer 1994;69:1052–1058.
Yoo J, Lee HK, Kang CS, et al. p53 gene mutations and p53 protein expression in human soft tissue sarcomas. Arch Pathol Lab Med 1997;121:395–399.
Mittal K, Demopoulos RI . MIB-1 (ki-67), p53, estrogen receptor, and progesterone receptor expression in uterine smooth muscle tumors. Hum Pathol 2001;32:984–987.
Miyajima K, Tamiya S, Oda Y, et al. Relative quantitation of p53 and MDM2 gene expression in leiomyosarcoma; real-time semi-quantitative reverse transcription-polymerase chain reaction. Cancer Lett 2001;164:177–188.
Wolf M, Tarkkanen M, Hulsebos T, et al. Characterization of the 17p amplicon in human sarcomas: microsatellite marker analysis. Int J Cancer 1999;82:329–333.
Van Dartel M, Cornelissen PW, Redeker S, et al. Amplification of 17p11.2 approximately p12, including PMP22, TOP3A, and MAPK7, in high-grade osteosarcoma. Cancer Genet Cytogenet 2002;139:91–96.
Tarkkanen M, Elomaa I, Blomqvist C, et al. DNA sequence copy number increase at 8q: a potential new prognostic marker in high-grade osteosarcoma. Int J Cancer 1999;84:114–121.
Rieker RJ, Joos S, Bartsch C, et al. Distinct chromosomal imbalances in pleomorphic and in high-grade dedifferentiated liposarcomas. Int J Cancer 200;99:68–73.
Weissman AM . Regulating protein degradation by ubiquitination. Immunol Today 1997;18:189–198.
Tarkkanen M, Kiuru-Kuhlefelt S, Blomqvist C, et al. Clinical correlations of genetic changes by comparative genomic hybridization in Ewing sarcoma and related tumors. Cancer Genet Cytogenet 1999;114:35–41.
Skytting BT, Szymanska J, Aalto Y, et al. Clinical importance of genomic imbalances in synovial sarcoma evaluated by comparative genomic hybridization. Cancer Genet Cytogenet 1999;115:39–46.
Acknowledgements
This study was supported by Finska Läkaresällskapet, K Albin Johansson Foundation, Kurt and Doris Palander Foundation, Helsinki University Central Hospital research funds, the Finnish Cancer Society, and Sigrid Jusélius Foundation in Finland, and by National Council of Scientific and Technological Research (CONICET) and National University of La Plata in Argentina (Grant 11/N325).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Modern Pathology website (http://www.nature.com/modpathol)
Supplementary information
Rights and permissions
About this article
Cite this article
Svarvar, C., Larramendy, M., Blomqvist, C. et al. Do DNA copy number changes differentiate uterine from non-uterine leiomyosarcomas and predict metastasis?. Mod Pathol 19, 1068–1082 (2006). https://doi.org/10.1038/modpathol.3800617
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800617